

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148816.5 + phase: 0 /pseudo
(211 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
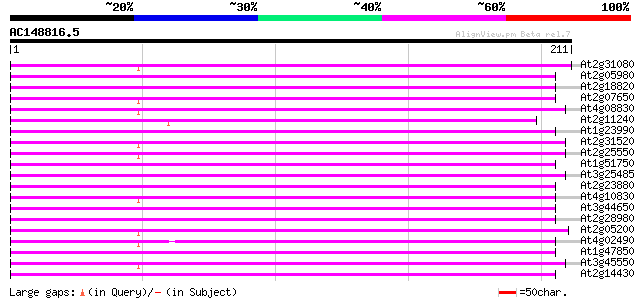
Score E
Sequences producing significant alignments: (bits) Value
At2g31080 putative non-LTR retroelement reverse transcriptase 122 1e-28
At2g05980 putative non-LTR retroelement reverse transcriptase 122 1e-28
At2g18820 putative non-LTR retroelement reverse transcriptase 120 5e-28
At2g07650 putative non-LTR retrolelement reverse transcriptase 119 9e-28
At4g08830 putative protein 117 6e-27
At2g11240 pseudogene 116 1e-26
At1g23990 reverse transcriptase, putative 114 4e-26
At2g31520 putative non-LTR retroelement reverse transcriptase 113 6e-26
At2g25550 putative non-LTR retroelement reverse transcriptase 113 6e-26
At1g51750 hypothetical protein 113 6e-26
At3g25485 unknown protein 112 1e-25
At2g23880 putative non-LTR retroelement reverse transcriptase 112 1e-25
At4g10830 putative protein 112 1e-25
At3g44650 putative protein 111 2e-25
At2g28980 putative non-LTR retroelement reverse transcriptase 110 7e-25
At2g05200 putative non-LTR retroelement reverse transcriptase 110 7e-25
At4g02490 109 9e-25
At1g47850 hypothetical protein 109 9e-25
At3g45550 putative protein 108 2e-24
At2g14430 putative non-LTR retroelement reverse transcriptase 108 2e-24
>At2g31080 putative non-LTR retroelement reverse transcriptase
Length = 1231
Score = 122 bits (306), Expect = 1e-28
Identities = 67/212 (31%), Positives = 112/212 (52%), Gaps = 1/212 (0%)
Query: 1 VSHLQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLV-GINVSEDFMAMA 59
+SH+ +ADD + +ASV + ++ +L F SG K++ KS + NVS + +
Sbjct: 539 LSHVCFADDLILFAEASVAQIRIIRRVLERFCEASGQKVSLEKSKIFFSHNVSREMEQLI 598
Query: 60 CDFLNCSAGSITFKYLGLPVGANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVLLNS 119
+ KYLG+P+ + T+ ++E + RL W R +S GRI L +
Sbjct: 599 SEESGIGCTKELGKYLGMPILQKRMNKETFGEVLERVSARLAGWKGRSLSLAGRITLTKA 658
Query: 120 VLNSMPIFYLSFLKMLVGVWKRIVRIQR*FLWGGVGGGKKISWVNWKSVCQQKENGGVRV 179
VL+S+P+ +S + + V + R R FLWG KK ++W+ +C+ K GG+ +
Sbjct: 659 VLSSIPVHVMSAILLPVSTLDTLDRYSRTFLWGSTMEKKKQHLLSWRKICKPKAEGGIGL 718
Query: 180 KDIRVMNISLLAKWRWRLIDGREALWK*VLVK 211
+ R MN +L+AK WRL+ +E+LW V+ K
Sbjct: 719 RSARDMNKALVAKVGWRLLQDKESLWARVVRK 750
>At2g05980 putative non-LTR retroelement reverse transcriptase
Length = 1352
Score = 122 bits (306), Expect = 1e-28
Identities = 65/205 (31%), Positives = 110/205 (52%)
Query: 1 VSHLQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLVGINVSEDFMAMAC 60
