

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
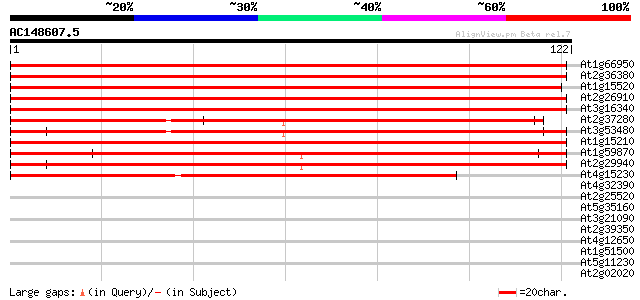
Query= AC148607.5 - phase: 0
(122 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At1g66950 ABC transporter, putative 159 3e-40
At2g36380 putative ABC transporter 154 1e-38
At1g15520 hypothetical protein 154 1e-38
At2g26910 putative ABC transporter 145 5e-36
At3g16340 putative ABC transporter 139 3e-34
At2g37280 putative ABC transporter 137 1e-33
At3g53480 ABC transporter - like protein 136 3e-33
At1g15210 putative ABC transporter 135 5e-33
At1g59870 putative ABC transporter (At1g59870) 134 1e-32
At2g29940 putative ABC transporter 117 1e-27
At4g15230 ABC transporter like protein 93 3e-20
At4g32390 putative protein 29 0.49
At2g25520 putative phosphate/phosphoenolpyruvate translocator pr... 29 0.49
At5g35160 unknown protein 28 0.83
At3g21090 ABC transporter, putative 28 0.83
At2g39350 ABC transporter like protein 28 1.4
At4g12650 putative protein 27 1.9
At1g51500 ATP-dependent transmembrane transporter, putative 27 1.9
At5g11230 putative protein 27 2.4
At2g02020 putative peptide/amino acid transporter 27 2.4
>At1g66950 ABC transporter, putative
Length = 1434
Score = 159 bits (402), Expect = 3e-40
Identities = 70/121 (57%), Positives = 93/121 (76%)
Query: 1 MKRNSFVYIFKLCQLAIMAMIAMTIFLRTEMHRDSVAHGDIYVGALFYGCIVILFIGVAE 60
MKRNSFVY+FK Q+ IM++I MT++LRTEMH +V G + GA+F+ I ++F G+AE
Sbjct: 539 MKRNSFVYVFKTVQITIMSLITMTVYLRTEMHVGTVRDGQKFYGAMFFSLINVMFNGLAE 598
Query: 61 LSMVVSRLPVFYKQRGYLFFPPWAYALPAWILKIPLTFVEVAVWVILTYYVIGFDPYIGR 120
L+ V RLPVFYKQR +LF+PPWA+ALPAW+LKIPL+ +E +W+ LTYY IGF P R
Sbjct: 599 LAFTVMRLPVFYKQRDFLFYPPWAFALPAWLLKIPLSLIESGIWIGLTYYTIGFAPSAAR 658
Query: 121 Y 121
+
Sbjct: 659 F 659
Score = 30.0 bits (66), Expect = 0.29
Identities = 23/111 (20%), Positives = 50/111 (44%), Gaps = 5/111 (4%)
Query: 4 NSFVYIFKLCQLAIMAMIAMTIFLRTEMHRD-SVAHGDIYVGALFYGCIVILFIGVAELS 62
N+ ++ + + +I I +TE +D + G +Y LF G L + +
Sbjct: 1178 NAIRFLMTVVIGVLFGLIFWQIGTKTENEQDLNNFFGAMYAAVLFLGA---LNAATVQPA 1234
Query: 63 MVVSRLPVFYKQRGYLFFPPWAYALPAWILKIPLTFVEVAVWVILTYYVIG 113
+ + R VFY+++ + YA+ ++I ++ V+ ++ Y +IG
Sbjct: 1235 IAIERT-VFYREKAAGMYSAIPYAISQVAVEIMYNTIQTGVYTLILYSMIG 1284
>At2g36380 putative ABC transporter
