

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148652.15 + phase: 0 /pseudo
(121 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
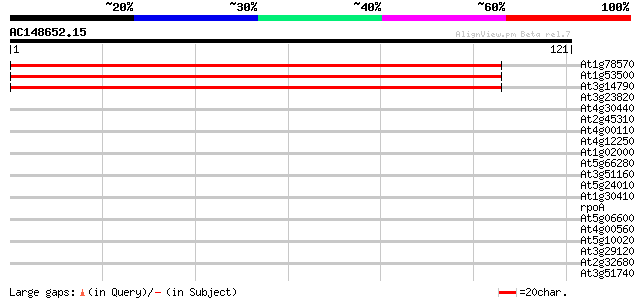
Score E
Sequences producing significant alignments: (bits) Value
At1g78570 Similar to dTDP-D-glucose 4,6-dehydratase 147 6e-37
At1g53500 dTDP-D-glucose 4,6-dehydratase, putative 147 8e-37
At3g14790 dTDP-glucose 4,6-dehydratase, putative 146 1e-36
At3g23820 NAD dependent epimerase, putative 37 0.002
At4g30440 nucleotide sugar epimerase-like protein 35 0.005
At2g45310 putative nucleotide sugar epimerase 33 0.023
At4g00110 nucleotide sugar epimerase like protein 33 0.029
At4g12250 nucleotide sugar epimerase -like protein 32 0.038
At1g02000 nucleotide sugar epimerase, putative 32 0.038
At5g66280 GDP-D-mannose 4,6-dehydratase; GMD1 (gb|AAF07199.1) 31 0.086
At3g51160 GDP-D-mannose-4,6-dehydratase (MUR1) 31 0.086
At5g24010 receptor-protein kinase-like protein 29 0.33
At1g30410 hypothetical protein 28 0.95
rpoA -chloroplast genome- RNA polymerase alpha subunit 27 1.2
At5g06600 ubiquitin carboxyl-terminal hydrolase 27 1.6
At4g00560 putative dTDP-6-deoxy-L-mannose-dehydrogenase 27 1.6
At5g10020 receptor protein kinase -like(fragment) 27 2.1
At3g29120 hypothetical protein 27 2.1
At2g32680 putative disease resistance protein 27 2.1
At3g51740 unknown protein 26 2.8
>At1g78570 Similar to dTDP-D-glucose 4,6-dehydratase
Length = 669
Score = 147 bits (372), Expect = 6e-37
Identities = 73/106 (68%), Positives = 83/106 (77%)
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKGFIHVSTDKFYGETDENAV 60
MHFAAQTHVDNS+ NSFEFT+ + Q++ FIHVSTD+ YGETDE+A+
Sbjct: 85 MHFAAQTHVDNSFGNSFEFTKNNIYGTHVLLEACKVTGQIRRFIHVSTDEVYGETDEDAL 144
Query: 61 VGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRVY 106
VGNHEASQLL TNPYSA K GAEMLVM+YGRSYGLPVITTRG+ VY
Sbjct: 145 VGNHEASQLLPTNPYSATKAGAEMLVMAYGRSYGLPVITTRGNNVY 190
>At1g53500 dTDP-D-glucose 4,6-dehydratase, putative
Length = 667
Score = 147 bits (371), Expect = 8e-37
Identities = 73/106 (68%), Positives = 82/106 (76%)
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKGFIHVSTDKFYGETDENAV 60
MHFAAQTHVDNS+ NSFEFT+ + Q++ FIHVSTD+ YGETDE+A
Sbjct: 87 MHFAAQTHVDNSFGNSFEFTKNNIYGTHVLLEACKVTGQIRRFIHVSTDEVYGETDEDAA 146
Query: 61 VGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRVY 106
VGNHEASQLL TNPYSA K GAEMLVM+YGRSYGLPVITTRG+ VY
Sbjct: 147 VGNHEASQLLPTNPYSATKAGAEMLVMAYGRSYGLPVITTRGNNVY 192
>At3g14790 dTDP-glucose 4,6-dehydratase, putative
Length = 664
Score = 146 bits (369), Expect = 1e-36
Identities = 73/106 (68%), Positives = 82/106 (76%)
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKGFIHVSTDKFYGETDENAV 60
MHFAAQTHVDNS+ NSFEFT+ + Q++ FIHVSTD+ YGETDE+A
Sbjct: 85 MHFAAQTHVDNSFGNSFEFTKNNIYGTHVLLEACKVTGQIRRFIHVSTDEVYGETDEDAS 144
Query: 61 VGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRVY 106