++HL +ADD + + +++ AI F +S LKI+ KS++ +S +
Sbjct: 845 LTHLCFADDIMVFSDGTSKSIQGTLAIFEKFAAMSWLKISLEKSTIFMAGISPNAKTSIL 904
Query: 61 DFLNCSAGSITFKYLGLPVGANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVLLNSV 120
G++ KYLGLP+ + S + PLVE I R+ +W+ R++SF GR+ L+ SV
Sbjct: 905 QQFPFELGTLPVKYLGLPLLTKRMTQSDYLPLVEKIRARITSWTNRFLSFAGRLQLIKSV 964
Query: 121 LNSMPIFYLSFLKMLVGVWKRIVRIQR*FLWGGVGGGKKISWVNWKSVCQQKENGGVRVK 180
L+S+ F+LS ++ + I ++ FLW G K + + W VC+ KE GG+ +K
Sbjct: 965 LSSITNFWLSVFRLPKACLQEIEKMFSAFLWSGPDLNTKKAKIAWSEVCKLKEEGGLGLK 1024
Query: 181 DIRVMNISLLAKWRWRLIDGREALW 205
++ N L K WR++ R++LW
Sbjct: 1025 PLKEANEVSLLKLIWRILSARDSLW 1049
>At2g18820 putative non-LTR retroelement reverse transcriptase
Length = 1412
Score = 120 bits (301), Expect = 5e-28
Identities = 64/205 (31%), Positives = 109/205 (52%)
Query: 1 VSHLQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLVGINVSEDFMAMAC 60
++HL +ADD + S +L + AI + F SGL I+ KS+L ++S + A
Sbjct: 953 LTHLCFADDIMVFSAGSAHSLEGVLAIFKDFAAFSGLNISLEKSTLFMASISSETCASIL 1012
Query: 61 DFLNCSAGSITFKYLGLPVGANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVLLNSV 120
+GS+ +YLGLP+ +++ PL+E I R+++W R++S+ GR+ LLNSV
Sbjct: 1013 ARFPFDSGSLPVRYLGLPLMTKRMTLADCLPLLEKIRSRISSWKNRFLSYAGRLQLLNSV 1072
Query: 121 LNSMPIFYLSFLKMLVGVWKRIVRIQR*FLWGGVGGGKKISWVNWKSVCQQKENGGVRVK 180
++S+ F++S ++ + I +I FLW G + V W VC+ K GG+ ++
Sbjct: 1073 ISSLTKFWISAFRLPRACIREIEQISAAFLWSGTDLNPHKAKVAWHDVCKPKSEGGLGLR 1132
Query: 181 DIRVMNISLLAKWRWRLIDGREALW 205
+ N K WRL+ + +LW
Sbjct: 1133 SLVDANKICCFKLIWRLVSAKHSLW 1157
>At2g07650 putative non-LTR retrolelement reverse transcriptase
Length = 732
Score = 119 bits (299), Expect = 9e-28
Identities = 65/206 (31%), Positives = 111/206 (53%), Gaps = 1/206 (0%)
Query: 1 VSHLQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLV-GINVSEDFMAMA 59
+SH+ +ADD + +ASV + ++ +L F + SG K++ KS + NVS D +
Sbjct: 201 LSHICFADDLILFAEASVAQIRVIRRVLERFCVASGQKVSLEKSKIFFSENVSRDLGKLI 260
Query: 60 CDFLNCSAGSITFKYLGLPVGANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVLLNS 119
D S+ KYLG+PV + T+ ++E + RL W R++S GR+ L +
Sbjct: 261 SDESGISSTRELGKYLGMPVLQRRINKDTFGDILEKLTTRLAGWKGRFLSLAGRVTLTKA 320
Query: 120 VLNSMPIFYLSFLKMLVGVWKRIVRIQR*FLWGGVGGGKKISWVNWKSVCQQKENGGVRV 179
VL+S+P+ +S + + + ++ R FLWG +K ++WK VC+ + GG+ +
Sbjct: 321 VLSSIPVHTMSTIALPKSTLDGLDKVSRSFLWGSSVTQRKQHLISWKRVCKPRSEGGLGI 380
Query: 180 KDIRVMNISLLAKWRWRLIDGREALW 205
+ + MN +LL+K WRLI +LW
Sbjct: 381 RKAQDMNKALLSKVGWRLIQDYHSLW 406
