

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148817.2 - phase: 0
(608 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
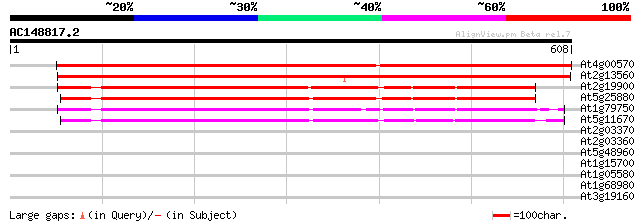
Score E
Sequences producing significant alignments: (bits) Value
At4g00570 putative malate oxidoreductase 918 0.0
At2g13560 malate oxidoreductase (malic enzyme) 748 0.0
At2g19900 malate oxidoreductase (malic enzyme) 416 e-116
At5g25880 malate dehydrogenase - like protein 414 e-116
At1g79750 malate oxidoreductase like protein 408 e-114
At5g11670 NADP dependent malic enzyme - like protein 407 e-114
At2g03370 unknown protein 38 0.019
At2g03360 unknown protein 32 1.0
At5g48960 unknown protein 30 3.0
At1g15700 unknown protein 30 3.0
At1g05580 unknown protein 30 3.9
At1g68980 unknown protein 29 6.7
At3g19160 tRNA isopentenyl transferase, putative 29 8.7
>At4g00570 putative malate oxidoreductase
Length = 607
Score = 918 bits (2373), Expect = 0.0
Identities = 445/558 (79%), Positives = 502/558 (89%), Gaps = 3/558 (0%)
Query: 51 GFPLTERDRLGLRGLLPPRIISFQEQYDRFMSSYRSLEKNTHGQSDKIVSLSKWRILNRL 110
GFPLTERDRLG+RGLLPPR+++ +Q DRF+ S+RSLE NT G+ + +V+L+KWR+LNRL
Sbjct: 53 GFPLTERDRLGIRGLLPPRVMTCVQQCDRFIESFRSLENNTKGEPENVVALAKWRMLNRL 112
Query: 111 HDRNETLYYRALIDNIKEFAPIIYTPTVGLVCQNYCGLYRRPRGMYFSAKDKGEMMSMIY 170
HDRNETLYYR LIDNIK+FAPIIYTPTVGLVCQNY GLYRRPRGMYFSAKDKGEMMSMIY
Sbjct: 113 HDRNETLYYRVLIDNIKDFAPIIYTPTVGLVCQNYSGLYRRPRGMYFSAKDKGEMMSMIY 172
Query: 171 NWPAPQVDMIVLTDGSRILGLGDLGVQGIGIPVGKLDMYVAAAGINPQRILPIMLDVGTN 230
NWPAPQVDMIV+TDGSRILGLGDLGVQGIGIP+GKLDMYVAAAGINPQR+LPIMLDVGTN
Sbjct: 173 NWPAPQVDMIVITDGSRILGLGDLGVQGIGIPIGKLDMYVAAAGINPQRVLPIMLDVGTN 232
Query: 231 NQKLLDDRLYLGLRQPRLEGEEYLSIVDEFMEAVHARWPKAIVQFEDFQMKWAFETLKRY 290
N+KLL + LYLG+RQPRLEGEEYL I+DEFMEA RWPKA+VQFEDFQ KWAF TL+RY
Sbjct: 233 NEKLLQNDLYLGVRQPRLEGEEYLEIIDEFMEAAFTRWPKAVVQFEDFQAKWAFGTLERY 292
Query: 291 RHKFCMFNDDIQGTAGVALAGLLGTVRAQGRPLSDFAKQKIVIVGAGSAGLGVLNMAVQA 350
R KFCMFNDD+QGTAGVALAGLLGTVRAQGRP+SDF QKIV+VGAGSAGLGV MAVQA
Sbjct: 293 RKKFCMFNDDVQGTAGVALAGLLGTVRAQGRPISDFVNQKIVVVGAGSAGLGVTKMAVQA 352
Query: 351 VSRMSGCSETAAKSQFFLIDKNGLVTTERSNLDPAAVPFAKNPRDLEGLAEGASIIEVVK 410
V+RM+G SE+ A F+LIDK+GLVTTER+ LDP AV FAKNP ++ EGASI+EVVK
