

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148651.5 + phase: 0
(496 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
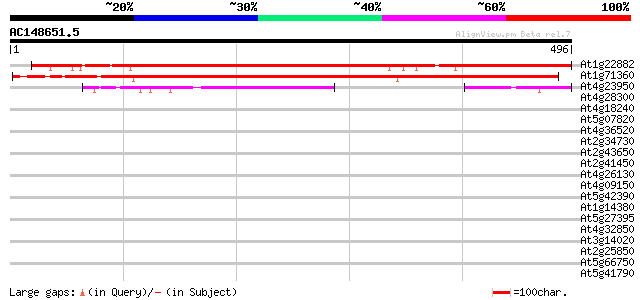
Score E
Sequences producing significant alignments: (bits) Value
At1g22882 unknown protein 395 e-110
At1g71360 hypothetical protein 386 e-107
At4g23950 putative protein 166 2e-41
At4g28300 predicted proline-rich protein 35 0.095
At4g18240 starch synthase-like protein 34 0.16
At5g07820 putative protein 33 0.28
At4g36520 trichohyalin like protein 32 1.1
At2g34730 putative myosin heavy chain 31 1.4
At2g43650 unknown protein 31 1.8
At2g41450 hypothetical protein 31 1.8
At4g26130 uncharacterized protein 30 2.3
At4g09150 unknown protein 30 2.3
At5g42390 pitrilysin 30 3.1
At1g14380 unknown protein 30 3.1
At5g27395 putative protein 30 4.0
At4g32850 poly(A) polymerase like protein 30 4.0
At3g14020 putative transcription factor 30 4.0
At2g25850 nuclear poly(A) polymerase 30 4.0
At5g66750 SWI2/SNF2-like protein (gb|AAD28303.1) 29 5.2
At5g41790 myosin heavy chain-like protein 29 5.2
>At1g22882 unknown protein
Length = 660
Score = 395 bits (1015), Expect = e-110
Identities = 236/519 (45%), Positives = 328/519 (62%), Gaps = 48/519 (9%)
Query: 20 VSLLYLYVDGSEELS----VGLSKWNEVNHGFCEISDTA--------DKYFIKE------ 61
+SL VD + +LS V LS+ +E EIS T D Y +K+
Sbjct: 80 LSLASASVDVTSDLSRNDDVNLSEESEDKEQEAEISSTVSGNDIESKDTYLLKQSEINKK 139
Query: 62 ---IDACFPSEALIYSKAGDAEANGLVNESHNGRESGAYAVPADINK-------ENTDSA 111
IDA S+ + K + G N++ G++ + + +NK E++D+
Sbjct: 140 DTGIDA--GSKYDDFPKKSEINNTGTWNDTE-GKDDNNFLKQSQLNKTGTGNDTESSDNE 196
Query: 112 NREDHVVENSEYAVKHENDVKKSDILSRAVPLGLNEFKSRAISSKVKSGTGQSRSVIHRL 171
E + + + E +V K D SRAVPLGL+EFKSRA +S+ KS + Q VIHR+
Sbjct: 197 FLEQNQMNKTVLGNGTEINVSKVDQPSRAVPLGLDEFKSRASNSRNKSLSDQVSGVIHRM 256
Query: 172 EPGGAEYNYASASKGAKVLGSNKEGKGASNILSRDKDKYLRNPCSVVGKFVIMELSEETL 231
EPGG EYNYASASKGAKVL SNKE KGA++ILSRD DKYLRNPCS GKFV++ELSEETL
Sbjct: 257 EPGGKEYNYASASKGAKVLSSNKEAKGAASILSRDNDKYLRNPCSTEGKFVVVELSEETL 316
Query: 232 VDTIEIANFEHHSSNLKDFEIHGSLNFPTNVWDLLGNFTASNVRHAQRFVLKEPKWVRYL 291
V+TI+IANFEH+SSNLK+FE+ G+L +PT+ W +GNFTASNV+H Q F L EPKWVRYL