>At4g08830 putative protein
Length = 947
Score = 117 bits (292), Expect = 6e-27
Identities = 67/210 (31%), Positives = 110/210 (51%), Gaps = 1/210 (0%)
Query: 1 VSHLQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLV-GINVSEDFMAMA 59
+SH+ +ADD + +ASV + ++ IL F + SG K++ KS + NVS D +
Sbjct: 353 ISHICFADDLILFAEASVSQIRVIRRILETFCIASGQKVSLDKSKIFFSKNVSRDLEKLI 412
Query: 60 CDFLNCSAGSITFKYLGLPVGANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVLLNS 119
+ KYLG+P+ + T+ ++E + RL W R +SF GR+ L S
Sbjct: 413 SKESGIKSTRELGKYLGMPILQRRINKDTFGEVLERVSSRLAGWKGRSLSFAGRLTLTKS 472
Query: 120 VLNSMPIFYLSFLKMLVGVWKRIVRIQR*FLWGGVGGGKKISWVNWKSVCQQKENGGVRV 179
VL+ +PI +S + + + + ++ R FL G KK+ V W VC K GG+ +
Sbjct: 473 VLSLIPIHTMSTISLPQSTLEGLDKLARVFLLGSSAEKKKLHLVAWDRVCLPKSEGGLGI 532
Query: 180 KDIRVMNISLLAKWRWRLIDGREALWK*VL 209
+ + MN +L++K WRLI+ R +LW +L
Sbjct: 533 RTSKCMNKALVSKVGWRLINDRYSLWARIL 562
>At2g11240 pseudogene
Length = 1044
Score = 116 bits (290), Expect = 1e-26
Identities = 67/199 (33%), Positives = 105/199 (52%), Gaps = 1/199 (0%)
Query: 1 VSHLQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLVGINVSEDFMAM-A 59
V+HL +ADDT+ ++ +++ T IL+ ++ SG IN SKS++ + D + A
Sbjct: 502 VNHLLFADDTIFFCRSDLKSCKTFLCILKKYEEASGQMINKSKSAITFSRKTPDHIKTEA 561
Query: 60 CDFLNCSAGSITFKYLGLPVGANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVLLNS 119
L KYLGLP + + +V+ I R +WS+R++S G+ +L S
Sbjct: 562 QQILGIQLVGGLGKYLGLPKMFGRKKRDLFNQIVDRIRQRSLSWSSRFLSTAGKTTMLKS 621
Query: 120 VLNSMPIFYLSFLKMLVGVWKRIVRIQR*FLWGGVGGGKKISWVNWKSVCQQKENGGVRV 179
VL SMP + +S K+LV + KRI F W KK+ W+ W + + K+ GG+
Sbjct: 622 VLASMPTYTMSCFKLLVSLCKRIQSALTHFWWDSSADKKKMCWIAWSKMAKNKKEGGLGF 681
Query: 180 KDIRVMNISLLAKWRWRLI 198
KDI N +LLAK WR++
Sbjct: 682 KDITNFNDALLAKLSWRIV 700
>At1g23990 reverse transcriptase, putative
Length = 657
Score = 114 bits (285), Expect = 4e-26
Identities = 62/205 (30%), Positives = 110/205 (53%)
Query: 1 VSHLQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLVGINVSEDFMAMAC 60
++HL +ADD + + V+++ + + F SGLKI+ KS++ V+ED
Sbjct: 147 LTHLSFADDLMVLSDGKVRSIDGIVEVFDIFAKFSGLKISMEKSTIYLAGVTEDVYHEIQ 206
Query: 61 DFLNCSAGSITFKYLGLPVGANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVLLNSV 120
+ G + +YLGLP+ + + + PL+E I ++ TW+TRY+S+ GR+ L+ SV
Sbjct: 207 NRYQFDVGQLPVRYLGLPLVTKRLTATDYSPLLEHIKKKIGTWTTRYLSYAGRLNLITSV 266
Query: 121 LNSMPIFYLSFLKMLVGVWKRIVRIQR*FLWGGVGGGKKISWVNWKSVCQQKENGGVRVK 180
L S+ F+L+ ++ + I +I FLW G + + V W VC+ K+ GG+ ++
