

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148655.10 + phase: 0
(250 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
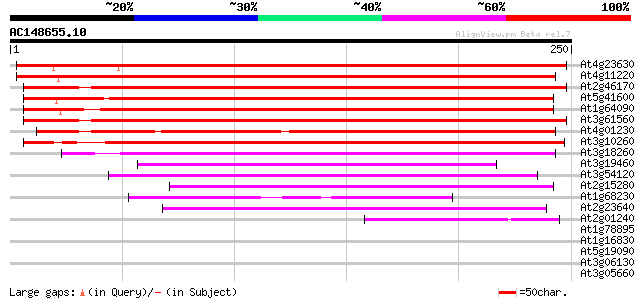
Score E
Sequences producing significant alignments: (bits) Value
At4g23630 unknown protein 299 1e-81
At4g11220 unknown protein 287 4e-78
At2g46170 unknown protein 280 7e-76
At5g41600 unknown protein 276 6e-75
At1g64090 unknown protein 274 3e-74
At3g61560 unknown protein 271 2e-73
At4g01230 predicted protein 200 5e-52
At3g10260 unknown protein 186 1e-47
At3g18260 unknown protein 162 2e-40
At3g19460 unknown protein 120 7e-28
At3g54120 putative protein 118 3e-27
At2g15280 hypothetical protein 117 6e-27
At1g68230 unknown protein 80 1e-15
At2g23640 putative seed maturation protein 72 2e-13
At2g01240 hypothetical protein 47 1e-05
At1g78895 unknown protein 40 0.001
At1g16830 unknown protein 38 0.006
At5g19090 unknown protein 33 0.19
At3g06130 unknown protein 30 1.6
At3g05660 putative disease resistance protein 28 6.0
>At4g23630 unknown protein
Length = 275
Score = 299 bits (765), Expect = 1e-81
Identities = 151/250 (60%), Positives = 189/250 (75%), Gaps = 5/250 (2%)
Query: 4 VDTESFDDKIEDKFH-GSDSDSFSDSEDDHNINNRPPIPLAKSNT----VYRLFGRDRPV 58
V+ ES DKI +K H G DS S S S DD + + P + S++ VYRLFGR++PV
Sbjct: 21 VERESLMDKISEKIHHGGDSSSSSSSSDDEDEKKKTKKPSSPSSSMKSKVYRLFGREQPV 80
Query: 59 HKVLGGGKPADILLWRNKKCTAIALGAGTALWVFFELMQYNLITLVCHLMILALAALFLW 118
HKVLGGGKPADI +W+NKK + LG TA WV FELM+Y+L+TL+CH+MI+ LA LFLW
Sbjct: 81 HKVLGGGKPADIFMWKNKKMSGGVLGGATAAWVVFELMEYHLLTLLCHVMIVVLAVLFLW 140
Query: 119 SNASVFIHKSPLQIPHVVIPQECVFEAASVLRIEINQVFAILREIGTGRDIKKFLTVIAG 178
SNA++FI+KSP +IP V IP+E + + AS LRIEIN+ F+ LREI +GRD+KKFL IAG
Sbjct: 141 SNATMFINKSPPKIPEVHIPEEPILQLASGLRIEINRGFSSLREIASGRDLKKFLIAIAG 200
Query: 179 LWFISVIGSCFNFLTLFYIFYVSLFTLPLVYEKNEDQVDALAEKAMIEIKKQYAVLDAKV 238
LW +S++G CFNFLTL YI V LFT+PL Y+K ED+VD L EKAMIE+KKQYAVLD KV
Sbjct: 201 LWVLSILGGCFNFLTLAYIALVLLFTVPLAYDKYEDKVDPLGEKAMIELKKQYAVLDEKV 260
Query: 239 LSQIPIAGFK 248
LS+IP+ K
Sbjct: 261 LSKIPLGPLK 270
>At4g11220 unknown protein
