

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
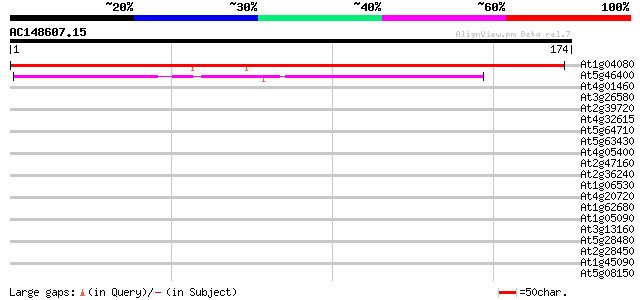
Query= AC148607.15 - phase: 0
(174 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At1g04080 unknown protein 209 8e-55
At5g46400 putative protein 90 7e-19
At4g01460 putative bHLH transcription factor (bHLH057) 39 0.001
At3g26580 unknown protein 32 0.17
At2g39720 unknown protein 30 0.66
At4g32615 putative protein 29 1.1
At5g64710 putative protein 29 1.5
At5g63430 putative protein 28 1.9
At4g05400 putative protein 28 1.9
At2g47160 putative anion exchange protein 28 1.9
At2g36240 putative salt-inducible protein 28 1.9
At1g06530 hypothetical protein 28 1.9
At4g20720 hypothetical protein 28 2.5
At1g62680 PPR-repeat protein 28 2.5
At1g05090 unknown protein 28 2.5
At3g13160 unknown protein 28 3.3
At5g28480 putative protein 27 4.3
At2g28450 RNA methyltransferase like protein 27 4.3
At1g45090 hypothetical protein 27 4.3
At5g08150 putative protein 27 7.3
>At1g04080 unknown protein
Length = 768
Score = 209 bits (531), Expect = 8e-55
Identities = 109/182 (59%), Positives = 131/182 (71%), Gaps = 10/182 (5%)
Query: 1 MQQDWARLAMIYTRILENPNQQLDRYFNSFKELASNRPLSELRTADEAAAVAGVV----- 55
MQQDW+R+A+IYTRILENP Q LDRYF+SFKELA RPLSELR+A+E+AA A V
Sbjct: 220 MQQDWSRVALIYTRILENPIQNLDRYFSSFKELAETRPLSELRSAEESAAAAVAVAGDAS 279
Query: 56 ----SEGIDQGVEGEVHPDGA-DNSPKPASAGLTEAEELEKYIAIREEMYKKAKEFDSKI 110
SE ++ EG DG+ + SPK SA TE EEL+KY+ IRE MY K+KEF+SKI
Sbjct: 280 ESAASESGEKADEGRSQVDGSTEQSPKLESASSTEPEELKKYVGIREAMYIKSKEFESKI 339
Query: 111 IGFETTIRRPYFHVRPLNIGELENWHNYLDFIEREGDLSKVTKLLSYKGTFCSIFSCYRL 170
IG+E IRRPYFHVRPLN+ ELENWHNYLDFIER+GD +KV KL C+ + Y +
Sbjct: 340 IGYEMAIRRPYFHVRPLNVAELENWHNYLDFIERDGDFNKVVKLYERCVVTCANYPEYWI 399
Query: 171 TY 172
Y
Sbjct: 400 RY 401
>At5g46400 putative protein
Length = 1022
Score = 89.7 bits (221), Expect = 7e-19
Identities = 55/154 (35%), Positives = 84/154 (53%), Gaps = 15/154 (9%)
Query: 2 QQDWARLAMIYTRILENPNQQLDRYFNSFKELASNRPLSELRTADEAAAVAGVVSEGIDQ 61
QQ W+ LA +Y R L+ P+++LD Y+ +F+++A++ D V G +S D
Sbjct: 167 QQQWSSLANVYLRTLKYPSKKLDLYYKNFRKIAASLKEKIKCRID----VNGDLSS--DP 220
Query: 62 GVEGEVHPDGADNSPK--------PASAGLTEAEELEKYIAIREEMYKKAKEFDSKIIGF 113
E VH D P+S+ ++ L Y++I E+ Y+ +++ KI F
Sbjct: 221 MEEDLVHTRHTDEEISIVVRELMGPSSSSAV-SKALHTYLSIGEQFYQDSRQLMEKISCF 279
Query: 114 ETTIRRPYFHVRPLNIGELENWHNYLDFIEREGD 147
ET IRRPYFHV+PL+ +L+NWH YL F E GD
Sbjct: 280 ETQIRRPYFHVKPLDTNQLDNWHAYLSFGETYGD 313
>At4g01460 putative bHLH transcription factor (bHLH057)
Length = 315
Score = 39.3 bits (90), Expect = 0.001
