

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148607.14 - phase: 0
(359 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
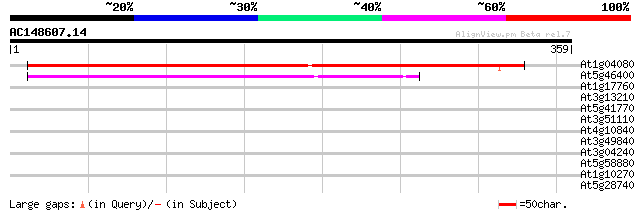
Score E
Sequences producing significant alignments: (bits) Value
At1g04080 unknown protein 424 e-119
At5g46400 putative protein 175 4e-44
At1g17760 unknown protein 39 0.004
At3g13210 crn-like protein 37 0.013
At5g41770 cell cycle control crn (crooked neck) protein-like 37 0.017
At3g51110 crooked neck-like protein 34 0.11
At4g10840 unknown protein 32 0.70
At3g49840 putative protein 30 2.7
At3g04240 O-linked GlcNAc transferase like protein 30 2.7
At5g58880 putative protein 29 3.5
At1g10270 unknown protein 29 4.5
At5g28740 putative protein 28 7.8
>At1g04080 unknown protein
Length = 768
Score = 424 bits (1091), Expect = e-119
Identities = 212/321 (66%), Positives = 254/321 (79%), Gaps = 5/321 (1%)
Query: 12 DQIVKLYERCVIACANYPEYWIRYVLCMEASESMDLANNVLARASQVFVKRQPEIHLFCA 71
+++VKLYERCV+ CANYPEYWIRYV MEAS S DLA N LARA+QVFVK+QPEIHLF A
Sbjct: 378 NKVVKLYERCVVTCANYPEYWIRYVTNMEASGSADLAENALARATQVFVKKQPEIHLFAA 437
Query: 72 RFKEQAGDIVGARAAYQLVHTEISPGLLEAIIRHANMEHRLGKLEDAFSLYEQAIAIEKG 131
R KEQ GDI GARAAYQLVH+EISPGLLEA+I+HANME+RLG L+DAFSLYEQ IA+EKG
Sbjct: 438 RLKEQNGDIAGARAAYQLVHSEISPGLLEAVIKHANMEYRLGNLDDAFSLYEQVIAVEKG 497
Query: 132 KEHSQTLPMLFAQYSRFVYLASGNSEKAREILVGGLENASLSKPLLEALLHFEAIQPQPK 191
KEHS LP+L+AQYSRF YL S ++EKAR I+V L++ SKPL+EAL+HFEAIQP P+
Sbjct: 498 KEHSTILPLLYAQYSRFSYLVSRDAEKARRIIVEALDHVQPSKPLMEALIHFEAIQPPPR 557
Query: 192 RVDIDFLESLVVKFITPNPENPGVASATEREELSNIFLEFLNLFGDVQSIKRAEDRHAKL 251
+ID+LE LV K I P+ + +AS+TEREELS I++EFL +FGDV+SIK+AED+H KL
Sbjct: 558 --EIDYLEPLVEKVIKPDADAQNIASSTEREELSLIYIEFLGIFGDVKSIKKAEDQHVKL 615
Query: 252 FLPNRGLSELKKRHAEDFLASDKTKVSRAYSAQSPAQSVAGAYPNGPNQWP-NYGVQPQT 310
F P+R SELKKR A+DFLASD+TK+++ Y+ PAQ V+ AYPN QW Y QPQT
Sbjct: 616 FYPHRSTSELKKRSADDFLASDRTKMAKTYNGTPPAQPVSNAYPNAQAQWSGGYAAQPQT 675
Query: 311 WP--ATTQAQGQQWPAGYTQQ 329
WP AQ QQW Y QQ
Sbjct: 676 WPPAQAAPAQPQQWNPAYGQQ 696
>At5g46400 putative protein
Length = 1022
Score = 175 bits (443), Expect = 4e-44
Identities = 98/252 (38%), Positives = 150/252 (58%), Gaps = 4/252 (1%)
Query: 12 DQIVKLYERCVIACANYPEYWIRYVLCMEASESMDLANNVLARASQVFVKRQPEIHLFCA 71
D + LYERC+I CANY E+W RYV +E+ +LAN LARASQ FVK IHLF A
Sbjct: 315 DWAINLYERCLIPCANYTEFWFRYVDFVESKGGRELANFALARASQTFVKSASVIHLFNA 374
Query: 72 RFKEQAGDIVGARAAYQLVHTEISPGLLEAIIRHANMEHRLGKLEDAFSLYEQAI-AIEK 130
RFKE GD A A E+ G +E + + ANME RLG E A + Y +A+
Sbjct: 375 RFKEHVGDASAASVALSRCGEELGFGFVENVTKKANMEKRLGNFEAAVTTYREALNKTLI 434
Query: 131 GKEHSQTLPMLFAQYSRFVYLASGNSEKAREILVGGLENASLSKPLLEALLHFEAIQPQP 190
GKE+ +T L+ Q+SR Y+ + +++ A +IL+ G EN K LLE L+ +