Length = 271
Score = 287 bits (734), Expect = 4e-78
Identities = 141/241 (58%), Positives = 183/241 (75%), Gaps = 1/241 (0%)
Query: 4 VDTESFDDKIEDKFHGS-DSDSFSDSEDDHNINNRPPIPLAKSNTVYRLFGRDRPVHKVL 62
V+ ES +K+ +K H DS S S S+D++ + P + + VYRLFGR+RPVHKVL
Sbjct: 21 VERESLMEKLSEKIHHKGDSSSSSSSDDENEKKSSSSSPKSLKSKVYRLFGRERPVHKVL 80
Query: 63 GGGKPADILLWRNKKCTAIALGAGTALWVFFELMQYNLITLVCHLMILALAALFLWSNAS 122
GGGKPADI +W++KK + G T WV FELM+Y+L+TL+CH+MI+ALA LFLWSNA+
Sbjct: 81 GGGKPADIFMWKDKKMSGGVFGGATVAWVLFELMEYHLLTLLCHVMIVALAVLFLWSNAT 140
Query: 123 VFIHKSPLQIPHVVIPQECVFEAASVLRIEINQVFAILREIGTGRDIKKFLTVIAGLWFI 182
+FIHKSP +IP V IP+E + + AS LRIEIN+ + LREI +GRDIKKFL+ IAGLW +
Sbjct: 141 MFIHKSPPKIPEVHIPEEPLLQLASGLRIEINRGISSLREIASGRDIKKFLSAIAGLWVL 200
Query: 183 SVIGSCFNFLTLFYIFYVSLFTLPLVYEKNEDQVDALAEKAMIEIKKQYAVLDAKVLSQI 242
S++G C++FLTL YI V LFT+PL Y+K ED+VD+ EKAM E+KKQYAVLDAKV S+I
Sbjct: 201 SILGGCYSFLTLAYIALVLLFTVPLFYDKYEDKVDSYGEKAMAELKKQYAVLDAKVFSKI 260
Query: 243 P 243
P
Sbjct: 261 P 261
>At2g46170 unknown protein
Length = 255
Score = 280 bits (715), Expect = 7e-76
Identities = 137/242 (56%), Positives = 176/242 (72%), Gaps = 5/242 (2%)
Query: 7 ESFDDKIEDKFHGSDSDSFSDSEDDHNINNRPPIPLAKSNTVYRLFGRDRPVHKVLGGGK 66
ES +KI +K H DS S S+SE + +P P A +YR+FGR++PVHKVLGGGK
Sbjct: 13 ESLMEKISEKIHHHDSSSSSESEYE-----KPDSPSAVKAKIYRMFGREKPVHKVLGGGK 67
Query: 67 PADILLWRNKKCTAIALGAGTALWVFFELMQYNLITLVCHLMILALAALFLWSNASVFIH 126
PAD+ LWR+KK + LG TA+WV FEL++Y+L++L+CH+ ILAL LFLWSNA I+
Sbjct: 68 PADVFLWRDKKLSGAVLGVATAIWVLFELVEYHLLSLLCHISILALGGLFLWSNAHTLIN 127
Query: 127 KSPLQIPHVVIPQECVFEAASVLRIEINQVFAILREIGTGRDIKKFLTVIAGLWFISVIG 186
K+ QIP + +P+E AS LR E+NQ F ILR I GRD+KKFL V+ GLW ISV+G
Sbjct: 128 KTSPQIPEIHVPEEAFLVVASSLRNELNQAFVILRSIALGRDLKKFLMVVVGLWIISVVG 187
Query: 187 SCFNFLTLFYIFYVSLFTLPLVYEKNEDQVDALAEKAMIEIKKQYAVLDAKVLSQIPIAG 246
+ FNFLTL YI +V L T+P++YEK+ED+VD LAEKAM E++KQY V D KVLS+IPIA
Sbjct: 188 NWFNFLTLVYICFVILHTVPMLYEKHEDKVDPLAEKAMKELQKQYVVFDEKVLSKIPIAS 247
Query: 247 FK 248
K
Sbjct: 248 LK 249
>At5g41600 unknown protein
Length = 257
Score = 276 bits (707), Expect = 6e-75