VGNHEASQLL TNPYSA K GAEMLVM+YGRSYGLPVITTRG+ VY
Sbjct: 145 VGNHEASQLLPTNPYSATKAGAEMLVMAYGRSYGLPVITTRGNNVY 190
>At3g23820 NAD dependent epimerase, putative
Length = 460
Score = 37.0 bits (84), Expect = 0.002
Identities = 25/106 (23%), Positives = 42/106 (39%), Gaps = 2/106 (1%)
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKGFIHVSTDKFYGETDENAV 60
+H AAQ V + +N + ++ ++ + + N + S+ YG EN
Sbjct: 191 LHLAAQAGVRYAMKNPQSYIASNIAGFVNLLEVAKAANPQPAIVWASSSSVYGLNTENPF 250
Query: 61 VGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRVY 106
H Q + Y+A K E + +Y YGL + R VY
Sbjct: 251 SEEHRTDQ--PASLYAATKKAGEEIAHTYNHIYGLSLTGLRFFTVY 294
>At4g30440 nucleotide sugar epimerase-like protein
Length = 429
Score = 35.4 bits (80), Expect = 0.005
Identities = 25/106 (23%), Positives = 43/106 (39%), Gaps = 2/106 (1%)
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKGFIHVSTDKFYGETDENAV 60
MH AAQ V + EN + ++ L++ + + N + S+ YG ++
Sbjct: 167 MHLAAQAGVRYALENPQSYVHSNIAGLVNLLEICKAANPQPAIVWASSSSVYGLNEKVPF 226
Query: 61 VGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRVY 106
+ Q + Y+A K E + +Y YGL + R VY
Sbjct: 227 SESDRTDQ--PASLYAATKKAGEEITHTYNHIYGLAITGLRFFTVY 270
>At2g45310 putative nucleotide sugar epimerase
Length = 437
Score = 33.1 bits (74), Expect = 0.023
Identities = 25/106 (23%), Positives = 41/106 (38%), Gaps = 2/106 (1%)
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKGFIHVSTDKFYGETDENAV 60
MH AAQ V + EN + ++ ++ + S N + S+ YG +
Sbjct: 176 MHLAAQAGVRYAMENPSSYVHSNIAGFVNLLEICKSVNPQPAIVWASSSSVYGLNTKVPF 235
Query: 61 VGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRVY 106
+ Q + Y+A K E + +Y YGL + R VY
Sbjct: 236 SEKDKTDQ--PASLYAATKKAGEEIAHTYNHIYGLSLTGLRFFTVY 279
>At4g00110 nucleotide sugar epimerase like protein
Length = 430
Score = 32.7 bits (73), Expect = 0.029
Identities = 25/106 (23%), Positives = 40/106 (37%), Gaps = 2/106 (1%)
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKGFIHVSTDKFYGETDENAV 60
MH AAQ V + EN + ++ ++ + S N + S+ YG +
Sbjct: 170 MHLAAQAGVRYAMENPSSYVHSNIAGFVNLLEVCKSANPQPAIVWASSSSVYGLNTKVPF 229
Query: 61 VGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRVY 106
Q + Y+A K E + +Y YGL + R VY
Sbjct: 230 SEKDRTDQ--PASLYAATKKAGEEIAHTYNHIYGLSLTGLRFFTVY 273
>At4g12250 nucleotide sugar epimerase -like protein
Length = 436
Score = 32.3 bits (72), Expect = 0.038
Identities = 24/106 (22%), Positives = 40/106 (37%), Gaps = 2/106 (1%)
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKGFIHVSTDKFYGETDENAV 60
MH AAQ V + +N + ++ ++ + S N + S+ YG +
Sbjct: 175 MHLAAQAGVRYAMQNPGSYVNSNIAGFVNLLEVSKSANPQPAIVWASSSSVYGLNSKVPF 234
Query: 61 VGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRVY 106
Q + Y+A K E + +Y YGL + R VY
Sbjct: 235 SEKDRTDQ--PASLYAATKKAGEGIAHTYNHIYGLSLTGLRFFTVY 278
>At1g02000 nucleotide sugar epimerase, putative