Identities = 24/76 (31%), Positives = 38/76 (49%), Gaps = 8/76 (10%)
Query: 10 MIYTRILENPNQQLDRYFNSFKELASNRPLSELRTADEAAAVAGVVS-----EGIDQGVE 64
M + + N +Q++ + NS + L P S L+ D+A+ V G + E + Q +E
Sbjct: 115 MTHIAVERNRRRQMNEHLNSLRSLM---PPSFLQRGDQASIVGGAIDFIKELEQLLQSLE 171
Query: 65 GEVHPDGADNSPKPAS 80
E DG D +PK AS
Sbjct: 172 AEKRKDGTDETPKTAS 187
>At3g26580 unknown protein
Length = 350
Score = 32.0 bits (71), Expect = 0.17
Identities = 24/84 (28%), Positives = 41/84 (48%), Gaps = 7/84 (8%)
Query: 26 YFNSFKELASNRPLSELRTADEAAAVAGVVSEGIDQGVEGEVHPDGADNSPKPAS---AG 82
+ + + L+SN+PL +D A V ++ G V D A +S + +S G
Sbjct: 45 HVSHYAALSSNKPLE----SDGEARVRRFLTRGKANSRANAVDFDDAGSSDEESSDGGGG 100
Query: 83 LTEAEELEKYIAIREEMYKKAKEF 106
+ EE E+Y + +E ++AKEF
Sbjct: 101 GGDEEEEEEYFDVEKERKRRAKEF 124
>At2g39720 unknown protein
Length = 401
Score = 30.0 bits (66), Expect = 0.66
Identities = 18/51 (35%), Positives = 26/51 (50%), Gaps = 1/51 (1%)
Query: 35 SNRPLSELRTADEAAAVAGVVSEGIDQGVEGEVHPDGADNSPKPASAGLTE 85
S P+ LR AA + VVSEG+D+ + DG D+ +P +TE
Sbjct: 93 SPNPVIVLR-GSAAAPSSDVVSEGLDRSAFQMYYDDGTDSGLRPLPPSMTE 142
>At4g32615 putative protein
Length = 315
Score = 29.3 bits (64), Expect = 1.1
Identities = 17/59 (28%), Positives = 30/59 (50%), Gaps = 1/59 (1%)
Query: 55 VSEGIDQGVEGEVHPDGADNSPKPASAGLTEAEELEKYIAIREEMYKKAKEFDSKIIGF 113
V E +QG E +VHP+ A+ K A +E E+ ++ +E K+ E ++ + F
Sbjct: 124 VEEDHEQGTETQVHPE-AEPEVKKAPEVRAPPKEAERQLSKKERKKKELAELEALLADF 181
>At5g64710 putative protein
Length = 841
Score = 28.9 bits (63), Expect = 1.5
Identities = 27/91 (29%), Positives = 37/91 (39%), Gaps = 8/91 (8%)
Query: 15 ILENPNQQLDRYFNSFKELASNRPLSELRTADEAAAVAGVVSEGIDQGVEGEVHPDGADN 74
I ENP+ L F E +S R S RT D A+ +SE + G G+ +
Sbjct: 601 IQENPSDALPFRVTRFTEESSCR--SNPRTTDGLRAIFVNMSESLCDGANGDKNSTNVGM 658
Query: 75 SPKPASAGLTEAEELEKYIAIREEMYKKAKE 105
S KP + K IA ++ KK E
Sbjct: 659 SQKP------KERSRSKVIADCHKLIKKITE 683
>At5g63430 putative protein
Length = 330
Score = 28.5 bits (62), Expect = 1.9
Identities = 24/102 (23%), Positives = 48/102 (46%), Gaps = 9/102 (8%)
Query: 9 AMIYTRILENP----NQQLDRYFNSFKELASNRPLSELRTADEAAAVAGVVSEGIDQGVE 64
A YT ++ P +++ D ++ SF +S+ SE ++ A +EG + +
Sbjct: 158 ASTYTEEVDTPVGSSSEESDDFWKSFINPSSSPSPSETENMNKVADTEPK-AEGKENSRD 216
Query: 65 GEVHPDGADN----SPKPASAGLTEAEELEKYIAIREEMYKK 102
+ D +D+ SPK + EE++K I +R E++ +
Sbjct: 217 DDELADASDSETKSSPKRVRKNKWKPEEIKKVIRMRGELHSR 258
>At4g05400 putative protein
Length = 250
Score = 28.5 bits (62), Expect = 1.9
Identities = 18/40 (45%), Positives = 24/40 (60%), Gaps = 1/40 (2%)