Sbjct: 317 VNTIKIANFEHYSSNLKEFELQGTLVYPTDTWVHMGNFTASNVKHEQNFTLLEPKWVRYL 376
Query: 292 KLNLQSHYGSEFYCTLSVVEVFGVDAVERMLEDLINTQDNLLA--SGEGNADKTILP--- 346
KLN SHYGSEFYCTLS++EV+GVDAVERMLEDLI+ QDN A EG+++ P
Sbjct: 377 KLNFISHYGSEFYCTLSLIEVYGVDAVERMLEDLISVQDNKNAYKPREGDSEHKEKPMQQ 436
Query: 347 ----HPDPAVIEHVHK---KPLEGINSVPASDISSSKHETANIKVPDPVEEIR--QQVGR 397
D + H+ K N + ++ S +K ++ K+ +PVEE+R Q R
Sbjct: 437 IESLEGDDGADKSTHREKEKEAPPENMLAKTEASMAK---SSNKLSEPVEEMRHHQPGSR 493
Query: 398 MPGDTVLKILMQKVRTLDVNLFVLERYMEDLNSRYVNIFKDYSKDTGEKDIVLQKIKEDI 457
MPGDTVLKILMQK+R+LD+NL +LERY+E+LN RY NIFK+ ++ G ++ + ++ D+
Sbjct: 494 MPGDTVLKILMQKLRSLDLNLSILERYLEELNLRYGNIFKEMDREAGVREKAIVALRLDL 553
Query: 458 KNLIDHQDVIAKDASDLISWKSQVSSQLNHLIQDNAVLR 496
+ + + Q+ + +A ++ W+ +V +++ ++ +R
Sbjct: 554 EGMKERQEGMVSEAEEMKEWRKRVEAEMEKAEKEKENIR 592
>At1g71360 hypothetical protein
Length = 622
Score = 386 bits (991), Expect = e-107
Identities = 225/502 (44%), Positives = 316/502 (62%), Gaps = 34/502 (6%)
Query: 3 TIFLLLWCRCISHSLILVSLLYL-YVDGSEELSVGLSKWNEVNHGFCEISDTADKYFIKE 61
++ L+W L+ +S L++ +VDG + G S + V G E D +
Sbjct: 57 SLVFLIW------GLVFLSTLWISHVDGDK----GRSLVDSVEKG--EPDDERADETAES 104
Query: 62 IDACFPSEALIYSKAGDAEANGLVNESHNGRESGAYAVPADINKENT------------D 109
+DA ++S G + V+ + G G+ + + +NT +
Sbjct: 105 VDATSLESTSVHSNPG---LSSDVDIAAAGESKGSETILKQLEVDNTIVIVGNVTESKDN 161
Query: 110 SANREDHVVENSEYAVKHENDVKKSDILSRAVPLGLNEFKSRAISSKVKSGTGQSRSVIH 169
++ + N+ E K D LSRAVPLGL+EFKSRA +S+ KS +GQ VIH
Sbjct: 162 VPMKQSEINNNTVPGNDTETTGSKLDQLSRAVPLGLDEFKSRASNSRDKSLSGQVTGVIH 221
Query: 170 RLEPGGAEYNYASASKGAKVLGSNKEGKGASNILSRDKDKYLRNPCSVVGKFVIMELSEE 229
R+EPGG EYNYA+ASKGAKVL SNKE KGAS+I+ RDKDKYLRNPCS GKFV++ELSEE
Sbjct: 222 RMEPGGKEYNYAAASKGAKVLSSNKEAKGASSIICRDKDKYLRNPCSTEGKFVVIELSEE 281
Query: 230 TLVDTIEIANFEHHSSNLKDFEIHGSLNFPTNVWDLLGNFTASNVRHAQRFVLKEPKWVR 289
TLV+TI+IANFEH+SSNLKDFEI G+L +PT+ W LGNFTA N++H Q F +PKWVR
Sbjct: 282 TLVNTIKIANFEHYSSNLKDFEILGTLVYPTDTWVHLGNFTALNMKHEQNFTFADPKWVR 341
Query: 290 YLKLNLQSHYGSEFYCTLSVVEVFGVDAVERMLEDLINTQD-NLLASGEGNAD----KTI 344
YLKLNL SHYGSEFYCTLS++EV+GVDAVERMLEDLI+ QD N+L EG+ + KT+