Length = 1450
Score = 154 bits (388), Expect = 1e-38
Identities = 68/121 (56%), Positives = 92/121 (75%)
Query: 1 MKRNSFVYIFKLCQLAIMAMIAMTIFLRTEMHRDSVAHGDIYVGALFYGCIVILFIGVAE 60
MKRNSFVY+FK Q+ IM++IAMT++ RTEMH +V G + GALF+ I ++F G+AE
Sbjct: 537 MKRNSFVYVFKTVQITIMSLIAMTVYFRTEMHVGTVQDGQKFYGALFFSLINLMFNGMAE 596
Query: 61 LSMVVSRLPVFYKQRGYLFFPPWAYALPAWILKIPLTFVEVAVWVILTYYVIGFDPYIGR 120
L+ V RLPVF+KQR +LF+PPWA+ALP ++LKIPL+ +E +W+ LTYY IGF P R
Sbjct: 597 LAFTVMRLPVFFKQRDFLFYPPWAFALPGFLLKIPLSLIESVIWIALTYYTIGFAPSAAR 656
Query: 121 Y 121
+
Sbjct: 657 F 657
Score = 31.2 bits (69), Expect = 0.13
Identities = 17/75 (22%), Positives = 37/75 (48%), Gaps = 6/75 (8%)
Query: 47 FYGCI--VILFIGVAELSMVVSRLP----VFYKQRGYLFFPPWAYALPAWILKIPLTFVE 100
F+G + +LF+G + V + VFY+++ + YA+ ++I ++
Sbjct: 1228 FFGAMYAAVLFLGATNAATVQPAVAIERTVFYREKAAGMYSAIPYAISQVAVEIMYNTIQ 1287
Query: 101 VAVWVILTYYVIGFD 115
V+ ++ Y +IG+D
Sbjct: 1288 TGVYTLILYSMIGYD 1302
>At1g15520 hypothetical protein
Length = 1423
Score = 154 bits (388), Expect = 1e-38
Identities = 67/120 (55%), Positives = 92/120 (75%)
Query: 1 MKRNSFVYIFKLCQLAIMAMIAMTIFLRTEMHRDSVAHGDIYVGALFYGCIVILFIGVAE 60
MKRNSFVY FK QL +MA + MT+F RTEM + + G +Y GALF+ ++++F G++E
Sbjct: 518 MKRNSFVYYFKFGQLLVMAFLTMTLFFRTEMQKKTEVDGSLYTGALFFILMMLMFNGMSE 577
Query: 61 LSMVVSRLPVFYKQRGYLFFPPWAYALPAWILKIPLTFVEVAVWVILTYYVIGFDPYIGR 120
LSM +++LPVFYKQR LF+P W Y+LP W+LKIP++F+E A+ +TYYVIGFDP +GR
Sbjct: 578 LSMTIAKLPVFYKQRDLLFYPAWVYSLPPWLLKIPISFMEAALTTFITYYVIGFDPNVGR 637
Score = 36.2 bits (82), Expect = 0.004
Identities = 25/90 (27%), Positives = 43/90 (47%), Gaps = 7/90 (7%)
Query: 28 RTEMHRD-SVAHGDIYVGALFYGCIVILFIGVAELSMVVS-RLPVFYKQRGYLFFPPWAY 85
+T+ +D S A G +Y LF G A + VV+ VFY+++ + Y
Sbjct: 1194 KTKTRQDLSNAMGSMYTAVLFLG-----LQNAASVQPVVNVERTVFYREQAAGMYSAMPY 1248
Query: 86 ALPAWILKIPLTFVEVAVWVILTYYVIGFD 115
A ++IP V+ V+ ++ Y +IGF+
Sbjct: 1249 AFAQVFIEIPYVLVQAIVYGLIVYAMIGFE 1278
>At2g26910 putative ABC transporter
Length = 1420
Score = 145 bits (366), Expect = 5e-36
Identities = 59/121 (48%), Positives = 89/121 (72%)
Query: 1 MKRNSFVYIFKLCQLAIMAMIAMTIFLRTEMHRDSVAHGDIYVGALFYGCIVILFIGVAE 60
MK+N+F+Y+FK QL ++A+I MT+F RT MH +++ G+IY+G+L++ ++ILF G E