Query: 80 SAGLTEAEELEKYIAIREEMYKKAKEFDSKIIGFETTIRR 119
S G+ EAEEL KY E+ + AKE+ SK++ E RR
Sbjct: 145 SLGIFEAEELPKYKLTEEDGERLAKEY-SKVLMREHRERR 183
>At2g47160 putative anion exchange protein
Length = 704
Score = 28.5 bits (62), Expect = 1.9
Identities = 20/71 (28%), Positives = 32/71 (44%)
Query: 12 YTRILENPNQQLDRYFNSFKELASNRPLSELRTADEAAAVAGVVSEGIDQGVEGEVHPDG 71
Y LE ++ + R F+ +S + S T +++ V S + G++ P
Sbjct: 624 YPGDLEILDEVMTRSRGEFRHTSSPKVTSSSSTPVNNRSLSQVFSPRVSGIRLGQMSPRV 683
Query: 72 ADNSPKPASAG 82
NSPKPAS G
Sbjct: 684 VGNSPKPASCG 694
>At2g36240 putative salt-inducible protein
Length = 497
Score = 28.5 bits (62), Expect = 1.9
Identities = 13/44 (29%), Positives = 23/44 (51%)
Query: 75 SPKPASAGLTEAEELEKYIAIREEMYKKAKEFDSKIIGFETTIR 118
+P P S+G+ ELE + Y +A++ D ++ F+T R
Sbjct: 142 NPCPCSSGIFSCPELEPIFRSAIDAYCRARKMDYALLAFDTMKR 185
>At1g06530 hypothetical protein
Length = 323
Score = 28.5 bits (62), Expect = 1.9
Identities = 24/76 (31%), Positives = 34/76 (44%), Gaps = 4/76 (5%)
Query: 43 RTADEAAAVAGVVSEGIDQGVEGEVHPDGADNSPKPASAGLTEAEELEKYIA----IREE 98
R ++ VA + E I EGE A+ S EELEK +A ++EE
Sbjct: 103 RASELETEVARLQHELITARTEGEEATAEAEKLRSEISQKGGGIEELEKEVAGLRTVKEE 162
Query: 99 MYKKAKEFDSKIIGFE 114
K+ KE +SK+ E
Sbjct: 163 NEKRMKELESKLGALE 178
>At4g20720 hypothetical protein
Length = 758
Score = 28.1 bits (61), Expect = 2.5
Identities = 27/96 (28%), Positives = 40/96 (41%), Gaps = 18/96 (18%)
Query: 56 SEGIDQGVEGEVHPDGADNSPKPASAGLTEAEELEKYIAIREEMYKKAKEFDSKIIGFET 115
SE ++ E + GA +P L++ +LE + EE + K
Sbjct: 138 SEMVEPLKETDGSSRGATKAPPSKGIALSKFLDLEIQWSALEEKSDDGQSVQKK------ 191
Query: 116 TIRRPYFHVRPLNIGELENWHNYLDFIEREGDLSKV 151
PLN+G + N +Y F+ER GDLSKV
Sbjct: 192 ---------NPLNLGGI-NLDDY--FVERRGDLSKV 215
>At1g62680 PPR-repeat protein
Length = 542
Score = 28.1 bits (61), Expect = 2.5
Identities = 16/37 (43%), Positives = 22/37 (59%), Gaps = 2/37 (5%)
Query: 82 GLTEAEELEKYIAIREEMYKKAKEFDSKIIGFETTIR 118
GL + ELEK + I E+M K +E D I+ + T IR
Sbjct: 403 GLCDNGELEKALVIFEDMQK--REMDLDIVTYTTVIR 437
>At1g05090 unknown protein
Length = 706
Score = 28.1 bits (61), Expect = 2.5
Identities = 27/96 (28%), Positives = 40/96 (41%), Gaps = 18/96 (18%)
Query: 56 SEGIDQGVEGEVHPDGADNSPKPASAGLTEAEELEKYIAIREEMYKKAKEFDSKIIGFET 115
SE ++ E + GA +P L++ +LE + EE + K
Sbjct: 109 SEMVEPLKETDGSSRGATKAPPSKGIALSKFLDLEIQWSALEEKSDDGQSVQKK------ 162
Query: 116 TIRRPYFHVRPLNIGELENWHNYLDFIEREGDLSKV 151
PLN+G + N +Y F+ER GDLSKV
Sbjct: 163 ---------NPLNLGGI-NLDDY--FVERRGDLSKV 186
>At3g13160 unknown protein
Length = 394
Score = 27.7 bits (60), Expect = 3.3