Sbjct: 435 GKENLETTARLYVQFSRLKYVITNSADDAAQILLEGNENVPHCKLLLEELMRLLMMHGGS 494
Query: 191 KRVDIDFLESLVVKFITPNPENPGVASATEREELSNIFLEFLNLFGDVQSIKRAEDRHAK 250
++VD+ L+ ++ K ++ ++ SA ++EE+SN+++EF++L G + +++A RH K
Sbjct: 495 RQVDL--LDPIIDKELSHQADSSDGLSAEDKEEISNLYMEFIDLSGTIHDVRKALGRHIK 552
Query: 251 LFLPNRGLSELK 262
LF P+ ++L+
Sbjct: 553 LF-PHSARAKLR 563
>At1g17760 unknown protein
Length = 734
Score = 38.9 bits (89), Expect = 0.004
Identities = 57/258 (22%), Positives = 91/258 (35%), Gaps = 67/258 (25%)
Query: 13 QIVKLYERCVIACANYPEYWIRYVLCMEASESMDLANNVLARASQVFVKRQPEIHLFCAR 72
+I+ YE+C++ +YP+ W Y S S D A V RA +K P+ +
Sbjct: 250 RIIYAYEQCLMCLYHYPDVWYDYAEWHVKSGSTDAAIKVFQRA----LKAIPDSEMLKYA 305
Query: 73 FKEQAGDIVGARAAYQLVHTEISPGLLEAIIRHANMEHRLGKLEDAFSLYEQAIAIEKGK 132
F A ME G ++ A LYE +
Sbjct: 306 F--------------------------------AEMEESRGAIQSAKKLYENILG----- 328
Query: 133 EHSQTLPMLFAQYSRFVYLASGNSEKAREILVGGLENASLSKPLLEALLHFE-AIQPQPK 191
+ T + QY RF+ A G E AR+ + ++ S + + A I +PK
Sbjct: 329 --ASTNSLAHIQYLRFLRRAEG-VEAARKYFLDARKSPSCTYHVYIAFATMAFCIDKEPK 385
Query: 192 RVDIDFLESL------------VVKFITPNPENPGVASATER----------EELSNIFL 229
F E L F+T ++ + + ER E+ F+
Sbjct: 386 VAHNIFEEGLKLYMSEPVYILKYADFLTRLNDDRNIRALFERALSTLPVEDSAEVWKRFI 445
Query: 230 EFLNLFGDVQSIKRAEDR 247
+F +GD+ SI + E R
Sbjct: 446 QFEQTYGDLASILKVEQR 463
>At3g13210 crn-like protein
Length = 657
Score = 37.4 bits (85), Expect = 0.013
Identities = 34/153 (22%), Positives = 68/153 (44%), Gaps = 10/153 (6%)
Query: 11 VDQIVKLYERCVIACANYPEYWIRYVLCMEASESMDLANNVLARASQVFVKRQPEIHLFC 70
V++ +++R V + W +++ E ++ A +L R + + L
Sbjct: 107 VNEARNVWDRAVSLLPRVDQLWYKFIHMEEKLGNIAGARQILER--WIHCSPDQQAWLCF 164
Query: 71 ARFKEQAGDIVGARAAYQ---LVHTEISPGLLEAIIRHANMEHRLGKLEDAFSLYEQAIA 127
+F+ + +I AR+ Y+ L H ++S A IR+A E + G++E A ++E+A
Sbjct: 165 IKFELKYNEIECARSIYERFVLCHPKVS-----AYIRYAKFEMKHGQVELAMKVFERAKK 219
Query: 128 IEKGKEHSQTLPMLFAQYSRFVYLASGNSEKAR 160
E ++ L + FA++ A K R
Sbjct: 220 ELADDEEAEILFVAFAEFEEQYKFALDQIPKGR 252
>At5g41770 cell cycle control crn (crooked neck) protein-like
Length = 705
Score = 37.0 bits (84), Expect = 0.017
Identities = 35/153 (22%), Positives = 67/153 (42%), Gaps = 19/153 (12%)
Query: 10 FVDQIVKLYERCVIACANYPEYWIRYVLCMEASESMDLANNVLARASQVFVKRQPEIHLF 69
FV+ +++R V + W +Y+ E ++ A + R +Q +
Sbjct: 141 FVNSARNVWDRAVTLLPRVDQLWYKYIHMEEILGNIAGARQIFERWMDWSPDQQGWLSFI 200
Query: 70 CARFKEQAGDIVGARAAYQ---LVHTEISPGLLEAIIRHANMEHRLGKLEDAFSLYEQAI 126
+F+ + +I AR Y+ L H ++S A IR+A E + G++ S+YE+A
Sbjct: 201 --KFELRYNEIERARTIYERFVLCHPKVS-----AYIRYAKFEMKGGEVARCRSVYERAT 253
Query: 127 AIEKGKEHSQTLPMLFAQY---------SRFVY 150
E ++ L + FA++ +RF+Y
Sbjct: 254 EKLADDEEAEILFVAFAEFEERCKEVERARFIY 286