Sbjct: 267 LWSICNFWLAAFRLPRECIREIDKICSAFLWSGPDLNPRKTRVCWGDVCKPKQEGGLGLR 326
Query: 181 DIRVMNISLLAKWRWRLIDGREALW 205
++ MN K WR++ +LW
Sbjct: 327 SLKEMNEVSCLKLIWRIVSHTNSLW 351
>At2g31520 putative non-LTR retroelement reverse transcriptase
Length = 1524
Score = 113 bits (283), Expect = 6e-26
Identities = 66/210 (31%), Positives = 108/210 (51%), Gaps = 1/210 (0%)
Query: 1 VSHLQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLV-GINVSEDFMAMA 59
++HLQ+ADD+L +A+V+N +K + ++ SG KIN KS + G V +
Sbjct: 832 ITHLQFADDSLFFCQANVRNCQALKDVFDVYEYYSGQKINVQKSMITFGSRVYGSTQSKL 891
Query: 60 CDFLNCSAGSITFKYLGLPVGANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVLLNS 119
L KYLGLP + +E +++ + R +TWS R++S G+ ++L S
Sbjct: 892 KQILEIPNQGGGGKYLGLPEQFGRKKKEMFEYIIDRVKKRTSTWSARFLSPAGKEIMLKS 951
Query: 120 VLNSMPIFYLSFLKMLVGVWKRIVRIQR*FLWGGVGGGKKISWVNWKSVCQQKENGGVRV 179
V +MP++ +S K+ G+ I + F W + I WV WK + K+ GG+
Sbjct: 952 VALAMPVYAMSCFKLPKGIVSEIESLLMNFWWEKASNQRGIPWVAWKRLQYSKKEGGLGF 1011
Query: 180 KDIRVMNISLLAKWRWRLIDGREALWK*VL 209
+D+ N +LLAK WRLI +L+ V+
Sbjct: 1012 RDLAKFNDALLAKQAWRLIQYPNSLFARVM 1041
>At2g25550 putative non-LTR retroelement reverse transcriptase
Length = 1750
Score = 113 bits (283), Expect = 6e-26
Identities = 66/210 (31%), Positives = 108/210 (51%), Gaps = 1/210 (0%)
Query: 1 VSHLQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLV-GINVSEDFMAMA 59
++HLQ+ADD+L +A+V+N +K + ++ SG KIN KS + G V +
Sbjct: 1058 ITHLQFADDSLFFCQANVRNCQALKDVFDVYEYYSGQKINVQKSMITFGSRVYGSTQSRL 1117
Query: 60 CDFLNCSAGSITFKYLGLPVGANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVLLNS 119
L KYLGLP + +E +++ + R +TWS R++S G+ ++L S
Sbjct: 1118 KQILEIPNQGGGGKYLGLPEQFGRKKKEMFEYIIDRVKKRTSTWSARFLSPAGKEIMLKS 1177
Query: 120 VLNSMPIFYLSFLKMLVGVWKRIVRIQR*FLWGGVGGGKKISWVNWKSVCQQKENGGVRV 179
V +MP++ +S K+ G+ I + F W + I WV WK + K+ GG+
Sbjct: 1178 VALAMPVYAMSCFKLPKGIVSEIESLLMNFWWEKASNQRGIPWVAWKRLQYSKKEGGLGF 1237
Query: 180 KDIRVMNISLLAKWRWRLIDGREALWK*VL 209
+D+ N +LLAK WRLI +L+ V+
Sbjct: 1238 RDLAKFNDALLAKQAWRLIQYPNSLFARVM 1267
>At1g51750 hypothetical protein
Length = 629
Score = 113 bits (283), Expect = 6e-26
Identities = 62/205 (30%), Positives = 108/205 (52%)
Query: 1 VSHLQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLVGINVSEDFMAMAC 60
++HL +ADD + + V+++ + ++ F SGL+IN K++L VS+ M
Sbjct: 103 LTHLCFADDLMILTDGKVRSVDGIVEVMNLFAKRSGLQINMEKTTLYTAGVSDHNRYMMI 162
Query: 61 DFLNCSAGSITFKYLGLPVGANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVLLNSV 120
G + +YLGLP+ + PL E I R+ TW++RY+SF GR+ L++SV