Sbjct: 499 MKQNAFIYVFKFVQLLLVALITMTVFCRTTMHHNTIDDGNIYLGSLYFSMVIILFNGFTE 558
Query: 61 LSMVVSRLPVFYKQRGYLFFPPWAYALPAWILKIPLTFVEVAVWVILTYYVIGFDPYIGR 120
+ M+V++LPV YK R F+P WAY LP+W+L IP + +E A WV +TYY IG+DP R
Sbjct: 559 VPMLVAKLPVLYKHRDLHFYPSWAYTLPSWLLSIPTSIIESATWVAVTYYTIGYDPLFSR 618
Query: 121 Y 121
+
Sbjct: 619 F 619
>At3g16340 putative ABC transporter
Length = 1416
Score = 139 bits (350), Expect = 3e-34
Identities = 63/121 (52%), Positives = 88/121 (72%)
Query: 1 MKRNSFVYIFKLCQLAIMAMIAMTIFLRTEMHRDSVAHGDIYVGALFYGCIVILFIGVAE 60
MKRN+F YI K Q+ IMA+IA T++LRTEM + + G +Y+GAL + IV +F G AE
Sbjct: 511 MKRNAFFYITKTVQIIIMALIASTVYLRTEMGTKNESDGAVYIGALMFSMIVNMFNGFAE 570
Query: 61 LSMVVSRLPVFYKQRGYLFFPPWAYALPAWILKIPLTFVEVAVWVILTYYVIGFDPYIGR 120
L++++ RLPVFYKQR LF PPW ++LP ++L IP++ E VWV +TYY+IGF P + R
Sbjct: 571 LALMIQRLPVFYKQRDLLFHPPWTFSLPTFLLGIPISIFESVVWVTITYYMIGFAPELSR 630
Query: 121 Y 121
+
Sbjct: 631 F 631
Score = 36.6 bits (83), Expect = 0.003
Identities = 22/107 (20%), Positives = 51/107 (47%), Gaps = 7/107 (6%)
Query: 19 AMIAMTIFLRTEMHRDSVAHGDIYVGALFYGCIVILFIGVAELS----MVVSRLPVFYKQ 74
A++ +IF + R++ +GA++ +LF+GV S ++ VFY++
Sbjct: 1170 AVMLGSIFWKVGTKRENANDLTKVIGAMY---AAVLFVGVNNSSSVQPLIAVERSVFYRE 1226
Query: 75 RGYLFFPPWAYALPAWILKIPLTFVEVAVWVILTYYVIGFDPYIGRY 121
R + YAL + +IP ++ + ++ Y ++ F+ + ++
Sbjct: 1227 RAAEMYSALPYALAQVVCEIPYVLIQTTYYTLIIYAMMCFEWTLAKF 1273
>At2g37280 putative ABC transporter
Length = 1413
Score = 137 bits (345), Expect = 1e-33
Identities = 65/116 (56%), Positives = 83/116 (71%), Gaps = 1/116 (0%)
Query: 1 MKRNSFVYIFKLCQLAIMAMIAMTIFLRTEMHRDSVAHGDIYVGALFYGCIVILFIGVAE 60
MKRN FVY+FK QL + A+I MT+F+RT M D + HG+ Y+ LF+ +V+L G+ E
Sbjct: 503 MKRNYFVYLFKTFQLVLAAIITMTVFIRTRMDID-IIHGNSYMSCLFFATVVLLVDGIPE 561
Query: 61 LSMVVSRLPVFYKQRGYLFFPPWAYALPAWILKIPLTFVEVAVWVILTYYVIGFDP 116
LSM V RL VFYKQ+ F+P WAYA+PA +LKIPL+F E VW LTYYVIG+ P
Sbjct: 562 LSMTVQRLSVFYKQKQLCFYPAWAYAIPATVLKIPLSFFESLVWTCLTYYVIGYTP 617
Score = 38.9 bits (89), Expect = 6e-04
Identities = 25/76 (32%), Positives = 44/76 (57%), Gaps = 7/76 (9%)
Query: 43 VGALFYGCIVILFIGV----AELSMVVSRLPVFYKQRGYLFFPPWAYALPAWILKIPLTF 98
+GA+ YG ++LF+G+ + L + V Y++R + +AYAL + +IP F
Sbjct: 1193 LGAI-YG--LVLFVGINNCTSALQYFETERNVMYRERFAGMYSAFAYALAQVVTEIPYIF 1249