Identities = 140/237 (59%), Positives = 172/237 (72%), Gaps = 3/237 (1%)
Query: 7 ESFDDKIEDKFHG-SDSDSFSDSEDDHNINNRPPIPLAKSNTVYRLFGRDRPVHKVLGGG 65
ES +KI +K HG DS S SDS+DD + + +YRLFGR++PVHKVLGGG
Sbjct: 9 ESILEKIVEKIHGHGDSSSLSDSDDDKKSTSSSSSSF--KSKIYRLFGREKPVHKVLGGG 66
Query: 66 KPADILLWRNKKCTAIALGAGTALWVFFELMQYNLITLVCHLMILALAALFLWSNASVFI 125
KPADI LWRNKK + LGA TA WV FEL +Y+L+ +CH I ALAALFLWSNA FI
Sbjct: 67 KPADIFLWRNKKVSGGVLGAVTASWVLFELFEYHLLAFLCHFAIFALAALFLWSNACTFI 126
Query: 126 HKSPLQIPHVVIPQECVFEAASVLRIEINQVFAILREIGTGRDIKKFLTVIAGLWFISVI 185
HKS IP V IP++ + + S LRIEIN+ +LR I +G+D+KKF+ VIAGLW +S+I
Sbjct: 127 HKSTPHIPEVHIPEDPILQLVSGLRIEINRGLTLLRNIASGKDVKKFILVIAGLWVLSII 186
Query: 186 GSCFNFLTLFYIFYVSLFTLPLVYEKNEDQVDALAEKAMIEIKKQYAVLDAKVLSQI 242
GSC+NFLTLFY V LFT+P++YEK ED+VDA EKAM EIKKQYAVLD KVL ++
Sbjct: 187 GSCYNFLTLFYTATVLLFTIPVLYEKYEDKVDAYGEKAMREIKKQYAVLDEKVLRKV 243
>At1g64090 unknown protein
Length = 255
Score = 274 bits (701), Expect = 3e-74
Identities = 140/238 (58%), Positives = 172/238 (71%), Gaps = 9/238 (3%)
Query: 7 ESFDDKIEDKFHGSD--SDSFSDSEDDHNINNRPPIPLAKSNTVYRLFGRDRPVHKVLGG 64
ES +KI +K HG D S S SDS+DD N + +YRLFGR++P+HK+ GG
Sbjct: 9 ESIMEKISEKIHGHDDSSSSSSDSDDDKN-------SASLKTKIYRLFGREQPLHKLFGG 61
Query: 65 GKPADILLWRNKKCTAIALGAGTALWVFFELMQYNLITLVCHLMILALAALFLWSNASVF 124
GKPADI LWRNKK + LGA T W+ FEL++YNL+TL H+ ILALA LFLWS+AS F
Sbjct: 62 GKPADIFLWRNKKVSGGVLGAATVSWILFELLEYNLLTLFGHISILALAVLFLWSSASTF 121
Query: 125 IHKSPLQIPHVVIPQECVFEAASVLRIEINQVFAILREIGTGRDIKKFLTVIAGLWFISV 184
IHKSPL IP V IP++ V + AS LRIEIN+ F +LR+I +GRD+KKFL VIAGLW +S
Sbjct: 122 IHKSPLHIPEVHIPEDVVLQLASGLRIEINRGFTVLRDIASGRDLKKFLLVIAGLWVLSK 181
Query: 185 IGSCFNFLTLFYIFYVSLFTLPLVYEKNEDQVDALAEKAMIEIKKQYAVLDAKVLSQI 242
+GS NFLTL YI V LFT+P++YEK ED+VD EKAM EIKKQY D KVLS++
Sbjct: 182 VGSSCNFLTLIYIATVLLFTIPVLYEKYEDKVDDFGEKAMREIKKQYVEFDVKVLSKV 239
>At3g61560 unknown protein
Length = 253
Score = 271 bits (693), Expect = 2e-73
Identities = 134/242 (55%), Positives = 174/242 (71%), Gaps = 5/242 (2%)
Query: 7 ESFDDKIEDKFHGSDSDSFSDSEDDHNINNRPPIPLAKSNTVYRLFGRDRPVHKVLGGGK 66