Sbjct: 342 YLKLNLLSHYGSEFYCTLSLLEVYGVDAVERMLEDLISIQDKNILKLQEGDTEQKEKKTM 401
Query: 345 LPHPDPAVIEHVHKKPLEGINSVPASDISSSKHETANIKVPDPVEEIRQQVG-RMPGDTV 403
E K+ + + P + + + K+PDPVEEI+ Q G RMPGDTV
Sbjct: 402 QAKESFESDEDKSKQKEKEQEASPENAVVKDEVSLEKRKLPDPVEEIKHQPGSRMPGDTV 461
Query: 404 LKILMQKVRTLDVNLFVLERYMEDLNSRYVNIFKDYSKDTGEKDIVLQKIKEDIKNLIDH 463
LKILMQK+R+LDV+L VLE Y+E+ + +Y IFK+ + +++ ++ ++ +++ + +
Sbjct: 462 LKILMQKIRSLDVSLSVLESYLEERSLKYGMIFKEMDLEASKREKEVETMRLEVEGMKER 521
Query: 464 QDVIAKDASDLISWKSQVSSQL 485
++ K+A ++ W+ +V ++L
Sbjct: 522 EENTKKEAMEMRKWRMRVETEL 543
>At4g23950 putative protein
Length = 466
Score = 166 bits (421), Expect = 2e-41
Identities = 104/239 (43%), Positives = 138/239 (57%), Gaps = 26/239 (10%)
Query: 65 CFPSEALIY--SKAGDAEANGLVNESHNGRESGAYAVPADINKENTDSANRE-------D 115
C P + IY + G+ +G V+++ N S P KEN R+ +
Sbjct: 23 CVPLKKSIYFVDRIGNY-TDGSVSKTLNSTSS---VFPQATEKENNFCLLRKGQLQDVYE 78
Query: 116 HVVENSEY----AVKHENDVKKSDILSRA---VPLGLNEFKSRAISSKVKSGTGQSRSVI 168
HV+ N+ V E + K + +R V L K S V +GT
Sbjct: 79 HVLVNNALLICKVVLPERRISKKTLEARDPRYVNLEDKSLKVNGSSQLVNNGTR------ 132
Query: 169 HRLEPGGAEYNYASASKGAKVLGSNKEGKGASNILSRDKDKYLRNPCSVVGKFVIMELSE 228
+RLEP G YNYASA KGAKV+ NKE KGASN+L +D DKYLRNPCSV K+V++EL+E
Sbjct: 133 YRLEPDGNGYNYASAMKGAKVVDHNKEAKGASNVLGKDHDKYLRNPCSVSDKYVVIELAE 192
Query: 229 ETLVDTIEIANFEHHSSNLKDFEIHGSLNFPTNVWDLLGNFTASNVRHAQRFVLKEPKW 287
ETLVDT+ IANFEH+SSN K+F + GSL+FP+++W G+F A+NV+ Q F L EPKW
Sbjct: 193 ETLVDTVRIANFEHYSSNPKEFSLSGSLSFPSDMWTPAGSFAAANVKQIQSFRLPEPKW 251
Score = 47.0 bits (110), Expect = 2e-05
Identities = 28/105 (26%), Positives = 57/105 (53%), Gaps = 15/105 (14%)
Query: 403 VLKILMQKVRTLDVNLFVLERYMEDLNSRYVNIFKDYSKDTGEKDIVLQKIKEDIKNLID 462
VLK++MQKV+ +++NL +LE ++ +N + + + K ++++K K DI+ + +
Sbjct: 288 VLKVMMQKVKLIEMNLSLLEDSVKKMNDKQPEVSLEMKKTL----VLVEKSKADIREITE 343
Query: 463 HQDVI-----------AKDASDLISWKSQVSSQLNHLIQDNAVLR 496
+ + K+ DL WK+ V+S++ L + N+ LR
Sbjct: 344 WKGKMKLPMNLIFFEQEKELRDLELWKTLVASRVESLARGNSALR 388
>At4g28300 predicted proline-rich protein
Length = 496
Score = 35.0 bits (79), Expect = 0.095
Identities = 31/167 (18%), Positives = 68/167 (40%), Gaps = 9/167 (5%)