Sbjct: 163 SRYPFGLGQLPVRYLGLPLVTKRLTKEDLSPLFEQIRNRIGTWTSRYLSFAGRLNLISSV 222
Query: 121 LNSMPIFYLSFLKMLVGVWKRIVRIQR*FLWGGVGGGKKISWVNWKSVCQQKENGGVRVK 180
L S F++S ++ K I I FLW G ++ + V+W +C+ K+ GG+ ++
Sbjct: 223 LWSTMNFWMSAFRLPSACLKEINSICSAFLWSGPELHRRKAKVSWDDICKPKQEGGLGLR 282
Query: 181 DIRVMNISLLAKWRWRLIDGREALW 205
+ N+ + K WR+ ++LW
Sbjct: 283 SLTEANVVSVLKLIWRVTSNDDSLW 307
>At3g25485 unknown protein
Length = 979
Score = 112 bits (281), Expect = 1e-25
Identities = 59/209 (28%), Positives = 115/209 (54%)
Query: 1 VSHLQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLVGINVSEDFMAMAC 60
++HL +ADD + + V+++ + ++ F SGLKI+ KS++ + +
Sbjct: 614 LTHLTFADDLMVLSDGKVRSVEGIVSVFDEFAKKSGLKISMEKSTIYLAGSANMYRREIE 673
Query: 61 DFLNCSAGSITFKYLGLPVGANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVLLNSV 120
+ + GS+ +YLGLP+ S + + PLVE I ++ TW+ RY+S+ GR+ L++SV
Sbjct: 674 ERFKFAVGSLPVRYLGLPLVTKRFSSADYRPLVEQIKRKIGTWTARYLSYAGRLNLISSV 733
Query: 121 LNSMPIFYLSFLKMLVGVWKRIVRIQR*FLWGGVGGGKKISWVNWKSVCQQKENGGVRVK 180
+ S+ F+++ ++ + I ++ FLW G K + VNW+++C+ K GG+ ++
Sbjct: 734 IWSICNFWIAAFRLPRECIREIDKLCSAFLWSGPELNSKRAKVNWEAICKPKREGGLGLR 793
Query: 181 DIRVMNISLLAKWRWRLIDGREALWK*VL 209
++ N K WR + ++LW+ +L
Sbjct: 794 SLKEANDVSCLKLIWRWVSRGDSLWRKLL 822
>At2g23880 putative non-LTR retroelement reverse transcriptase
Length = 1216
Score = 112 bits (281), Expect = 1e-25
Identities = 61/205 (29%), Positives = 109/205 (52%)
Query: 1 VSHLQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLVGINVSEDFMAMAC 60
++HL +ADD + + ++++ + +L F GLKI K++L VS+ +
Sbjct: 419 LTHLCFADDLMILTDGKIRSVDGIVKVLNQFAAKLGLKICMEKTTLYLAGVSDHSRQLMS 478
Query: 61 DFLNCSAGSITFKYLGLPVGANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVLLNSV 120
+ G + +YLGLP+ + S + PL++ I R+ W++RY+SF GR+ L+NSV
Sbjct: 479 SRYSFGVGKLPVRYLGLPLVTKRLTTSDYSPLIDQIRRRIGMWTSRYLSFAGRLSLINSV 538
Query: 121 LNSMPIFYLSFLKMLVGVWKRIVRIQR*FLWGGVGGGKKISWVNWKSVCQQKENGGVRVK 180
L S+ F+++ ++ I RI LW G K + V+W +C+ K+ GG+ ++
Sbjct: 539 LWSITNFWMNAFRLPRECINEINRISSALLWSGPELNPKKAKVSWDEICKPKKEGGLGLQ 598
Query: 181 DIRVMNISLLAKWRWRLIDGREALW 205
+R N K WRL+ +++LW
Sbjct: 599 SLREANKVSSLKLIWRLLSCQDSLW 623
>At4g10830 putative protein
Length = 1294
Score = 112 bits (280), Expect = 1e-25
Identities = 63/206 (30%), Positives = 109/206 (52%), Gaps = 1/206 (0%)
Query: 1 VSHLQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLV-GINVSEDFMAMA 59
V+HLQ+ADD+L +++V+N +K + ++ SG KIN SKS + G V
Sbjct: 1038 VTHLQFADDSLFFCQSNVRNCQALKDVFDVYEYYSGQKINMSKSMITFGSRVHGTTQNRL 1097