Sbjct: 353 VARMAGISESEATKNFYLIDKDGLVTTERTKLDPGAVLFAKNPAEIR---EGASIVEVVK 409
Query: 411 KVKPHVLLGLSGVGGVFNEQVLKAMRESVSTKPAIFAMSNPTMNAECTAIDAFNHAGENI 470
KV+PHVLLGLSGVGG+FNE+VLKAMRES S KPAIFAMSNPT+NAECTA DAF HAG NI
Sbjct: 410 KVRPHVLLGLSGVGGIFNEEVLKAMRESDSCKPAIFAMSNPTLNAECTAADAFKHAGGNI 469
Query: 471 VFASGSPFENVDLGNGRAGHVNQANNMYLFPGIGLGSLLSGAHHITDGMLQAASECLASY 530
VFASGSPFENV+L NG+ GHVNQANNMYLFPGIGLG+LLSGA +TDGMLQAASECLASY
Sbjct: 470 VFASGSPFENVELENGKVGHVNQANNMYLFPGIGLGTLLSGARIVTDGMLQAASECLASY 529
Query: 531 MTEEDIQKGILYPSIDCIRNVTAEVGAAVLRAAIAEGLAEGHGGVGSKELEHMSEDDTVE 590
MT+E++QKGILYPSI+ IR++TAEVGAAVLRAA+ + +AEGHG VG K+L HMS++DTV
Sbjct: 530 MTDEEVQKGILYPSINNIRHITAEVGAAVLRAAVTDDIAEGHGDVGPKDLSHMSKEDTVN 589
Query: 591 YVRGNMWYPEYCPLVHEK 608
Y+ NMW+P Y PLVHEK
Sbjct: 590 YITRNMWFPVYSPLVHEK 607
>At2g13560 malate oxidoreductase (malic enzyme)
Length = 623
Score = 748 bits (1932), Expect = 0.0
Identities = 359/561 (63%), Positives = 459/561 (80%), Gaps = 5/561 (0%)
Query: 52 FPLTERDRLGLRGLLPPRIISFQEQYDRFMSSYRSLEKNTHGQSDKIVSLSKWRILNRLH 111
F +TER+RL LRGLLPP ++ ++Q RFM+ + LE+ +L+KWRILNRLH
Sbjct: 61 FTMTERNRLDLRGLLPPNVMDSEQQIFRFMTDLKRLEEQARDGPSDPNALAKWRILNRLH 120
Query: 112 DRNETLYYRALIDNIKEFAPIIYTPTVGLVCQNYCGLYRRPRGMYFSAKDKGEMMSMIYN 171
DRNET+YY+ LI+NI+E+API+YTPTVGLVCQNY GL+RRPRGMYFSA+D+GEMMSM+YN
Sbjct: 121 DRNETMYYKVLINNIEEYAPIVYTPTVGLVCQNYSGLFRRPRGMYFSAEDRGEMMSMVYN 180
Query: 172 WPAPQVDMIVLTDGSRILGLGDLGVQGIGIPVGKLDMYVAAAGINPQRILPIMLDVGTNN 231
WPA QVDMIV+TDGSRILGLGDLGV GIGI VGKLD+YVAAAGINPQR+LP+M+DVGTNN
Sbjct: 181 WPAEQVDMIVVTDGSRILGLGDLGVHGIGIAVGKLDLYVAAAGINPQRVLPVMIDVGTNN 240
Query: 232 QKLLDDRLYLGLRQPRLEGEEYLSIVDEFMEAVHARWPKAIVQFEDFQMKWAFETLKRYR 291
+KL +D +YLGL+Q RLE ++Y+ ++DEFMEAV+ RWP IVQFEDFQ KWAF+ L+RYR
Sbjct: 241 EKLRNDPMYLGLQQRRLEDDDYIDVIDEFMEAVYTRWPHVIVQFEDFQSKWAFKLLQRYR 300
Query: 292 HKFCMFNDDIQGTAGVALAGLLGTVRAQGRPLSDFAKQKIVIVGAGSAGLGVLNMAVQAV 351
+ MFNDD+QGTAGVA+AGLLG VRAQGRP+ DF K KIV+ GAGSAG+GVLN A + +
Sbjct: 301 CTYRMFNDDVQGTAGVAIAGLLGAVRAQGRPMIDFPKMKIVVAGAGSAGIGVLNAARKTM 360
Query: 352 SRMSGCSETA---AKSQFFLIDKNGLVTTERSNLDPAAVPFAKNPRDLE--GLAEGASII 406
+RM G +ETA A+SQF+++D GL+T R N+DP A PFA+ +++E GL EGA+++