ES +KI DK H DS S SDSE + +P P A +YRLFGR++PVHKVLGGG
Sbjct: 13 ESLMEKIADKIHDHDSSSSSDSEHE-----KPESPSALKAKIYRLFGREKPVHKVLGGGL 67
Query: 67 PADILLWRNKKCTAIALGAGTALWVFFELMQYNLITLVCHLMILALAALFLWSNASVFIH 126
PAD+ LWR+KK +A LG TA+WV FEL++Y+ ++LVCH++I ALAALFL SNA F++
Sbjct: 68 PADVFLWRDKKLSASVLGVATAIWVLFELVEYHFLSLVCHILIFALAALFLLSNAHAFMN 127
Query: 127 KSPLQIPHVVIPQECVFEAASVLRIEINQVFAILREIGTGRDIKKFLTVIAGLWFISVIG 186
K+P +IP + I +E S LR E+NQ F ILR I GRD+KKFL V+ GLW ISV+G
Sbjct: 128 KTPPKIPEIHIKEEHFLMIVSALRNELNQAFVILRSIALGRDLKKFLMVVFGLWIISVVG 187
Query: 187 SCFNFLTLFYIFYVSLFTLPLVYEKNEDQVDALAEKAMIEIKKQYAVLDAKVLSQIPIAG 246
+ FNFLTL YI +V L T+P++YEK+ED+VD +AEK + E+KK Y V D KVLS++P+A
Sbjct: 188 NWFNFLTLVYICFVVLHTVPMLYEKHEDKVDPVAEKTLKELKKHYMVFDEKVLSKLPVAS 247
Query: 247 FK 248
K
Sbjct: 248 LK 249
>At4g01230 predicted protein
Length = 242
Score = 200 bits (509), Expect = 5e-52
Identities = 104/231 (45%), Positives = 152/231 (65%), Gaps = 10/231 (4%)
Query: 13 IEDKFHGSDSDSFSDSEDDHNINNRPPIPLAKSNTVYRLFGRDRPVHKVLGGGKPADILL 72
+ ++ +G DS + SDS+ + +P P+ + +YR+FGR+RP+H VLGG AD+LL
Sbjct: 21 VPEEINGLDSLTSSDSDSE-----KPDSPVPINAPIYRMFGRERPIHMVLGGA--ADVLL 73
Query: 73 WRNKKCTAIALGAGTALWVFFELMQYNLITLVCHLMILALAALFLWSNASVFIHKSPLQI 132
WR+KK T L A T +W+ F L+T +C IL L FLWSNA ++KSP +
Sbjct: 74 WRDKKVTLGLLSAVTVIWLLFGFGGRRLLTSLCRGSILFLLLSFLWSNA---LNKSPENM 130
Query: 133 PHVVIPQECVFEAASVLRIEINQVFAILREIGTGRDIKKFLTVIAGLWFISVIGSCFNFL 192
+ IP++ + +AAS + E+N FA LR I RDIK F+ + GLW +SVIG+ F+FL
Sbjct: 131 MDIYIPEKPLLQAASAMTFELNCAFATLRSIALERDIKNFVMAVIGLWLVSVIGNWFSFL 190
Query: 193 TLFYIFYVSLFTLPLVYEKNEDQVDALAEKAMIEIKKQYAVLDAKVLSQIP 243
+L YI +V + T+P++YEK ED++D +AEKA+IE+KK Y V +AK LS+IP
Sbjct: 191 SLLYICFVLIHTVPMLYEKYEDEIDPIAEKAVIEMKKHYQVFEAKFLSKIP 241
>At3g10260 unknown protein
Length = 247
Score = 186 bits (472), Expect = 1e-47
Identities = 90/241 (37%), Positives = 147/241 (60%), Gaps = 15/241 (6%)
Query: 7 ESFDDKIEDKFHGSDSDSFSDSEDDHNINNRPPIPLAKSNTVYRLFGRDRPVHKVLGGGK 66
++F + ++ HGS SF + ED + S+ RLF R +P+H VLGGGK
Sbjct: 16 DNFTETVQKNKHGS---SFFEQED------------SVSSRFNRLFDRQKPIHHVLGGGK 60