Length = 434
Score = 32.3 bits (72), Expect = 0.038
Identities = 25/106 (23%), Positives = 40/106 (37%), Gaps = 2/106 (1%)
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKGFIHVSTDKFYGETDENAV 60
MH AAQ V + EN + ++ ++ + S N + S+ YG +
Sbjct: 171 MHLAAQAGVRYAMENPGSYVHSNIAGFVNLLEVCKSANPQPAIVWASSSSVYGLNTKVPF 230
Query: 61 VGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRVY 106
Q + Y+A K E + +Y YGL + R VY
Sbjct: 231 SEKDRTDQ--PASLYAATKKAGEEIAHTYNHIYGLSLTGLRFFTVY 274
>At5g66280 GDP-D-mannose 4,6-dehydratase; GMD1 (gb|AAF07199.1)
Length = 361
Score = 31.2 bits (69), Expect = 0.086
Identities = 23/98 (23%), Positives = 43/98 (43%), Gaps = 8/98 (8%)
Query: 2 HFAAQTHVDNSYE---NSFEFTQTSFMELM-SFRSLQVSKNQVKGFIHVSTDKFYGETDE 57
+ AAQ+HV S+E + + T + L+ + RS + + + + + +G T
Sbjct: 100 NLAAQSHVAVSFEIPDYTADVVATGALRLLEAVRSHNIDNGRAIKYYQAGSSEMFGSTPP 159
Query: 58 NAVVGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGL 95
E + +PY+A K A ++Y +YGL
Sbjct: 160 P----QSETTPFHPRSPYAASKCAAHWYTVNYREAYGL 193
>At3g51160 GDP-D-mannose-4,6-dehydratase (MUR1)
Length = 373
Score = 31.2 bits (69), Expect = 0.086
Identities = 23/98 (23%), Positives = 44/98 (44%), Gaps = 8/98 (8%)
Query: 2 HFAAQTHVDNSYE---NSFEFTQTSFMELM-SFRSLQVSKNQVKGFIHVSTDKFYGETDE 57
+ AAQ+HV S+E + + T + L+ + RS + + + + + +G T
Sbjct: 112 NLAAQSHVAVSFEIPDYTADVVATGALRLLEAVRSHTIDSGRTVKYYQAGSSEMFGSTPP 171
Query: 58 NAVVGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGL 95
E + + +PY+A K A ++Y +YGL
Sbjct: 172 P----QSETTPIHLRSPYAASKCAAHWYTVNYREAYGL 205
>At5g24010 receptor-protein kinase-like protein
Length = 824
Score = 29.3 bits (64), Expect = 0.33
Identities = 21/58 (36%), Positives = 27/58 (46%), Gaps = 4/58 (6%)
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKGFIHVSTDKFYGETDEN 58
+ FAA T DN NS T TSF SF +S + G +STD+ +D N
Sbjct: 21 LSFAAFTPTDNYLINSGSNTNTSFFTTRSF----LSDSSEPGSSFLSTDRSISISDTN 74
>At1g30410 hypothetical protein
Length = 1495
Score = 27.7 bits (60), Expect = 0.95
Identities = 29/103 (28%), Positives = 47/103 (45%), Gaps = 7/103 (6%)
Query: 21 QTSFMELMSFRSLQVSKNQVKG---FIHVSTDKF-YGETDENAVVGNHEASQLLETNPYS 76
Q F+ LM + + VS+ +K F+ +S + + ENA G +A+Q + TN +
Sbjct: 799 QLHFLPLMD-KIILVSEGMIKEEGTFVELSKSGILFKKLMENA--GKMDATQEVNTNDEN 855
Query: 77 AMKIGAEMLVMSYGRSYGLPVITTRGDRVYPFSHERKESIDSW 119
+K+G + V R+ G R V ER+ I SW
Sbjct: 856 ILKLGPTVTVDVSERNLGSTKQGKRRRSVLIKQEERETGIISW 898
>rpoA -chloroplast genome- RNA polymerase alpha subunit
Length = 329
Score = 27.3 bits (59), Expect = 1.2
Identities = 20/63 (31%), Positives = 31/63 (48%), Gaps = 10/63 (15%)
Query: 8 HVDNSYENSFEFTQTSFMELMSFRSL-------QVSKNQVKGFI---HVSTDKFYGETDE 57
H +SY N E + F+E+ + SL Q S+N + FI HV + FY E ++
Sbjct: 184 HSIHSYGNGNEKQEILFLEIWTNGSLTPKEALHQASRNLINLFIPFLHVEEETFYLENNQ 243
Query: 58 NAV 60
+ V
Sbjct: 244 HQV 246
>At5g06600 ubiquitin carboxyl-terminal hydrolase