Sbjct: 361 ARMLGNTETAFDSAQSQFWVVDAQGLITEGRENIDPEAQPFARKTKEMERQGLKEGATLV 420
Query: 407 EVVKKVKPHVLLGLSGVGGVFNEQVLKAMRESVSTKPAIFAMSNPTMNAECTAIDAFNHA 466
EVV++VKP VLLGLS VGG+F+++VL+AM+ S ST+PAIFAMSNPT NAECT DAF+
Sbjct: 421 EVVREVKPDVLLGLSAVGGLFSKEVLEAMKGSTSTRPAIFAMSNPTKNAECTPQDAFSIL 480
Query: 467 GENIVFASGSPFENVDLGNGRAGHVNQANNMYLFPGIGLGSLLSGAHHITDGMLQAASEC 526
GEN++FASGSPF+NV+ GNG GH NQ NNMYLFPGIGLG+LLSGA ++DGMLQAASEC
Sbjct: 481 GENMIFASGSPFKNVEFGNGHVGHCNQGNNMYLFPGIGLGTLLSGAPIVSDGMLQAASEC 540
Query: 527 LASYMTEEDIQKGILYPSIDCIRNVTAEVGAAVLRAAIAEGLAEGHGGVGSKELEHMSED 586
LA+YM+EE++ +GI+YP I IR++T + AAV++ AI E L EG+ + ++E++ + E+
Sbjct: 541 LAAYMSEEEVLEGIIYPPISRIRDITKRIAAAVIKEAIEEDLVEGYREMDAREIQKLDEE 600
Query: 587 DTVEYVRGNMWYPEYCPLVHE 607
+EYV NMW PEY LV++
Sbjct: 601 GLMEYVENNMWNPEYPTLVYK 621
>At2g19900 malate oxidoreductase (malic enzyme)
Length = 581
Score = 416 bits (1069), Expect = e-116
Identities = 221/520 (42%), Positives = 321/520 (61%), Gaps = 23/520 (4%)
Query: 52 FPLTERDRLGLRGLLPPRIISFQEQYDRFMSSYRSLEKNTHGQSDKIVSLSKWRILNRLH 111
F ERD LRGLLPP ++ + Q R +++ R + L K+ L L
Sbjct: 57 FTEKERDTHYLRGLLPPVVLDQKLQEKRLLNNIRQYQ----------FPLQKYMALTELQ 106
Query: 112 DRNETLYYRALIDNIKEFAPIIYTPTVGLVCQNYCGLYRRPRGMYFSAKDKGEMMSMIYN 171
+RNE L+Y+ LIDN++E PI+YTPTVG CQ + ++RRP+G++ S KDKG+++ ++ N
Sbjct: 107 ERNERLFYKLLIDNVEELLPIVYTPTVGEACQKFGSIFRRPQGLFISLKDKGKILDVLKN 166
Query: 172 WPAPQVDMIVLTDGSRILGLGDLGVQGIGIPVGKLDMYVAAAGINPQRILPIMLDVGTNN 231
WP + +IV+TDG RILGLGDLG QG+GIPVGKL +Y A G+ P LP+ +DVGTNN
Sbjct: 167 WPERNIQVIVVTDGERILGLGDLGCQGMGIPVGKLALYSALGGVRPSACLPVTIDVGTNN 226
Query: 232 QKLLDDRLYLGLRQPRLEGEEYLSIVDEFMEAVHARW-PKAIVQFEDFQMKWAFETLKRY 290
+KLL+D Y+GLRQ R G+EY +++EFM AV + K ++QFEDF AFE L +Y
Sbjct: 227 EKLLNDEFYIGLRQKRATGQEYSELLNEFMSAVKQNYGEKVLIQFEDFANHNAFELLAKY 286
Query: 291 RHKFCMFNDDIQGTAGVALAGLLGTVRAQGRPLSDFAKQKIVIVGAGSAGLGVLNMAVQA 350
+FNDDIQGTA V LAGL+ + PL A+ + +GAG AG G+ +
Sbjct: 287 SDTHLVFNDDIQGTASVVLAGLVSAQKLTNSPL---AEHTFLFLGAGEAGTGIAELIALY 343
Query: 351 VSRMSGCSETAAKSQFFLIDKNGLVTTER-SNLDPAAVPFAKNPRDLEGLAEGASIIEVV 409
+S+ S ++ + +L+D GL+ R +L P+A ++ L + +
Sbjct: 344 MSKQMNASVEESRKKIWLVDSKGLIVNSRKDSLQDFKKPWAHEHEPVKDL------LGAI 397