Query: 330 DNLLASGEG--NADKTILPHPDPAVIEHVHKKPLEGINSVPASDISSSKHETANIKVPDP 387
D++L S + N D + PH DPA+ K +S +S + +
Sbjct: 21 DDILCSYDDYTNQDSSNGPHSDPAIAASNSNKEFHKTRMARSSVFPTSSYSPPEDSLSQD 80
Query: 388 VEEIRQQVGRMPGDTVLKI---LMQKVRTLDVNLFVLERYMEDLNSRYVNIFKDYSKDTG 444
+ + ++ +M D +++ L ++ L++ + L++ + ++ S + +D
Sbjct: 81 ITDTVERTMKMYADNMMRFLEGLSSRLSQLELYCYNLDKTIGEMRSELTHAHEDADVKLR 140
Query: 445 EKDIVLQKIKEDIKNLIDHQDVIAKDAS----DLISWKSQVSSQLNH 487
D LQ++ ++ L D Q++ L+ +S SS H
Sbjct: 141 SLDKHLQEVHRSVQILRDKQELADTQKELAKLQLVQKESSSSSHSQH 187
>At4g18240 starch synthase-like protein
Length = 1071
Score = 34.3 bits (77), Expect = 0.16
Identities = 29/113 (25%), Positives = 57/113 (49%), Gaps = 11/113 (9%)
Query: 372 ISSSKHETANIKV-PDPVEEIRQQ-VGRMPGDTVLKILMQKVRTLDVNLFVLERYMEDLN 429
I ++ E A++++ + +E++R + + + D + L +++ TL + L +E L
Sbjct: 241 IKTAAQEKAHVELLEEQLEKLRHEMISPIESDGYVLALSKELETLKLENLSLRNDIEMLK 300
Query: 430 SRYVNIFKDYSKDTGEKDIVLQK----IKEDIKNLIDHQDVIAKDASDLISWK 478
S D KDTGE+ +VL+K ++ +K+L V +D S L + K
Sbjct: 301 SEL-----DSVKDTGERVVVLEKECSGLESSVKDLESKLSVSQEDVSQLSTLK 348
>At5g07820 putative protein
Length = 561
Score = 33.5 bits (75), Expect = 0.28
Identities = 36/186 (19%), Positives = 78/186 (41%), Gaps = 9/186 (4%)
Query: 28 DGSEELSVGLSKWNEVNHGFCEISDTADKYFIKEIDACFPSEALIYSKAGDAEANGLVNE 87
D E+ + + K + ++ DT +ID + ++ + DA+ ++E
Sbjct: 217 DVLEKTDLEVKKVSRISENKSSKEDTLKNKEKAKIDEPVRCDDVLEKTSLDAQKVSRISE 276
Query: 88 SHNGRESGAYAVPADINKENTDSANREDHVVENSEYAVKHENDVKKSDILSRAVPLGLNE 147
+ N +E + + K N D R D VE + Y V+ + KK + +++V ++E
Sbjct: 277 NKNSKEERLKNLK-NKEKTNIDEPVRPDDAVEKTLYVVESSVEKKKKKMSTKSVK--ISE 333
Query: 148 FKSRAISSKVKSGTGQSRSVIHRLEP------GGAEYNYASASKGAKVLGSNKEGKGASN 201
+ + ++S +S S++ L P G + +K K+ G++N
Sbjct: 334 TQQSSEKKIIRSTGKKSLSLLPSLPPPSEVVTGSDPRPIRQTTSRSKTSLPEKKQSGSAN 393
Query: 202 ILSRDK 207
+++ K
Sbjct: 394 LVTNPK 399
>At4g36520 trichohyalin like protein
Length = 1432
Score = 31.6 bits (70), Expect = 1.1
Identities = 45/189 (23%), Positives = 76/189 (39%), Gaps = 23/189 (12%)
Query: 24 YLYVDGSEELSVGLSKWNEVNHGFCEISDTADKYFIKEID-ACFPSEALIYSKAGDAEAN 82
Y +DGS E ++ +K ++++ G +S + I+ D P + G + N