Query: 67 PADILLWRNKKCTAIALGAGTALWVFFELMQYNLITLVCHLMILALAALFLWSNASVFIH 126
AD+LLWRNKK +A L TA+WV FE + ++ ++LVC+ ++L + A F+WSNAS F++
Sbjct: 61 SADVLLWRNKKISASVLMGATAIWVLFEWINFHFLSLVCYALLLGMIAQFVWSNASGFLN 120
Query: 127 KSPLQIPHVVIPQECVFEAASVLRIEINQVFAILREIGTGRDIKKFLTVIAGLWFISVIG 186
+S ++P +V+P++ E + E+N+ L+++ ++K+FL + GLW +++G
Sbjct: 121 RSQSRVPRLVLPKDFFAEVGVAVGKEVNRGLLFLQDLACKGNLKQFLMAVIGLWVAAMVG 180
Query: 187 SCFNFLTLFYIFYVSLFTLPLVYEKNEDQVDALAEKAMIEIKKQYAVLDAKVLSQIPIAG 246
SC NFLT+ YI +V T+P++YE+ ED+VD + +++ Y LD LS+IP
Sbjct: 181 SCCNFLTVLYIGFVGAHTMPVLYERYEDEVDGFMDSMIMKFHSHYKKLDTGFLSRIPSGK 240
Query: 247 F 247
F
Sbjct: 241 F 241
>At3g18260 unknown protein
Length = 225
Score = 162 bits (410), Expect = 2e-40
Identities = 85/220 (38%), Positives = 123/220 (55%), Gaps = 11/220 (5%)
Query: 24 SFSDSEDDHNINNRPPIPLAKSNTVYRLFGRDRPVHKVLGGGKPADILLWRNKKCTAIAL 83
S SDSED+ I+ +LF R R +H + GGGK ADILLWR K A +
Sbjct: 7 SSSDSEDERTIHKTT-----------KLFTRQRSIHSIFGGGKVADILLWREPKIAATLV 55
Query: 84 GAGTALWVFFELMQYNLITLVCHLMILALAALFLWSNASVFIHKSPLQIPHVVIPQECVF 143
+ LW E+++YN ITL+CH + ++ F+WS AS F++ IP VV+ +
Sbjct: 56 IGVSILWFLMEVVEYNFITLICHASMTSMLFFFIWSTASDFLNWERPLIPEVVLDESSFK 115
Query: 144 EAASVLRIEINQVFAILREIGTGRDIKKFLTVIAGLWFISVIGSCFNFLTLFYIFYVSLF 203
+ A + NQ+ L ++ GRD F L+ +S+IG+ FNF+ L +I +VS+
Sbjct: 116 QLARSFHVRFNQILTKLLDVACGRDPPLFFLTTISLYIVSIIGTYFNFVNLLFIGFVSMQ 175
Query: 204 TLPLVYEKNEDQVDALAEKAMIEIKKQYAVLDAKVLSQIP 243
TLP++YE ED VD A K M ++KK Y +D VLS+IP
Sbjct: 176 TLPVMYEMYEDDVDTAAGKLMRKMKKLYKKVDTNVLSKIP 215
>At3g19460 unknown protein
Length = 200
Score = 120 bits (301), Expect = 7e-28
Identities = 61/160 (38%), Positives = 95/160 (59%)
Query: 58 VHKVLGGGKPADILLWRNKKCTAIALGAGTALWVFFELMQYNLITLVCHLMILALAALFL 117
VH+ LG G AD+LLWRN+ I L + T W FE YNL++ V ++++L +A FL
Sbjct: 13 VHQSLGAGSVADLLLWRNRTGAVILLISSTGFWFLFERAGYNLLSFVSNVLLLLVAIFFL 72
Query: 118 WSNASVFIHKSPLQIPHVVIPQECVFEAASVLRIEINQVFAILREIGTGRDIKKFLTVIA 177
W+ ++ +++ +P++ IP+E +AA LR+ IN V +I +I R+ + L V
Sbjct: 73 WAKSATVLNRPLPPVPNMEIPEEFANKAADDLRVWINYVLSIASDITIARNPIRLLQVSL 132