Query: 410 KKVKPHVLLGLSGVGGVFNEQVLKAMRESVSTKPAIFAMSNPTMNAECTAIDAFNHAGEN 469
K +KP VL+G SGVG F ++V++AM S++ +P I A+SNPT +ECTA +A+ +
Sbjct: 398 KAIKPTVLIGSSGVGRSFTKEVIEAM-SSINERPLIMALSNPTTQSECTAEEAYTWSKGR 456
Query: 470 IVFASGSPFENVDLGNGRAGHVNQANNMYLFPGIGLGSLLSGAHHITDGMLQAASECLAS 529
+FASGSPF+ V+ G+ QANN Y+FPG GLG ++SGA + D ML AA+E LA
Sbjct: 457 AIFASGSPFDPVEY-EGKVFVSTQANNAYIFPGFGLGLVISGAIRVHDDMLLAAAEALAG 515
Query: 530 YMTEEDIQKGILYPSIDCIRNVTAEVGAAVLRAAIAEGLA 569
+++E+ +KG++YPS IR ++A++ A V A GLA
Sbjct: 516 QVSKENYEKGMIYPSFSSIRKISAQIAANVATKAYELGLA 555
>At5g25880 malate dehydrogenase - like protein
Length = 588
Score = 414 bits (1064), Expect = e-116
Identities = 224/516 (43%), Positives = 317/516 (61%), Gaps = 23/516 (4%)
Query: 56 ERDRLGLRGLLPPRIISFQEQYDRFMSSYRSLEKNTHGQSDKIVSLSKWRILNRLHDRNE 115
ERD + GLLPP ++S Q + M + R V L ++ L L +RNE
Sbjct: 68 ERDAHYITGLLPPVVLSQDVQERKVMHNLRQYT----------VPLQRYMALMDLQERNE 117
Query: 116 TLYYRALIDNIKEFAPIIYTPTVGLVCQNYCGLYRRPRGMYFSAKDKGEMMSMIYNWPAP 175
L+Y+ LIDN++E P++YTPTVG CQ Y +YRRP+G+Y S K+KG+++ ++ NWP
Sbjct: 118 RLFYKLLIDNVEELLPVVYTPTVGEACQKYGSIYRRPQGLYISLKEKGKILEVLKNWPQR 177
Query: 176 QVDMIVLTDGSRILGLGDLGVQGIGIPVGKLDMYVAAAGINPQRILPIMLDVGTNNQKLL 235
+ +IV+TDG RILGLGDLG QG+GIPVGKL +Y A GI P LPI +DVGTNN+KLL
Sbjct: 178 GIQVIVVTDGERILGLGDLGCQGMGIPVGKLSLYTALGGIRPSACLPITIDVGTNNEKLL 237
Query: 236 DDRLYLGLRQPRLEGEEYLSIVDEFMEAVHARW-PKAIVQFEDFQMKWAFETLKRYRHKF 294
++ Y+GL+Q R GEEY + EFM AV + K +VQFEDF AFE L +Y
Sbjct: 238 NNEFYIGLKQKRANGEEYAEFLQEFMCAVKQNYGEKVLVQFEDFANHHAFELLSKYCSSH 297
Query: 295 CMFNDDIQGTAGVALAGLLGTVRAQGRPLSDFAKQKIVIVGAGSAGLGVLNMAVQAVSRM 354
+FNDDIQGTA V LAGL+ + G+ L+D + +GAG AG G+ + +S+
Sbjct: 298 LVFNDDIQGTASVVLAGLIAAQKVLGKSLAD---HTFLFLGAGEAGTGIAELIALKISKE 354
Query: 355 SGCSETAAKSQFFLIDKNGLVTTER-SNLDPAAVPFAKNPRDLEGLAEGASIIEVVKKVK 413
+G + + +L+D GL+ +ER +L P+A + + ++ ++ V +K
Sbjct: 355 TGKPIDETRKKIWLVDSKGLIVSERKESLQHFKQPWAHDHKPVK------ELLAAVNAIK 408
Query: 414 PHVLLGLSGVGGVFNEQVLKAMRESVSTKPAIFAMSNPTMNAECTAIDAFNHAGENIVFA 473
P VL+G SGVG F ++V++AM +++ KP I A+SNPT AECTA +A+ +FA
Sbjct: 409 PTVLIGTSGVGKTFTKEVVEAM-ATLNEKPLILALSNPTSQAECTAEEAYTWTKGRAIFA 467