Sbjct: 177 YDSIDGSTEFNISYNKASQISGGETNMSSSG---VIRVADLGAIPGYTVAVD--GTTKLN 231
Query: 83 GLVNESHNGRESGAYAVPADINKENTDSANREDHVVENSEYAVK-HENDVKKSDILSRAV 141
+ S GR + + D + V SE +K H + SRA
Sbjct: 232 KVTGPSSVGRGTSRFVFE--------DKYYSSEPFVTVSEIGLKTHPTGIPPP---SRAA 280
Query: 142 PLGLNEFKSRAISSKVKSGTGQSRSVIHRLEPG--GAEYNYASASKGAKVLGSNKEGKGA 199
P+ ++F+S A +SK TG SV P E + SA+ +L + + K A
Sbjct: 281 PILKSDFRSSASNSKT---TGSQGSVDSSSSPTFFDVEVDANSAAVREAMLKAEAKLKSA 337
Query: 200 SNILSRDKD 208
+L R +D
Sbjct: 338 KELLERKRD 346
>At2g34730 putative myosin heavy chain
Length = 829
Score = 31.2 bits (69), Expect = 1.4
Identities = 35/150 (23%), Positives = 65/150 (43%), Gaps = 9/150 (6%)
Query: 353 IEHVHKKPLEGINSVPASDISSSKHETANIKVPDPVEEIRQQVGRMPGDTV--LKILMQK 410
+E +H+K +NSV + + E++ +P+ E +R P + + KI M K
Sbjct: 284 VEQLHRKMSGSLNSVSSVWENGKHEESSTGLIPEHNETLRHM---SPDEMINHFKIEMNK 340
Query: 411 V-RTLDVNLFVLERYMEDLNSRYVNIFKDYSKDTGEKDIVLQKIKEDIKNLIDHQDVIAK 469
+ R D + L +Y+N+ + S KD L +K+ I +I D I
Sbjct: 341 MKRDHDYKIQELTEQCFTFKRKYLNLTERGSFSFVGKDKELGALKKKIPFVISKLDKILM 400
Query: 470 DASDLISW---KSQVSSQLNHLIQDNAVLR 496
+ +S + + QL+ L+ +N L+
Sbjct: 401 EDEKFVSEGKNDAGLKRQLDSLLLENRQLK 430
>At2g43650 unknown protein
Length = 654
Score = 30.8 bits (68), Expect = 1.8
Identities = 18/71 (25%), Positives = 39/71 (54%), Gaps = 4/71 (5%)
Query: 422 ERYMEDLNSRYVNIFKDYS--KDTGEKDIVLQKIKEDIKNLI--DHQDVIAKDASDLISW 477
E ME+++ + K + K+ G+KD +++IK+DI +L + DV+ A +++
Sbjct: 183 ELTMEEISDKGKQATKSITDKKEKGDKDTHVEEIKKDINSLSKEEQMDVVYSSAPEIVGL 242
Query: 478 KSQVSSQLNHL 488
S+++ + L
Sbjct: 243 LSELNDAVEEL 253
>At2g41450 hypothetical protein
Length = 991
Score = 30.8 bits (68), Expect = 1.8
Identities = 16/55 (29%), Positives = 30/55 (54%), Gaps = 3/55 (5%)
Query: 94 SGAYAVPADINKENTDS---ANREDHVVENSEYAVKHENDVKKSDILSRAVPLGL 145
SGA+ V KE+ AN H++ + +Y +K++ D+K + + ++A P L
Sbjct: 391 SGAWIVSPSWLKESVREGRFANEASHILHDEDYQLKYDTDLKSTVLRAKARPNSL 445
>At4g26130 uncharacterized protein
Length = 286
Score = 30.4 bits (67), Expect = 2.3
Identities = 49/227 (21%), Positives = 88/227 (38%), Gaps = 36/227 (15%)
Query: 230 TLVDTIEIANFEHHSSNLKDFEIH--GSLNFPTNVWDLLGNFTASNVRHAQRFVLKEPKW 287