Query: 99 VEVAVWVILTYYVIGF 114
++ A +VI+ Y +IGF
Sbjct: 1250 IQSAEFVIVIYPMIGF 1265
>At3g53480 ABC transporter - like protein
Length = 1450
Score = 136 bits (342), Expect = 3e-33
Identities = 65/121 (53%), Positives = 83/121 (67%), Gaps = 1/121 (0%)
Query: 1 MKRNSFVYIFKLCQLAIMAMIAMTIFLRTEMHRDSVAHGDIYVGALFYGCIVILFIGVAE 60
MKRN FVYIFK QL + A I MT+F+RT M D + HG+ Y+ ALF+ I++L G E
Sbjct: 538 MKRNYFVYIFKTAQLVMAAFITMTVFIRTRMGID-IIHGNSYMSALFFALIILLVDGFPE 596
Query: 61 LSMVVSRLPVFYKQRGYLFFPPWAYALPAWILKIPLTFVEVAVWVILTYYVIGFDPYIGR 120
LSM RL VFYKQ+ F+P WAYA+PA +LK+PL+F E VW L+YYVIG+ P R
Sbjct: 597 LSMTAQRLAVFYKQKQLCFYPAWAYAIPATVLKVPLSFFESLVWTCLSYYVIGYTPEASR 656
Query: 121 Y 121
+
Sbjct: 657 F 657
Score = 43.1 bits (100), Expect = 3e-05
Identities = 30/112 (26%), Positives = 57/112 (50%), Gaps = 7/112 (6%)
Query: 9 IFKLCQLAIMAMIAMTIFLRTEMHRDSVAHGDIYVGALFYGCIVILFIGV----AELSMV 64
+ ++ + ++I +F + + D+ GA+ YG ++LF+G+ + L
Sbjct: 1196 LMRMMHTLVSSLIFGALFWKQGQNLDTQQSMFTVFGAI-YG--LVLFLGINNCASALQYF 1252
Query: 65 VSRLPVFYKQRGYLFFPPWAYALPAWILKIPLTFVEVAVWVILTYYVIGFDP 116
+ V Y++R + AYAL + +IP F++ A +VI+TY +IGF P
Sbjct: 1253 ETERNVMYRERFAGMYSATAYALGQVVTEIPYIFIQAAEFVIVTYPMIGFYP 1304
>At1g15210 putative ABC transporter
Length = 1442
Score = 135 bits (340), Expect = 5e-33
Identities = 61/121 (50%), Positives = 84/121 (69%)
Query: 1 MKRNSFVYIFKLCQLAIMAMIAMTIFLRTEMHRDSVAHGDIYVGALFYGCIVILFIGVAE 60
MKRNSF Y+FK Q+ I+A I T++LRTEMH + +IYVG+L + IV +F G+AE
Sbjct: 533 MKRNSFFYVFKTVQIIIIAAITSTLYLRTEMHTRNEIDANIYVGSLLFAMIVNMFNGLAE 592
Query: 61 LSMVVSRLPVFYKQRGYLFFPPWAYALPAWILKIPLTFVEVAVWVILTYYVIGFDPYIGR 120
++M + RLPVFYKQR LF PPW Y LP ++L IP++ E W+++TYY IG+ P R
Sbjct: 593 MAMTIQRLPVFYKQRDLLFHPPWTYTLPTFLLGIPISIFESTAWMVVTYYSIGYAPDAER 652
Query: 121 Y 121
+
Sbjct: 653 F 653
Score = 35.8 bits (81), Expect = 0.005
Identities = 23/105 (21%), Positives = 50/105 (46%), Gaps = 8/105 (7%)
Query: 15 LAIMAMIAMTIFLRTEMHRDSVAHGDIYVGALFYGCIVILFIGVAELS----MVVSRLPV 70
LA MI ++F + R +V + +GA++ ++F+G+ S MV V
Sbjct: 1193 LATSLMIG-SVFWQIGGKRSNVQDLTMVIGAIY---AAVVFVGINNCSTVQPMVAVERTV 1248
Query: 71 FYKQRGYLFFPPWAYALPAWILKIPLTFVEVAVWVILTYYVIGFD 115
FY+++ + YA+ ++P ++ + ++ Y ++GF+
Sbjct: 1249 FYREKAAGMYSAIPYAISQVTCELPYVLIQTTYYSLIIYSMVGFE 1293