Query: 474 SGSPFENVDLGNGRAGHVNQANNMYLFPGIGLGSLLSGAHHITDGMLQAASECLASYMTE 533
SGSPF+ V +G+ QANN Y+FPG+GLG ++SGA + D ML AASE LAS +TE
Sbjct: 468 SGSPFDPVQY-DGKKFTPGQANNCYIFPGLGLGLIMSGAIRVRDDMLLAASEALASQVTE 526
Query: 534 EDIQKGILYPSIDCIRNVTAEVGAAVLRAAIAEGLA 569
E+ G++YP IR ++A + A+V GLA
Sbjct: 527 ENFANGLIYPPFANIRKISANIAASVGAKTYELGLA 562
>At1g79750 malate oxidoreductase like protein
Length = 646
Score = 408 bits (1049), Expect = e-114
Identities = 231/553 (41%), Positives = 324/553 (57%), Gaps = 35/553 (6%)
Query: 52 FPLTERDRLGLRGLLPPRIISFQEQYDRFMSSYRSLEKNTHGQSDKIVSLSKWRILNRLH 111
F ERD LRGLLPP +IS Q + M + R + V L K+ + L
Sbjct: 122 FSHRERDAHYLRGLLPPTVISQDLQVKKIMHTLRQYQ----------VPLQKYMAMMDLQ 171
Query: 112 DRNETLYYRALIDNIKEFAPIIYTPTVGLVCQNYCGLYRRPRGMYFSAKDKGEMMSMIYN 171
+ NE L+Y+ LID+++E P+IYTPTVG CQ Y ++ RP+G++ S K+KG++ ++ N
Sbjct: 172 ETNERLFYKLLIDHVEELLPVIYTPTVGEACQKYGSIFLRPQGLFISLKEKGKIHEVLRN 231
Query: 172 WPAPQVDMIVLTDGSRILGLGDLGVQGIGIPVGKLDMYVAAAGINPQRILPIMLDVGTNN 231
WP + +IV+TDG RILGLGDLG QG+GIPVGKL +Y A G+ P LP+ +DVGTNN
Sbjct: 232 WPEKNIQVIVVTDGERILGLGDLGCQGMGIPVGKLSLYTALGGVRPSACLPVTIDVGTNN 291
Query: 232 QKLLDDRLYLGLRQPRLEGEEYLSIVDEFMEAVHARW-PKAIVQFEDFQMKWAFETLKRY 290
+KLL+D Y+GLRQ R GEEY ++ EFM AV + K ++QFEDF AF+ L +Y
Sbjct: 292 EKLLNDEFYIGLRQRRATGEEYSELMHEFMTAVKQNYGEKVVIQFEDFANHNAFDLLAKY 351
Query: 291 RHKFCMFNDDIQGTAGVALAGLLGTVRAQGRPLSDFAKQKIVIVGAGSAGLGVLNMAVQA 350
+FNDDIQGTA V LAGL+ +R G LSD + + +GAG AG G+ +
Sbjct: 352 GTTHLVFNDDIQGTASVVLAGLIAALRFVGGSLSD---HRFLFLGAGEAGTGIAELIALE 408
Query: 351 VSRMSGCSETAAKSQFFLIDKNGLVTTERSNLDPAAVPFAKNP--RDLEGLAEGASIIEV 408
+S+ S A+ +L+D GL+ + R ++ K P D E + E +++
Sbjct: 409 ISKKSHIPLEEARKNIWLVDSKGLIVSSRKE----SIQHFKKPWAHDHEPIRE---LVDA 461
Query: 409 VKKVKPHVLLGLSGVGGVFNEQVLKAMRESVSTKPAIFAMSNPTMNAECTAIDAFNHAGE 468
VK +KP VL+G SGVG F + V++ M + ++ KP I ++SNPT +ECTA +A+ +
Sbjct: 462 VKAIKPTVLIGTSGVGQTFTQDVVETMAK-LNEKPIILSLSNPTSQSECTAEEAYTWSQG 520
Query: 469 NIVFASGSPFENVDLGNGRAGHVNQANNMYLFPGIGLGSLLSGAHHITDGMLQAASECLA 528
+FASGSPF V+ G+ QANN Y+FPG GLG ++SG + D ML AASE LA
Sbjct: 521 RAIFASGSPFAPVEY-EGKTFVPGQANNAYIFPGFGLGLIMSGTIRVHDDMLLAASEALA 579
Query: 529 SYMTEEDIQKGILYPSIDCIRNVTAEVGAAVLRAAIAEGLAEGHGGVGSKELEHMSEDDT 588