+L+D ++ NF ++S + EIH GS P LL + N+ +
Sbjct: 64 SLIDRVKSINFHLYNSPSPESEIHYSGSDPNPNPPPSLLQRVKSINM-----------PY 112
Query: 288 VRYLKLNLQSHYGSEFYCTLSVVEVFGVDAVERMLEDLINTQDNLLASGEGNADKTIL-- 345
++ + N + Y + + E VD ++++ ED + T+ A K+I
Sbjct: 113 FKFPQHNSEGDYAA-YELMTQPDETNRVDPIDKIPEDDVTTEPRFGAPSLLQRVKSIKLP 171
Query: 346 ------PHPDPAVIEHVHKKPLEGINSVPASDISSSKHETANIKVPDPVEEIRQQVGRMP 399
P P P V H K +S ++ K + A K+ E + +GR
Sbjct: 172 SLYRSDPDPTPEVQTHTRTKS-------ESSKPATKKKKKATKKMMKSASE--RHIGREE 222
Query: 400 GDTVLKILMQKVRTLDVNLFVL-----ERYMEDLNSRYVNIFKDYSK 441
+TV + ++ T+ V E ++D S ++N FK K
Sbjct: 223 EETVEAVEKRRPETMRVERTTSIGDGGEEGVDDKASNFINKFKQQLK 269
>At4g09150 unknown protein
Length = 1097
Score = 30.4 bits (67), Expect = 2.3
Identities = 42/218 (19%), Positives = 90/218 (41%), Gaps = 33/218 (15%)
Query: 281 VLKEPKWVRYLKLNLQSHYGSEFYCTLS-------VVEVFGVDAVE-RMLEDLINTQDNL 332
+LK P + YLK + S +GS + S ++ V G E + +D ++ N
Sbjct: 792 LLKGPAGLEYLKKSFSSRHGSPDQASSSLPLTKRWLLSVRGEAEREWKEHKDALSAVINN 851
Query: 333 LASGEGNADKTILPHPDPAVIEHVHKKPLEGINSVPASDISSSKHETANIKVPDPVEEIR 392
+ G T+ + + + V+ + P ++S K ET ++ V + ++
Sbjct: 852 HSGSSGLPSTTMRTGGNVSSVSKVNTPS----SPFPGIELSECKGETVDLLVRIGLLKMV 907
Query: 393 QQVGRMPGDTV-------------LKILMQKVRTLDVNLFVLERYMEDLNSRYVN----- 434
++G + +TV ++ +QK+ + +++ +L++ + NS ++
Sbjct: 908 SEIGGLTLETVPETFQLNLSRLRDVQSQIQKITLVSISVLILQQTLVSENSSSIDMEAIT 967
Query: 435 ---IFKDYSKDTGEKDIVLQKIKEDIKNLIDHQDVIAK 469
I + Y + D L +I E + L+D D K
Sbjct: 968 RTCINRLYEMLDAKPDAGLSEIMETLSELLDSNDAETK 1005
>At5g42390 pitrilysin
Length = 1265
Score = 30.0 bits (66), Expect = 3.1
Identities = 54/205 (26%), Positives = 85/205 (41%), Gaps = 28/205 (13%)
Query: 150 SRAISSKVKSGTGQSRSVIHRLEPGGAEYNYASASKGAKVLG--SNKEGKGASNILSRDK 207
S I+ K T +SR+ + RL GG S SKGA V+G + EG +
Sbjct: 768 SNGIAVNYKKSTTESRAGVMRLIVGGGRAAETSDSKGAVVVGVRTLSEGGRVGDFSREQV 827
Query: 208 DKYLRN---PCSV--VGKFVIMELSEETLVDTIEIANFE------HHSSNLKDFEIHGS- 255
+ + N CS+ +F+ ME TL D A F+ S L+D
Sbjct: 828 ELFCVNHLINCSLESTEEFIAMEF-RFTLRDNGMQAAFQLLHMVLERSVWLEDAFDRARQ 886
Query: 256 --LNFPTNVWDLLGNFTASNVRHA-----QRFVLKEPKWVRYLKLN-----LQSHYGSEF 303