Query: 178 GLWFISVIGSCFNFLTLFYIFYVSLFTLPLVYEKNEDQVD 217
LW IS +G+ N LTL YI + + P+VYEK +D +D
Sbjct: 133 VLWAISYVGTLINSLTLVYIGVLLSLSFPIVYEKYQDHID 172
>At3g54120 putative protein
Length = 203
Score = 118 bits (296), Expect = 3e-27
Identities = 60/191 (31%), Positives = 106/191 (55%)
Query: 45 SNTVYRLFGRDRPVHKVLGGGKPADILLWRNKKCTAIALGAGTALWVFFELMQYNLITLV 104
S++ +LF RDR +H++LGGG AD++LWR K + + A W+ FE Y + TL+
Sbjct: 2 SSSSDKLFNRDRTIHEILGGGIVADVMLWRKKNVSVGIVTVTIASWMVFEAFAYTIFTLI 61
Query: 105 CHLMILALAALFLWSNASVFIHKSPLQIPHVVIPQECVFEAASVLRIEINQVFAILREIG 164
+++L L+ LFLWS ++ +++ +P I + EA+ LRI +N++ + +I
Sbjct: 62 SSVLLLLLSILFLWSKSASILNRPSPPLPEFQISEAMAEEASIWLRIHVNKLLQVSHDIA 121
Query: 165 TGRDIKKFLTVIAGLWFISVIGSCFNFLTLFYIFYVSLFTLPLVYEKNEDQVDALAEKAM 224
RD + + V L+ +S+IGS +F TL + + + T+P YE+ ED + E
Sbjct: 122 MARDSELYTKVAVSLFLLSLIGSLMDFQTLCHTSVLVVMTVPAFYERYEDYIVGSIEFIC 181
Query: 225 IEIKKQYAVLD 235
+ ++ Y L+
Sbjct: 182 NKTRELYLRLE 192
>At2g15280 hypothetical protein
Length = 183
Score = 117 bits (293), Expect = 6e-27
Identities = 56/171 (32%), Positives = 97/171 (55%)
Query: 72 LWRNKKCTAIALGAGTALWVFFELMQYNLITLVCHLMILALAALFLWSNASVFIHKSPLQ 131
+W+N++ + LG+ T LW FE Y+ V + +L++ LFLW+ +++ ++ Q
Sbjct: 1 MWKNRRGGFLLLGSTTLLWFLFEKCGYSFFPFVVNTQLLSVVILFLWAKSAILFNRPMPQ 60
Query: 132 IPHVVIPQECVFEAASVLRIEINQVFAILREIGTGRDIKKFLTVIAGLWFISVIGSCFNF 191
+P++ I +E VF A +R+ IN V A+ REI GR+ K+ V LW +S +G+ NF
Sbjct: 61 LPNLEITEEFVFMVADAIRVWINTVLAVAREIYVGRNAKQLFRVSVVLWTVSFVGNFLNF 120
Query: 192 LTLFYIFYVSLFTLPLVYEKNEDQVDALAEKAMIEIKKQYAVLDAKVLSQI 242
LT+ Y+ V +P +YE+ +D +D I+ QY +D ++L +I
Sbjct: 121 LTILYLGVVLSLLIPFLYERYQDLIDEKLSLTHRVIQTQYRKIDERLLQKI 171
>At1g68230 unknown protein
Length = 151
Score = 80.1 bits (196), Expect = 1e-15
Identities = 46/144 (31%), Positives = 75/144 (51%), Gaps = 13/144 (9%)
Query: 54 RDRPVHKVLGGGKPADILLWRNKKCTAIALGAGTALWVFFELMQYNLITLVCHLMILALA 113
R R ++ LGGG+ ADI+ W+NKK + LG T +W FE+++Y IT +C +++L +
Sbjct: 18 RRRSLYHNLGGGRFADIMFWKNKKESGTILGVFTLIWFLFEVVEYPFITFLCQILLLFI- 76
Query: 114 ALFLWSNASVFIHKSPLQIPHVVIPQECVFEAASVLRIEINQVFAILREIGTGRDIKKFL 173