Query: 60 CDFLNCSAGSITFKYLGLPVGANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVLLNS 119
+ L + KYLGLP + + ++E + R ++WS +Y+S G+ ++L S
Sbjct: 1098 KNILGIQSHGGGGKYLGLPEQFGRKKRDMFNYIIERVKKRTSSWSAKYLSPAGKEIMLKS 1157
Query: 120 VLNSMPIFYLSFLKMLVGVWKRIVRIQR*FLWGGVGGGKKISWVNWKSVCQQKENGGVRV 179
V SMP++ +S K+ + + I + F W ++I W+ WK + K+ GG+
Sbjct: 1158 VAMSMPVYAMSCFKLPLNIVSEIEALLMNFWWEKNAKKREIPWIAWKRLQYSKKEGGLGF 1217
Query: 180 KDIRVMNISLLAKWRWRLIDGREALW 205
+D+ N +LLAK WR+I+ +L+
Sbjct: 1218 RDLAKFNDALLAKQVWRMINNPNSLF 1243
>At3g44650 putative protein
Length = 762
Score = 111 bits (278), Expect = 2e-25
Identities = 59/205 (28%), Positives = 105/205 (50%)
Query: 1 VSHLQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLVGINVSEDFMAMAC 60
++HL +ADD + + +++ + + F SGLKI+ KS++ +S A
Sbjct: 251 LTHLSFADDLMILTDGQCRSIEGIIEVFDLFSKWSGLKISMEKSTIFSAGLSSTSRAQLH 310
Query: 61 DFLNCSAGSITFKYLGLPVGANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVLLNSV 120
G + +YLGLP+ S + PL+E I R+ +WS+R++SF GR L++S+
Sbjct: 311 THFPFEVGELPIRYLGLPLVTKRLSSVDYAPLIEQIRKRIGSWSSRFLSFAGRFNLISSI 370
Query: 121 LNSMPIFYLSFLKMLVGVWKRIVRIQR*FLWGGVGGGKKISWVNWKSVCQQKENGGVRVK 180
+ S F+LS ++ + I ++ FLW G K + ++W VC+ K GG+ ++
Sbjct: 371 IWSSCNFWLSAFQLPRACIQEIEKLCSSFLWSGTNLNSKKAKISWNQVCKPKSEGGLGLR 430
Query: 181 DIRVMNISLLAKWRWRLIDGREALW 205
++ N K WR+I ++LW
Sbjct: 431 SLKEANDVCCLKLVWRIISHGDSLW 455
>At2g28980 putative non-LTR retroelement reverse transcriptase
Length = 1529
Score = 110 bits (274), Expect = 7e-25
Identities = 58/205 (28%), Positives = 107/205 (51%)
Query: 1 VSHLQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLVGINVSEDFMAMAC 60
++HL +ADD + ++ + + + F SGL+I+ KS++ VS
Sbjct: 995 LTHLCFADDLMVFVDGHQWSIEGVINVFKEFAGRSGLQISLEKSTIYLAGVSASDRVQTL 1054
Query: 61 DFLNCSAGSITFKYLGLPVGANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVLLNSV 120
+ G + +YLGLP+ + + + PL+E + ++++W+ R +S+ GR+ LLNSV
Sbjct: 1055 SSFPFANGQLPVRYLGLPLLTKQMTTADYSPLIEAVKTKISSWTARSLSYAGRLALLNSV 1114
Query: 121 LNSMPIFYLSFLKMLVGVWKRIVRIQR*FLWGGVGGGKKISWVNWKSVCQQKENGGVRVK 180
+ S+ F++S ++ G + I ++ FLW G K + + W S+CQ K+ GG+ +K
Sbjct: 1115 IVSIANFWMSAYRLPAGCIREIEKLCSAFLWSGPVLNPKKAKIAWSSICQPKKEGGLGIK 1174
Query: 181 DIRVMNISLLAKWRWRLIDGREALW 205
+ N K WRL+ + +LW
Sbjct: 1175 SLAEANKVSCLKLIWRLLSTQPSLW 1199
>At2g05200 putative non-LTR retroelement reverse transcriptase
Length = 1229
Score = 110 bits (274), Expect = 7e-25
Identities = 66/211 (31%), Positives = 106/211 (49%), Gaps = 1/211 (0%)