>At1g59870 putative ABC transporter (At1g59870)
Length = 1469
Score = 134 bits (336), Expect = 1e-32
Identities = 59/121 (48%), Positives = 87/121 (71%)
Query: 1 MKRNSFVYIFKLCQLAIMAMIAMTIFLRTEMHRDSVAHGDIYVGALFYGCIVILFIGVAE 60
M+RN+F Y+FK Q+ I+A I T+FLRTEM+ + ++Y+GAL +G I+ +F G AE
Sbjct: 535 MQRNAFFYVFKTVQIVIIAAITSTLFLRTEMNTRNEGDANLYIGALLFGMIINMFNGFAE 594
Query: 61 LSMVVSRLPVFYKQRGYLFFPPWAYALPAWILKIPLTFVEVAVWVILTYYVIGFDPYIGR 120
++M+VSRLPVFYKQR LF+P W ++LP ++L IP + +E W+++TYY IGF P R
Sbjct: 595 MAMMVSRLPVFYKQRDLLFYPSWTFSLPTFLLGIPSSILESTAWMVVTYYSIGFAPDASR 654
Query: 121 Y 121
+
Sbjct: 655 F 655
Score = 40.4 bits (93), Expect = 2e-04
Identities = 22/101 (21%), Positives = 49/101 (47%), Gaps = 7/101 (6%)
Query: 19 AMIAMTIFLRTEMHRDSVAHGDIYVGALFYGCIVILFIGVAELS----MVVSRLPVFYKQ 74
+++ T+F + +R + + +GAL+ I+F+G+ S MV VFY++
Sbjct: 1223 SLLIGTVFWQIGGNRSNAGDLTMVIGALY---AAIIFVGINNCSTVQPMVAVERTVFYRE 1279
Query: 75 RGYLFFPPWAYALPAWILKIPLTFVEVAVWVILTYYVIGFD 115
R + YA+ ++P ++ + ++ Y ++GF+
Sbjct: 1280 RAAGMYSAMPYAISQVTCELPYVLIQTVYYSLIVYAMVGFE 1320
>At2g29940 putative ABC transporter
Length = 1443
Score = 117 bits (294), Expect = 1e-27
Identities = 46/121 (38%), Positives = 81/121 (66%)
Query: 1 MKRNSFVYIFKLCQLAIMAMIAMTIFLRTEMHRDSVAHGDIYVGALFYGCIVILFIGVAE 60
+KR+ F+Y F+ CQ+ + ++ T+FL+T +H S G+ Y+ LF+G + ++F G +E
Sbjct: 542 IKRHKFLYTFRTCQVGFVGLVTATVFLKTRLHPTSEQFGNEYLSCLFFGLVHMMFNGFSE 601
Query: 61 LSMVVSRLPVFYKQRGYLFFPPWAYALPAWILKIPLTFVEVAVWVILTYYVIGFDPYIGR 120
L +++SRLPVFYKQR F P W++++ +W+L++P + +E VW + Y+ +G P GR
Sbjct: 602 LPLMISRLPVFYKQRDNSFHPAWSWSIASWLLRVPYSVLEAVVWSGVVYFTVGLAPSAGR 661
Query: 121 Y 121
+
Sbjct: 662 F 662
Score = 47.0 bits (110), Expect = 2e-06
Identities = 31/117 (26%), Positives = 55/117 (46%), Gaps = 7/117 (5%)
Query: 9 IFKLCQLAIMAMIAMTIFLRTEMHRDSVAHGDIYVGALFYGCIVILFIGVAELS----MV 64
+ +L I A I T+F R S +GAL+ C LF+GV+ S +V
Sbjct: 1189 LVRLVFTTIAAFILGTVFWDIGSKRTSSQDLITVMGALYSAC---LFLGVSNASSVQPIV 1245
Query: 65 VSRLPVFYKQRGYLFFPPWAYALPAWILKIPLTFVEVAVWVILTYYVIGFDPYIGRY 121
VFY+++ + P YA +++IP + ++ ++TY+ IGF+ ++
Sbjct: 1246 SIERTVFYREKAAGMYAPIPYAAAQGLVEIPYILTQTILYGVITYFTIGFERTFSKF 1302
>At4g15230 ABC transporter like protein
Length = 979
Score = 93.2 bits (230), Expect = 3e-20