+F+ P I + I + L +IN L +I +G+D +
Sbjct: 77 --------FIFLICKPPSINDLRISE----SNWRFLFNKINWFIIKLYDISSGKDFRLLF 124
Query: 174 TVIAGLWFISVIGSCFNFLTLFYI 197
+ LW +SV+G+ F+ LTL YI
Sbjct: 125 LAVVSLWILSVVGNYFSSLTLLYI 148
>At2g23640 putative seed maturation protein
Length = 206
Score = 72.4 bits (176), Expect = 2e-13
Identities = 42/171 (24%), Positives = 76/171 (43%)
Query: 69 DILLWRNKKCTAIALGAGTALWVFFELMQYNLITLVCHLMILALAALFLWSNASVFIHKS 128
DI LWR KK L T+ W+ + IT+V + I ++ +FLW + + K
Sbjct: 18 DIYLWRRKKLAFSTLLVSTSTWILLSFYGFTTITIVSWIGIAVVSMIFLWGSLLRLLSKV 77
Query: 129 PLQIPHVVIPQECVFEAASVLRIEINQVFAILREIGTGRDIKKFLTVIAGLWFISVIGSC 188
++ + + +E V E R+ + ++ + +G + F + G W +S IG+
Sbjct: 78 EPELSGLEVSEEFVVETVRSCRMLMEEMVRWMFRVGAESEWFVFARTVLGFWILSRIGNL 137
Query: 189 FNFLTLFYIFYVSLFTLPLVYEKNEDQVDALAEKAMIEIKKQYAVLDAKVL 239
+F T +I V T+P ++E+ DQ+ + K Y K+L
Sbjct: 138 LDFHTCLFIGLVMGLTVPKLWEEYGDQIQKHLGSLKDKSKGAYNTTHEKIL 188
>At2g01240 hypothetical protein
Length = 160
Score = 47.0 bits (110), Expect = 1e-05
Identities = 31/87 (35%), Positives = 48/87 (54%), Gaps = 1/87 (1%)
Query: 159 ILREIGTGRDIKKFLTVIAGLWFISVIGSCFNFLTLFYIFYVSLFTLPLVYEKNEDQVDA 218
+L EI G+D K FL I + I IGS + LT+ YI V T+P+VY + ++ +D+
Sbjct: 71 MLYEIAYGKDNKTFLKTILYVAIIYNIGSYISLLTILYICLVCSMTIPVVYMQFQELIDS 130
Query: 219 LAEKAMIEIKKQYAVLDAKVLSQIPIA 245
K + E K + +V+S+IP A
Sbjct: 131 FMGK-VSEEKNNLLEVFKQVVSKIPRA 156
>At1g78895 unknown protein
Length = 164
Score = 40.4 bits (93), Expect = 0.001
Identities = 36/135 (26%), Positives = 63/135 (46%), Gaps = 8/135 (5%)
Query: 101 ITLVCHLMILALAALFLWSNASVFIHKSPLQIPHVVIPQECVFEAASVLRIEINQVFAIL 160
I L+C L IL L LF N SV + QI Q+ + L + +L
Sbjct: 37 IVLLCSLAILGL--LFRQLNVSVPVDPLEWQIS-----QDTASNIVARLANTVGAAEGVL 89
Query: 161 REIGTGRDIKKFLTVIAGLWFISVIGSCFNFLTLFYIFYVSLFTLPLVYEKNEDQVDALA 220
R TG D + F+ V+ L+F+S +G + +T+ Y + LF L ++ + ++ + +
Sbjct: 90 RVAATGHDKRLFVKVVICLYFLSALGRLISGVTVAYA-GLCLFCLSMLCQTSQSLGNCVL 148
Query: 221 EKAMIEIKKQYAVLD 235
++ +I +Q A D
Sbjct: 149 KRGNGQILEQEAHSD 163
>At1g16830 unknown protein
Length = 152
Score = 37.7 bits (86), Expect = 0.006