Identities = 21/86 (24%), Positives = 38/86 (43%)
Query: 76 PKPASAGLTEAEELEKYIAIREEMYKKAKEFDSKIIGFETTIRRPYFHVRPLNIGELENW 135
PKP+ L E K+I + + +A+ F I +E T+RR + + E+
Sbjct: 37 PKPSLITLVNDERDPKFITEKFKKACQAEWFRKNIAVYERTVRRLAAAKKFEWVEEILEE 96
Query: 136 HNYLDFIEREGDLSKVTKLLSYKGTF 161
N + +EG ++++ L G F
Sbjct: 97 QNKYPNMSKEGFVARIINLYGRVGMF 122
>At5g28480 putative protein
Length = 1248
Score = 27.3 bits (59), Expect = 4.3
Identities = 21/75 (28%), Positives = 30/75 (40%), Gaps = 2/75 (2%)
Query: 46 DEAAAVAGVVSEGIDQGVEGEVHPDGADNSPKPASA-GLTEAEELEKYIAIREEMYKKAK 104
D +G EG G +GE P+GAD P A G E +KY +K+
Sbjct: 444 DGEGGPSGGDGEGGPSGGDGEGGPNGADGEGGPNGADGEVGDEAFDKYAGELRSFFKRQT 503
Query: 105 E-FDSKIIGFETTIR 118
+ D K + +R
Sbjct: 504 QLLDKKFEDLKVFVR 518
>At2g28450 RNA methyltransferase like protein
Length = 809
Score = 27.3 bits (59), Expect = 4.3
Identities = 11/31 (35%), Positives = 21/31 (67%)
Query: 85 EAEELEKYIAIREEMYKKAKEFDSKIIGFET 115
+++EL+K++ + +YK AK+ I+GF T
Sbjct: 182 QSDELKKFLGEQGVLYKSAKKRRGMIVGFVT 212
>At1g45090 hypothetical protein
Length = 1210
Score = 27.3 bits (59), Expect = 4.3
Identities = 21/75 (28%), Positives = 30/75 (40%), Gaps = 2/75 (2%)
Query: 46 DEAAAVAGVVSEGIDQGVEGEVHPDGADNSPKPASA-GLTEAEELEKYIAIREEMYKKAK 104
D +G EG G +GE P+GAD P A G E +KY +K+
Sbjct: 438 DGEGGPSGGDGEGGPSGGDGEGGPNGADGEGGPNGADGEVGDEAFDKYAGELRSFFKRQT 497
Query: 105 E-FDSKIIGFETTIR 118
+ D K + +R
Sbjct: 498 QLLDKKFEDLKVFVR 512
>At5g08150 putative protein
Length = 144
Score = 26.6 bits (57), Expect = 7.3
Identities = 18/56 (32%), Positives = 29/56 (51%), Gaps = 3/56 (5%)
Query: 64 EGEVHPDGADNSPKPASAGLTEAE---ELEKYIAIREEMYKKAKEFDSKIIGFETT 116
E E DG D+ AS+G EA +L + I + + KK K+ + +++ ETT
Sbjct: 42 EEEEDSDGGDSMDSDASSGPMEATSYLKLAQEIEEQNSIKKKKKKTNEEMVLVETT 97
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.316 0.135 0.390
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,922,649
Number of Sequences: 26719
Number of extensions: 163291
Number of successful extensions: 459
Number of sequences better than 10.0: 25
Number of HSP's better than 10.0 without gapping: 9
Number of HSP's successfully gapped in prelim test: 17
Number of HSP's that attempted gapping in prelim test: 439
Number of HSP's gapped (non-prelim): 32
length of query: 174
length of database: 11,318,596
effective HSP length: 92
effective length of query: 82
effective length of database: 8,860,448
effective search space: 726556736
effective search space used: 726556736
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 56 (26.2 bits)
Medicago: description of AC148607.15