Query: 1 VSHLQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLV-GINVSEDFMAMA 59
V+HL +ADDT+ K+ ++ + IL + SG INF KSS+
Sbjct: 571 VNHLLFADDTMFFSKSDPESCNKLSEILSRYGKASGQSINFHKSSVTFSSKTPRSVKGQV 630
Query: 60 CDFLNCSAGSITFKYLGLPVGANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVLLNS 119
L T KYLGLP R + +++ I + ++W++R++S G+ V+L +
Sbjct: 631 KRILKIRKEGGTGKYLGLPEHFGRRKRDIFGAIIDKIRQKSHSWASRFLSQAGKQVMLKA 690
Query: 120 VLNSMPIFYLSFLKMLVGVWKRIVRIQR*FLWGGVGGGKKISWVNWKSVCQQKENGGVRV 179
VL SMP++ +S K+ + ++I + F W +K SWV W + K GG+
Sbjct: 691 VLASMPLYSMSCFKLPSALCRKIQSLLTRFWWDTKPDVRKTSWVAWSKLTNPKNAGGLGF 750
Query: 180 KDIRVMNISLLAKWRWRLIDGREALWK*VLV 210
+DI N SLLAK WRL++ E+L +L+
Sbjct: 751 RDIERCNDSLLAKLGWRLLNSPESLLSRILL 781
>At4g02490
Length = 657
Score = 109 bits (273), Expect = 9e-25
Identities = 66/207 (31%), Positives = 107/207 (50%), Gaps = 4/207 (1%)
Query: 1 VSHLQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLV--GINVSEDFMAM 58
++HL +ADD L S+ +L + IL F+ SGL IN K++L+ G N + +
Sbjct: 239 ITHLSFADDILVFCDGSLSSLVAILDILDVFKKGSGLGINLQKTALLLDGGNFERNRIMA 298
Query: 59 ACDFLNCSAGSITFKYLGLPVGANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVLLN 118
A L S GS+ +YLG+P+ + ++PLV+ I R +W+ R++SF GR+ LL
Sbjct: 299 AS--LGVSQGSLPVRYLGVPLMSQKMKKHDYQPLVDRINSRFTSWTARHLSFAGRLQLLK 356
Query: 119 SVLNSMPIFYLSFLKMLVGVWKRIVRIQR*FLWGGVGGGKKISWVNWKSVCQQKENGGVR 178
SV+ S F+ S + ++ ++ FLW G + + ++W VC KE+GG+
Sbjct: 357 SVIYSTINFWASIFILPNQCLHKLEQMCNAFLWSGAPNSAREAKISWDIVCSSKESGGLG 416
Query: 179 VKDIRVMNISLLAKWRWRLIDGREALW 205
+K + N L K W L +LW
Sbjct: 417 LKRLSSWNKVLALKLIWLLFTASGSLW 443
>At1g47850 hypothetical protein
Length = 799
Score = 109 bits (273), Expect = 9e-25
Identities = 59/205 (28%), Positives = 109/205 (52%)
Query: 1 VSHLQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLVGINVSEDFMAMAC 60
++HL +ADD + +++ + I + F SGL I+ KS+L VSE
Sbjct: 271 LTHLCFADDLMVFIDGQQRSVEGVINIFKEFAGKSGLHISLEKSTLYLAGVSELNRNNIL 330
Query: 61 DFLNCSAGSITFKYLGLPVGANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVLLNSV 120
++G + +YLGLP+ + + + PL++ + ++++W+ R +S+ GR+ L+NSV
Sbjct: 331 SAFPFASGQLPVRYLGLPLLTKQMTTADYSPLLDKVRSKISSWTARSLSYAGRLALINSV 390
Query: 121 LNSMPIFYLSFLKMLVGVWKRIVRIQR*FLWGGVGGGKKISWVNWKSVCQQKENGGVRVK 180
+ S+ F++S ++ G K I ++ FLW G K + + W S+C+ K+ GG+ +K
Sbjct: 391 IVSLSNFWMSAYRLPAGCIKEIEKLCSAFLWSGPELNPKKAKITWTSLCKLKQEGGLGIK 450
Query: 181 DIRVMNISLLAKWRWRLIDGREALW 205
+ N K WRL+ + +LW
Sbjct: 451 SLLEANKVSCLKLIWRLVSRQSSLW 475
>At3g45550 putative protein
Length = 851