+ EE +KG++YP IR ++A + A V A GLA KELE +E
Sbjct: 580 EELMEEHYEKGMIYPPFRNIRKISARIAAKVAAKAYELGLATRL--PQPKELEQCAE--- 634
Query: 589 VEYVRGNMWYPEY 601
+M+ P Y
Sbjct: 635 -----SSMYSPSY 642
>At5g11670 NADP dependent malic enzyme - like protein
Length = 588
Score = 407 bits (1046), Expect = e-114
Identities = 227/548 (41%), Positives = 320/548 (57%), Gaps = 33/548 (6%)
Query: 56 ERDRLGLRGLLPPRIISFQEQYDRFMSSYRSLEKNTHGQSDKIVSLSKWRILNRLHDRNE 115
ERD L GLLPP I+S Q + M + R V L ++ L L +RNE
Sbjct: 68 ERDAHYLTGLLPPVILSQDVQERKVMHNLRQYT----------VPLQRYMALMDLQERNE 117
Query: 116 TLYYRALIDNIKEFAPIIYTPTVGLVCQNYCGLYRRPRGMYFSAKDKGEMMSMIYNWPAP 175
L+Y+ LIDN++E P++YTPTVG CQ Y ++R+P+G+Y S +KG+++ ++ NWP
Sbjct: 118 RLFYKLLIDNVEELLPVVYTPTVGEACQKYGSIFRKPQGLYISLNEKGKILEVLKNWPQR 177
Query: 176 QVDMIVLTDGSRILGLGDLGVQGIGIPVGKLDMYVAAAGINPQRILPIMLDVGTNNQKLL 235
+ +IV+TDG RILGLGDLG QG+GIPVGKL +Y A GI P LPI +DVGTNN+KLL
Sbjct: 178 GIQVIVVTDGERILGLGDLGCQGMGIPVGKLSLYTALGGIRPSACLPITIDVGTNNEKLL 237
Query: 236 DDRLYLGLRQPRLEGEEYLSIVDEFMEAVHARW-PKAIVQFEDFQMKWAFETLKRYRHKF 294
+D Y+GL+Q R G+EY + EFM AV + K +VQFEDF AF+ L +Y
Sbjct: 238 NDEFYIGLKQRRATGQEYAEFLHEFMCAVKQNYGEKVLVQFEDFANHNAFDLLSKYSDSH 297
Query: 295 CMFNDDIQGTAGVALAGLLGTVRAQGRPLSDFAKQKIVIVGAGSAGLGVLNMAVQAVSRM 354
+FNDDIQGTA V LAGL+ + G+ L+D + +GAG AG G+ + +S+
Sbjct: 298 LVFNDDIQGTASVVLAGLIAAQKVLGKKLAD---HTFLFLGAGEAGTGIAELIALKISKE 354
Query: 355 SGCSETAAKSQFFLIDKNGLVTTER-SNLDPAAVPFAKNPRDLEGLAEGASIIEVVKKVK 413
+G T + + +L+D GL+ + R +L P+A + ++ L I V +K
Sbjct: 355 TGAPITETRKKIWLVDSKGLIVSSRKESLQHFKQPWAHEHKPVKDL------IGAVNAIK 408
Query: 414 PHVLLGLSGVGGVFNEQVLKAMRESVSTKPAIFAMSNPTMNAECTAIDAFNHAGENIVFA 473
P VL+G SGVG F ++V++AM + + KP I A+SNPT AECTA A+ +F
Sbjct: 409 PTVLIGTSGVGQTFTKEVVEAMATN-NEKPLILALSNPTSQAECTAEQAYTWTKGRAIFG 467
Query: 474 SGSPFENVDLGNGRAGHVNQANNMYLFPGIGLGSLLSGAHHITDGMLQAASECLASYMTE 533
SGSPF+ V + +G+ QANN Y+FPG+GLG ++SGA + D ML AASE LA+ +TE
Sbjct: 468 SGSPFDPV-VYDGKTYLPGQANNCYIFPGLGLGLIMSGAIRVRDDMLLAASEALAAQVTE 526
Query: 534 EDIQKGILYPSIDCIRNVTAEVGAAVLRAAIAEGLAEGHGGVGSKELEHMSEDDTVEYVR 593
E G++YP IR ++A + A V GLA D V++
Sbjct: 527 EHYANGLIYPPFSNIREISANIAACVAAKTYDLGLAS----------NLPRAKDLVKFAE 576