Identities = 29/132 (21%), Positives = 65/132 (48%), Gaps = 8/132 (6%)
Query: 85 AGTALWVFFELMQYNLITLVCHLMILALAAL----FLWSNASVFIHKSPLQIPHVVIPQE 140
+GT ++ L++L+ ++I+ L++L L+ + +V + PL+ I Q+
Sbjct: 10 SGTLVYHHCANRNATLLSLISDVLIVLLSSLAILGLLFRHLNVSVPVDPLEWQ---ISQD 66
Query: 141 CVFEAASVLRIEINQVFAILREIGTGRDIKKFLTVIAGLWFISVIGSCFNFLTLFYIFYV 200
+ L + ++LR TG D + F+ V+ L+F++ +G + +T+ Y +
Sbjct: 67 TACNIVARLANTVGAAESVLRVAATGHDKRLFVKVVICLYFLAALGRIISGVTIAYA-GL 125
Query: 201 SLFTLPLVYEKN 212
LF L +++ +
Sbjct: 126 CLFCLSMLFRSS 137
>At5g19090 unknown protein
Length = 587
Score = 32.7 bits (73), Expect = 0.19
Identities = 14/37 (37%), Positives = 20/37 (53%), Gaps = 1/37 (2%)
Query: 5 DTESFDDKIEDKFHGSDSDSFSDSEDDHNINNRPPIP 41
D E F D+ +D+F D D F D +D ++ PP P
Sbjct: 219 DEEDFSDEFDDEF-DEDDDEFDDDLEDDEFDDHPPPP 254
>At3g06130 unknown protein
Length = 473
Score = 29.6 bits (65), Expect = 1.6
Identities = 11/27 (40%), Positives = 15/27 (54%)
Query: 5 DTESFDDKIEDKFHGSDSDSFSDSEDD 31
D + F D+ +D+F D D F D DD
Sbjct: 191 DDDDFSDEFDDEFTDDDDDEFDDEFDD 217
>At3g05660 putative disease resistance protein
Length = 883
Score = 27.7 bits (60), Expect = 6.0
Identities = 18/49 (36%), Positives = 29/49 (58%), Gaps = 1/49 (2%)
Query: 182 ISVIGSCFNFLTLFYIFYVSLFTLPLVYEKNEDQVDALAE-KAMIEIKK 229
+S+I F FL LF+ + +F +P ++ + +Q DAL E K +IKK
Sbjct: 1 MSLIPITFYFLFLFFSNFRGVFAVPNIHLCHFEQRDALLEFKNEFKIKK 49
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.325 0.141 0.424
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,550,540
Number of Sequences: 26719
Number of extensions: 234972
Number of successful extensions: 1066
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 19
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 1030
Number of HSP's gapped (non-prelim): 27
length of query: 250
length of database: 11,318,596
effective HSP length: 97
effective length of query: 153
effective length of database: 8,726,853
effective search space: 1335208509
effective search space used: 1335208509
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 59 (27.3 bits)
Medicago: description of AC148655.10