Score = 108 bits (271), Expect = 2e-24
Identities = 64/210 (30%), Positives = 106/210 (50%), Gaps = 1/210 (0%)
Query: 1 VSHLQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLV-GINVSEDFMAMA 59
++HLQ+ADD+L +A+V+N +K + ++ SG KIN KS + G V
Sbjct: 297 ITHLQFADDSLFFCQANVRNCQALKDVFDVYEYYSGQKINVQKSLITFGSRVYGSTQTRL 356
Query: 60 CDFLNCSAGSITFKYLGLPVGANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVLLNS 119
LN KYLGLP + + +++ + R +WS +++S G+ +LL S
Sbjct: 357 KTLLNIPNQGGGGKYLGLPEQFGRKKKEMFNYIIDRVKERTASWSAKFLSPAGKEILLKS 416
Query: 120 VLNSMPIFYLSFLKMLVGVWKRIVRIQR*FLWGGVGGGKKISWVNWKSVCQQKENGGVRV 179
V +MP++ +S K+ G+ I + F W + I WV WK + K+ GG+
Sbjct: 417 VALAMPVYAMSCFKLPQGIVSEIESLLMNFWWEKASNKRGIPWVAWKRLQYSKKEGGLGF 476
Query: 180 KDIRVMNISLLAKWRWRLIDGREALWK*VL 209
+D+ N +LLAK WR+I +L+ V+
Sbjct: 477 RDLAKFNDALLAKQAWRIIQYPNSLFARVM 506
>At2g14430 putative non-LTR retroelement reverse transcriptase
Length = 1277
Score = 108 bits (270), Expect = 2e-24
Identities = 58/205 (28%), Positives = 108/205 (52%)
Query: 1 VSHLQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLVGINVSEDFMAMAC 60
++HL +ADD + +++ + I + F SGL I+ KS+L VSE
Sbjct: 840 LTHLCFADDLMVFIDGQQRSVEGVINIFKDFAGKSGLHISLEKSTLYLAEVSELNRNNIL 899
Query: 61 DFLNCSAGSITFKYLGLPVGANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVLLNSV 120
++G + +YLG P+ + + + PL++ + ++++W+ R +S+ GR+ L+NSV
Sbjct: 900 SAFPFASGQLPVRYLGFPLLTKQMTTADYSPLLDKVRSKISSWTARSLSYAGRLALINSV 959
Query: 121 LNSMPIFYLSFLKMLVGVWKRIVRIQR*FLWGGVGGGKKISWVNWKSVCQQKENGGVRVK 180
+ S+ F++S ++ G K I ++ FLW G K + + W S+C+ K+ GG+ +K
Sbjct: 960 IVSLSNFWMSAYRLPAGCIKEIEKLCSAFLWSGPELNPKKAKITWTSLCKLKQEGGLGIK 1019
Query: 181 DIRVMNISLLAKWRWRLIDGREALW 205
+ N K WRL+ + +LW
Sbjct: 1020 SLLEANKVSCLKLIWRLVSRQSSLW 1044
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.330 0.142 0.465
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,696,076
Number of Sequences: 26719
Number of extensions: 183426
Number of successful extensions: 553
Number of sequences better than 10.0: 62
Number of HSP's better than 10.0 without gapping: 60
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 441
Number of HSP's gapped (non-prelim): 72
length of query: 211
length of database: 11,318,596
effective HSP length: 95
effective length of query: 116
effective length of database: 8,780,291
effective search space: 1018513756
effective search space used: 1018513756
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 58 (26.9 bits)
Medicago: description of AC148816.5