Identities = 46/97 (47%), Positives = 66/97 (67%), Gaps = 1/97 (1%)
Query: 1 MKRNSFVYIFKLCQLAIMAMIAMTIFLRTEMHRDSVAHGDIYVGALFYGCIVILFIGVAE 60
MKRNSF+Y+FK L A++ MT+FL+ DS+ HG+ +G+LF +L G+ E
Sbjct: 236 MKRNSFIYLFKSALLVFNALVTMTVFLQVGATTDSL-HGNYLMGSLFTALFRLLADGLPE 294
Query: 61 LSMVVSRLPVFYKQRGYLFFPPWAYALPAWILKIPLT 97
L++ +SRL VF KQ+ F+P WAYA+P+ ILKIPL+
Sbjct: 295 LTLTISRLGVFCKQKDLYFYPAWAYAIPSIILKIPLS 331
Score = 35.4 bits (80), Expect = 0.007
Identities = 13/56 (23%), Positives = 31/56 (55%)
Query: 59 AELSMVVSRLPVFYKQRGYLFFPPWAYALPAWILKIPLTFVEVAVWVILTYYVIGF 114
A ++ + + VFY++R + WAY+ ++++P + ++ + I+ Y IG+
Sbjct: 776 AVINFIAAERNVFYRERFARMYSSWAYSFSQVLIEVPYSLLQSLLCTIIVYPTIGY 831
>At4g32390 putative protein
Length = 350
Score = 29.3 bits (64), Expect = 0.49
Identities = 12/28 (42%), Positives = 20/28 (70%)
Query: 91 ILKIPLTFVEVAVWVILTYYVIGFDPYI 118
I KI L++ VA+W+ L++ VI ++ YI
Sbjct: 12 IKKILLSYTYVAIWIFLSFTVIVYNKYI 39
>At2g25520 putative phosphate/phosphoenolpyruvate translocator
protein
Length = 347
Score = 29.3 bits (64), Expect = 0.49
Identities = 12/28 (42%), Positives = 20/28 (70%)
Query: 91 ILKIPLTFVEVAVWVILTYYVIGFDPYI 118
I KI L++ VA+W+ L++ VI ++ YI
Sbjct: 12 IKKIILSYTYVAIWIFLSFTVIVYNKYI 39
>At5g35160 unknown protein
Length = 627
Score = 28.5 bits (62), Expect = 0.83
Identities = 15/40 (37%), Positives = 20/40 (49%), Gaps = 5/40 (12%)
Query: 80 FPPW-----AYALPAWILKIPLTFVEVAVWVILTYYVIGF 114
+P W A LP L I L F+ ++W+ YYV GF
Sbjct: 481 YPSWLLVLGAGTLPFGTLFIELFFIMSSIWMGRVYYVFGF 520
>At3g21090 ABC transporter, putative
Length = 680
Score = 28.5 bits (62), Expect = 0.83
Identities = 15/67 (22%), Positives = 27/67 (39%)
Query: 55 FIGVAELSMVVSRLPVFYKQRGYLFFPPWAYALPAWILKIPLTFVEVAVWVILTYYVIGF 114
F+ + + + VFYK+R ++ Y L +I P + +TY ++ F
Sbjct: 424 FMSIGGFPSFLEEMKVFYKERLSGYYGVSVYILSNYISSFPFLVAISVITGTITYNLVKF 483
Query: 115 DPYIGRY 121
P Y
Sbjct: 484 RPGFSHY 490
>At2g39350 ABC transporter like protein
Length = 740
Score = 27.7 bits (60), Expect = 1.4
Identities = 23/115 (20%), Positives = 47/115 (40%), Gaps = 5/115 (4%)
Query: 2 KRNSFVYIFKLCQLAIMAMIAMTIFLRTEMHRDSVAHGDIYVGALFYGCIVILFIGVAEL 61
+R ++ ++ + I I T+F R + V +G + + + L
Sbjct: 448 RRQPELFGIRIASVVITGFILATVFWRLDNSPKGVQER---LGFFAFAMSTMFYTCADAL 504