Query: 594 GNMWYPEY 601
+M+ P Y
Sbjct: 577 SSMYSPVY 584
>At2g03370 unknown protein
Length = 458
Score = 37.7 bits (86), Expect = 0.019
Identities = 34/113 (30%), Positives = 50/113 (44%), Gaps = 7/113 (6%)
Query: 359 ETAAKSQFFLIDKNGLVTTERSNLDPAAVPFAKNPRDLEGLAEGA--SIIEVVKKVKPHV 416
E+ A + F GL+T +DP +P +K+ D L E A + I K KP +
Sbjct: 224 ESVATTHCFTSAIVGLITHWPMTIDPTQIPNSKSLVDFHNLLEKAFTTNISTPKTHKPRL 283
Query: 417 LLGLSGVGG----VFNEQVLKAMRESVSTKPAIFAMSNPTMNAECTAIDAFNH 465
+L +S G + NEQ +K M E V + IF S T E + +H
Sbjct: 284 ML-VSRYGNIGRVILNEQEIKEMLEDVGFEVIIFRPSKTTNLKEAYKLIKSSH 335
>At2g03360 unknown protein
Length = 393
Score = 32.0 bits (71), Expect = 1.0
Identities = 28/112 (25%), Positives = 49/112 (43%), Gaps = 5/112 (4%)
Query: 359 ETAAKSQFFLIDKNGLVTTERSNLDPAAVPFAKNPRDLEGLAEGA--SIIEVVKKVKPHV 416
E A+ + F GL++ +DP +P +K+ D L + A + ++K KP +
Sbjct: 159 ENASITHCFTSATVGLISHGPMTIDPTQIPNSKSLVDFHNLLDKALNPNLSIIKINKPRL 218
Query: 417 LL--GLSGVGGV-FNEQVLKAMRESVSTKPAIFAMSNPTMNAECTAIDAFNH 465
+L +G V NE+ ++ M E V + F S T E + +H
Sbjct: 219 ILVRRYGNIGRVILNEEEIREMLEDVGFEVITFRPSKTTSLREAYKLIKSSH 270
>At5g48960 unknown protein
Length = 642
Score = 30.4 bits (67), Expect = 3.0
Identities = 36/138 (26%), Positives = 58/138 (41%), Gaps = 21/138 (15%)
Query: 51 GFP---LTERDRLGLRGLLPPRIISFQEQYDRFMSSYRSLEKNTHGQSDKIVS------- 100
GFP L L +RGL+ + + DRF R++ T S+K VS
Sbjct: 211 GFPVDGLAFDPELVIRGLMIDKEKGNLVKADRFGYVKRAMH-GTKMLSNKAVSEIYGREL 269
Query: 101 -----LSKWRILNRLHDRNETLYYRALIDNIKEFAPIIYTPTVGLVCQNYCGLYRRPRGM 155
S+W LN +E L Y ++D + + + +G + +Y GLY+
Sbjct: 270 VDLRNQSRWEFLNTFFSVSEALAYAQMVDRLDDG---FISADLGTL--DYKGLYKAVAKA 324
Query: 156 YFSAKDKGEMMSMIYNWP 173
F A +G++ S I + P
Sbjct: 325 LFRAHVEGQLKSEIMSKP 342
>At1g15700 unknown protein
Length = 386
Score = 30.4 bits (67), Expect = 3.0
Identities = 29/117 (24%), Positives = 53/117 (44%), Gaps = 24/117 (20%)
Query: 344 LNMAVQAVSRMSGCSETAAKSQFFLIDKNGLVTTERSNLD---PAAVPFAKNPRDLEGLA 400
L+M ++ C + F L K+G + ER+ L+ P P + +D
Sbjct: 253 LSMKGESCDVKGECVDAIEDEMFRLTSKDGKLAVERTKLEVEKPEISPLMQFEQD----- 307
Query: 401 EGASIIEVVKKVKPHVLLGLSGVGGVFNEQVLKAMRESVSTKPA--IFAMSNPTMNA 455
++++ + P L N Q+L+A++ES++++ A + AMSN T NA