Length = 1126
Score = 26.9 bits (58), Expect = 1.6
Identities = 14/50 (28%), Positives = 27/50 (54%), Gaps = 3/50 (6%)
Query: 2 HFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKGFIHVSTDKF 51
HFA +T + +N F + F+ + +L+ KN+++ +HVS + F
Sbjct: 1011 HFAKETGQNQQVQN---FGEPFFLVIHEGETLEEIKNRIQKKLHVSDEDF 1057
>At4g00560 putative dTDP-6-deoxy-L-mannose-dehydrogenase
Length = 267
Score = 26.9 bits (58), Expect = 1.6
Identities = 18/59 (30%), Positives = 26/59 (43%), Gaps = 18/59 (30%)
Query: 44 IHVSTDK-------FYGETDENAVVGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGL 95
IH+STD+ FY E DE V N Y K+ AE+L+ +S+ +
Sbjct: 126 IHLSTDQVYQGVKSFYKEEDETVAV-----------NVYGKSKVAAELLIKDKCQSFAI 173
>At5g10020 receptor protein kinase -like(fragment)
Length = 1048
Score = 26.6 bits (57), Expect = 2.1
Identities = 22/72 (30%), Positives = 30/72 (41%), Gaps = 15/72 (20%)
Query: 9 VDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKGFIHVSTDKFYGETDENAVVGNHEASQ 68
+D S F SF S RSL +S+N ++G I F G AS+
Sbjct: 416 IDLSSNKFSGFIPVSFFTFASLRSLNLSRNNLEGPI-----PFRGS----------RASE 460
Query: 69 LLETNPYSAMKI 80
LL N Y M++
Sbjct: 461 LLVLNSYPQMEL 472
>At3g29120 hypothetical protein
Length = 154
Score = 26.6 bits (57), Expect = 2.1
Identities = 10/31 (32%), Positives = 19/31 (61%)
Query: 31 RSLQVSKNQVKGFIHVSTDKFYGETDENAVV 61
++L ++N +GF V TD+ G+ D N ++
Sbjct: 123 QALLCTRNWFRGFQEVETDEIQGQEDTNTIL 153
>At2g32680 putative disease resistance protein
Length = 890
Score = 26.6 bits (57), Expect = 2.1
Identities = 16/63 (25%), Positives = 26/63 (40%)
Query: 9 VDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKGFIHVSTDKFYGETDENAVVGNHEASQ 68
+D SY F +S + L L + +N + G + VS + + NH Q
Sbjct: 272 LDLSYNKFFGVIPSSLLTLPFLAHLALRENNLAGSVEVSNSSTSSRLEIMYLGSNHFEGQ 331
Query: 69 LLE 71
+LE
Sbjct: 332 ILE 334
>At3g51740 unknown protein
Length = 836
Score = 26.2 bits (56), Expect = 2.8
Identities = 16/50 (32%), Positives = 24/50 (48%)
Query: 9 VDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKGFIHVSTDKFYGETDEN 58
+D SY + SF L S SL + N +KG I + D+ + T+ N
Sbjct: 292 LDFSYNSINGTIPDSFSNLSSLVSLNLESNHLKGPIPDAIDRLHNLTELN 341
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.317 0.130 0.380
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,497,401
Number of Sequences: 26719
Number of extensions: 82090
Number of successful extensions: 200
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 19
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 172
Number of HSP's gapped (non-prelim): 29
length of query: 121
length of database: 11,318,596
effective HSP length: 97
effective length of query: 24
effective length of database: 8,726,853
effective search space: 209444472
effective search space used: 209444472
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 52 (24.6 bits)
Medicago: description of AC148652.15