L++ ++ L TA + A +RFV PK ++ L L + SH+ +
Sbjct: 887 LYLSYFRSIPKSLERATAHKLMIAMLNGDERFVEPTPKSLQSLNLESVKDAVMSHFVGD- 945
Query: 304 YCTLSVVEVFGVDAVERMLEDLINT 328
+S+V F + +ER + D + T
Sbjct: 946 NMEVSIVGDFSEEEIERCILDYLGT 970
>At1g14380 unknown protein
Length = 664
Score = 30.0 bits (66), Expect = 3.1
Identities = 41/189 (21%), Positives = 68/189 (35%), Gaps = 20/189 (10%)
Query: 75 KAGDAEANGLVNES----HNGRESGAYAV---PADINKENTDSANREDHVVENSEYAVKH 127
KA + L NES HN R+S + + P +I E + + + S+ A
Sbjct: 298 KASTLSKDPLRNESDKANHNSRKSRSGSKEGSPLEIKDEKPSPSLKRSSLSNGSKKATLR 357
Query: 128 ENDVKKSDILSRAVPLGLNEFKSRAISSKVKSGTGQSRSVIHRLEPGGAEYNYASASKGA 187
+ KK DI +V + KV + I E +
Sbjct: 358 SAEKKKKDIPDSSVQI--------QPEGKVSENVLEGGDNIEFAEKEKDTTDSVQIESEG 409
Query: 188 KVLGSNKEGKGASNILSRDKDKYLRNPCSVVGKFVIMELSEETLVDTIEIANFEHHSSNL 247
KV G+ EG NI+ +K+K +P + + ++E D IE + E + +
Sbjct: 410 KVSGNVLEGGEGENIVFTEKEKDTADPVQIEPERKVLEEG-----DNIESSGKEKDTGDS 464
Query: 248 KDFEIHGSL 256
E G +
Sbjct: 465 VQIESEGKV 473
>At5g27395 putative protein
Length = 241
Score = 29.6 bits (65), Expect = 4.0
Identities = 28/83 (33%), Positives = 37/83 (43%), Gaps = 5/83 (6%)
Query: 28 DGSEELSVGLSKWNEVNHGFCEISDTADKYFIKEIDACF-PSEALIYS--KAGDAEANGL 84
DGSE LSV + N++ F E D +D +K F P A IYS E GL
Sbjct: 127 DGSEVLSVVIRPSNQLKITFLEAKDISDLGSLKAAARLFVPGAATIYSARTIKVKEEEGL 186
Query: 85 VNE--SHNGRESGAYAVPADINK 105
N GR+ A+ A +N+
Sbjct: 187 RNYYFYEFGRDEERIALVASVNR 209
>At4g32850 poly(A) polymerase like protein
Length = 741
Score = 29.6 bits (65), Expect = 4.0
Identities = 11/24 (45%), Positives = 16/24 (65%)
Query: 465 DVIAKDASDLISWKSQVSSQLNHL 488
D++A DA DL++WK V S+ L
Sbjct: 380 DIVAADAEDLLAWKGWVESRFRQL 403
>At3g14020 putative transcription factor
Length = 308
Score = 29.6 bits (65), Expect = 4.0
Identities = 18/59 (30%), Positives = 32/59 (53%), Gaps = 2/59 (3%)
Query: 201 NILSRDKDKYLRNPCSVVGKFVIMELSEETLVDTIEIANFEHHSSNLKDFEIHGSLNFP 259
NI+ Y NP + +G I ++S++++V IE A++ H + F +G L+FP
Sbjct: 85 NIVVTHLSGYKENPENPIGSHSISKVSQDSVVLPIEAASWPLHGNVTPHF--NGFLSFP 141
>At2g25850 nuclear poly(A) polymerase
Length = 759
Score = 29.6 bits (65), Expect = 4.0