Sbjct: 308 ----PVQILDAMMPLYL----------NSQILRALQESLASELASRMNAMSNATDNA 350
>At1g05580 unknown protein
Length = 867
Score = 30.0 bits (66), Expect = 3.9
Identities = 24/79 (30%), Positives = 36/79 (45%), Gaps = 9/79 (11%)
Query: 487 RAGHVNQANNMYLFPGIGLGSLLS---GAHHITDGMLQAAS---ECLASYMTEE---DIQ 537
+AGHV + ++ G+ L L++ G H IT L S + + M EE D
Sbjct: 275 KAGHVGDTHVWFILGGVVLCGLITDACGVHSITGAFLFGLSIPHDHIIRNMIEEKLHDFL 334
Query: 538 KGILYPSIDCIRNVTAEVG 556
GIL P I + A++G
Sbjct: 335 SGILMPLFYIICGLRADIG 353
>At1g68980 unknown protein
Length = 619
Score = 29.3 bits (64), Expect = 6.7
Identities = 32/133 (24%), Positives = 54/133 (40%), Gaps = 8/133 (6%)
Query: 426 VFNEQVLKAMRESVSTKPAIFAMSNPTMNAECTAIDAFNHAGENIVFASGSPFENVDLGN 485
VF E A+ E + SN + A C +++ A EN++ E++D+
Sbjct: 167 VFRESCRIAVDEKLDFMKPDLVASNAALEACCRQMESLADA-ENLI-------ESMDVLG 218
Query: 486 GRAGHVNQANNMYLFPGIGLGSLLSGAHHITDGMLQAASECLASYMTEEDIQKGILYPSI 545
+ ++ YL+ GL +S + DG+ A+ L S M ++ G L +
Sbjct: 219 VKPDELSFGFLAYLYARKGLREKISELEDLMDGLGFASRRILYSSMISGYVKSGDLDSAS 278
Query: 546 DCIRNVTAEVGAA 558
D I VG A
Sbjct: 279 DVILCSLKGVGEA 291
>At3g19160 tRNA isopentenyl transferase, putative
Length = 330
Score = 28.9 bits (63), Expect = 8.7
Identities = 19/61 (31%), Positives = 32/61 (52%), Gaps = 3/61 (4%)
Query: 328 KQKIVIVGAGSAGLGVLNMAVQAVSRMSGCSETAAKSQFFLIDKNGLVTTERSNLDPAAV 387
KQK+V++ G+ G G +++ +R SG + K QF+ D + T + S L+ V
Sbjct: 42 KQKVVVI-MGATGSGKSCLSIDLATRFSGEIVNSDKIQFY--DGLKVTTNQMSILERCGV 98
Query: 388 P 388
P
Sbjct: 99 P 99
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.322 0.139 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,466,602
Number of Sequences: 26719
Number of extensions: 578690
Number of successful extensions: 1217
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 1182
Number of HSP's gapped (non-prelim): 15
length of query: 608
length of database: 11,318,596
effective HSP length: 105
effective length of query: 503
effective length of database: 8,513,101
effective search space: 4282089803
effective search space used: 4282089803
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 63 (28.9 bits)
Medicago: description of AC148817.2