Query: 62 SMVVSRLPVFYKQRGYLFFPPWAYALPAWILKIP-LTFVEVAVWVILTYYVIGFD 115
+ + +F ++ Y + +Y L I+ P L F+ VA + TY+ +G D
Sbjct: 505 PVFLQERYIFMRETAYNAYRRSSYVLSHAIVSFPSLIFLSVA-FAATTYWAVGLD 558
>At4g12650 putative protein
Length = 527
Score = 27.3 bits (59), Expect = 1.9
Identities = 15/40 (37%), Positives = 20/40 (49%), Gaps = 5/40 (12%)
Query: 80 FPPW-----AYALPAWILKIPLTFVEVAVWVILTYYVIGF 114
+P W A LP L I L F+ ++W+ YYV GF
Sbjct: 381 YPSWLLVLGAGTLPFGTLFIELFFIFSSIWLGRFYYVFGF 420
>At1g51500 ATP-dependent transmembrane transporter, putative
Length = 687
Score = 27.3 bits (59), Expect = 1.9
Identities = 12/67 (17%), Positives = 28/67 (40%)
Query: 55 FIGVAELSMVVSRLPVFYKQRGYLFFPPWAYALPAWILKIPLTFVEVAVWVILTYYVIGF 114
F+ + + + VFYK+R ++ Y + ++ P + +TY ++ F
Sbjct: 425 FMSIGGFPSFIEEMKVFYKERLSGYYGVSVYIISNYVSSFPFLVAIALITGSITYNMVKF 484
Query: 115 DPYIGRY 121
P + +
Sbjct: 485 RPGVSHW 491
>At5g11230 putative protein
Length = 351
Score = 26.9 bits (58), Expect = 2.4
Identities = 11/28 (39%), Positives = 19/28 (67%)
Query: 91 ILKIPLTFVEVAVWVILTYYVIGFDPYI 118
I I L++ VA+W+ L++ VI ++ YI
Sbjct: 12 IKNIVLSYSYVAIWIFLSFTVIVYNKYI 39
>At2g02020 putative peptide/amino acid transporter
Length = 545
Score = 26.9 bits (58), Expect = 2.4
Identities = 17/56 (30%), Positives = 25/56 (44%)
Query: 19 AMIAMTIFLRTEMHRDSVAHGDIYVGALFYGCIVILFIGVAELSMVVSRLPVFYKQ 74
A I T+ L+ D V GDI +F+ +G A + V R+ FY+Q
Sbjct: 409 AAIVETVRLQLARDLDLVESGDIVPLNIFWQIPQYFLMGTAGVFFFVGRIEFFYEQ 464
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.336 0.149 0.486
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,511,909
Number of Sequences: 26719
Number of extensions: 87753
Number of successful extensions: 303
Number of sequences better than 10.0: 33
Number of HSP's better than 10.0 without gapping: 30
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 258
Number of HSP's gapped (non-prelim): 43
length of query: 122
length of database: 11,318,596
effective HSP length: 87
effective length of query: 35
effective length of database: 8,994,043
effective search space: 314791505
effective search space used: 314791505
T: 11
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 53 (25.0 bits)
Medicago: description of AC148607.5