Identities = 12/24 (50%), Positives = 16/24 (66%)
Query: 465 DVIAKDASDLISWKSQVSSQLNHL 488
DV+A DA DL++WK V S+ L
Sbjct: 382 DVLAADAEDLLAWKGWVESRFRQL 405
>At5g66750 SWI2/SNF2-like protein (gb|AAD28303.1)
Length = 764
Score = 29.3 bits (64), Expect = 5.2
Identities = 37/185 (20%), Positives = 72/185 (38%), Gaps = 28/185 (15%)
Query: 67 PSEALIYSKAGDAEANG-----LVNESHNGRESGAYAVPADIN-KENTDSANREDHVVEN 120
P+ ++ + +A+G ++NE N E V +I +N DS+ + + +
Sbjct: 11 PASEMVSDGKTEKDASGDSPTSVLNEEENCEEKSVTVVEEEILLAKNGDSSLISEAMAQE 70
Query: 121 SEYAVKHENDVKKSDILSRAVPLGLNEFKSRAISSKVKSGTGQSRSVIHRLEP---GGAE 177
E +K D +K++ AV LNE + + + S ++ ++E G E
Sbjct: 71 EEQLLKLREDEEKANNAGSAVAPNLNETQFTKLDELLTQTQLYSEFLLEKMEDITINGIE 130
Query: 178 YNYASAS----------KGAKVLGSNKEGKGASNILSRDKDKYLRNPCSVVGKFVIMELS 227
A K A + K + + ++SR K+ G+ + +L+
Sbjct: 131 SESQKAEPEKTGRGRKRKAASQYNNTKAKRAVAAMISRSKED---------GETINSDLT 181
Query: 228 EETLV 232
EE V
Sbjct: 182 EEETV 186
>At5g41790 myosin heavy chain-like protein
Length = 1305
Score = 29.3 bits (64), Expect = 5.2
Identities = 25/95 (26%), Positives = 43/95 (44%), Gaps = 14/95 (14%)
Query: 401 DTVLKILMQKVRTLDVNLFVLERYMEDLNSRYVNIFKDYSKDTGEKDIVLQKIKEDIKNL 460
+T LK+L Q+V L +L E + L+S + I T E K++E + L
Sbjct: 491 ETQLKLLEQRVVDLSASLNAAEEEKKSLSSMILEI-------TDELKQAQSKVQELVTEL 543
Query: 461 IDHQDVIAKDASDLISW-------KSQVSSQLNHL 488
+ +D + + ++L S+ K SSQ+ L
Sbjct: 544 AESKDTLTQKENELSSFVEVHEAHKRDSSSQVKEL 578
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.314 0.133 0.377
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,285,286
Number of Sequences: 26719
Number of extensions: 503695
Number of successful extensions: 1218
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 22
Number of HSP's that attempted gapping in prelim test: 1204
Number of HSP's gapped (non-prelim): 33
length of query: 496
length of database: 11,318,596
effective HSP length: 103
effective length of query: 393
effective length of database: 8,566,539
effective search space: 3366649827
effective search space used: 3366649827
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 62 (28.5 bits)
Medicago: description of AC148651.5