Score = 30.0 bits (66), Expect = 2.0
Identities = 37/160 (23%), Positives = 66/160 (41%), Gaps = 14/160 (8%)
Query: 17 LYERCVIACANYPEYWIRYVLCMEASE---SMDLANNVLARASQVFVK-RQPEIHLFCAR 72
+YER A+ E I +V E E ++ A + A K R +++
Sbjct: 248 VYERATEKLADDEEAEILFVAFAEFEERCKEVERARFIYKFALDHIPKGRAEDLYRKFVA 307
Query: 73 FKEQAGD-------IVGARAAYQLVHTEISPGLLEAIIRHANMEHRLGKLEDAFSLYEQA 125
F++Q GD IVG R SP ++ + +E +G + +YE+A
Sbjct: 308 FEKQYGDKEGIEDAIVGKRRFQYEDEVRKSPSNYDSWFDYVRLEESVGNKDRIREIYERA 367
Query: 126 IA---IEKGKEHSQTLPMLFAQYSRFVYLASGNSEKAREI 162
IA + K + Q L+ Y+ F + + + E+ R++
Sbjct: 368 IANVPPAEEKRYWQRYIYLWINYALFEEIETEDIERTRDV 407
>At3g51110 crooked neck-like protein
Length = 599
Score = 34.3 bits (77), Expect = 0.11
Identities = 36/153 (23%), Positives = 69/153 (44%), Gaps = 13/153 (8%)
Query: 11 VDQIVKLYERCVIACANYPEYWIRYVLCMEASESMDLANNVLARASQVFVKRQPEIHLFC 70
V+ +++R V ++W +Y+ E ++D A + R +Q L
Sbjct: 116 VNHARNVWDRAVKILPRVDQFWYKYIHMEEILGNIDGARKIFERWMDWSPDQQ--AWLCF 173
Query: 71 ARFKEQAGDIVGARAAYQ---LVHTEISPGLLEAIIRHANMEHRLGKLEDAFSLYEQAIA 127
+F+ + +I +R+ Y+ L H + S + IR+A E + ++ A +YE+ A
Sbjct: 174 IKFELRYNEIERSRSIYERFVLCHPKAS-----SFIRYAKFEMKNSQVSLARIVYER--A 226
Query: 128 IEKGKEHSQTLPMLFAQYSRFVYLASGNSEKAR 160
IE K+ + M+F ++ F L E+AR
Sbjct: 227 IEMLKDVEEEAEMIFVAFAEFEELCK-EVERAR 258
Score = 32.0 bits (71), Expect = 0.54
Identities = 25/109 (22%), Positives = 49/109 (44%), Gaps = 1/109 (0%)
Query: 16 KLYERCVIACANYPEYWIRYVLCMEASESMDLANNVLARASQVFVKRQPEIHLFCARFKE 75
K +E + + W+RY E+ + D A +V RA + R + L A F+
Sbjct: 52 KEFEDQIRGAKTNSQVWVRYADWEESQKDHDRARSVWERALEDESYRNHTLWLKYAEFEM 111
Query: 76 QAGDIVGARAAYQLVHTEISPGLLEAIIRHANMEHRLGKLEDAFSLYEQ 124
+ + AR + +I P + + ++ +ME LG ++ A ++E+
Sbjct: 112 RNKSVNHARNVWDRA-VKILPRVDQFWYKYIHMEEILGNIDGARKIFER 159
>At4g10840 unknown protein
Length = 609
Score = 31.6 bits (70), Expect = 0.70
Identities = 18/63 (28%), Positives = 28/63 (43%)
Query: 66 IHLFCARFKEQAGDIVGARAAYQLVHTEISPGLLEAIIRHANMEHRLGKLEDAFSLYEQA 125
I++ RF E A ++ E P + +R A + HR GKL ++ S E A
Sbjct: 316 IYMSLCRFDEAVFSYQKALTVFKASKGETHPTVASVFVRLAELYHRTGKLRESKSYCENA 375
Query: 126 IAI 128
+ I
Sbjct: 376 LRI 378
>At3g49840 putative protein
Length = 651
Score = 29.6 bits (65), Expect = 2.7
Identities = 26/89 (29%), Positives = 33/89 (36%), Gaps = 22/89 (24%)
Query: 264 RHAEDFLASDK--TKVSRAYSAQSPAQSVAGAYPNG-------------------PNQWP 302
R+ E SDK T+ + S Q P V+ P G P Q+P
Sbjct: 509 RYKERNKPSDKSITEKKKKMSYQDPQHPVSAPPPQGYPPKEGYPPAGYPPPAGYPPPQYP 568
Query: 303 NYGVQPQTWPATTQAQGQQWPA-GYTQQQ 330
G P +P Q GQ +PA GY Q
Sbjct: 569 QAGYPPAGYPPPQQGYGQGYPAQGYPPPQ 597
>At3g04240 O-linked GlcNAc transferase like protein
Length = 977
Score = 29.6 bits (65), Expect = 2.7
Identities = 28/122 (22%), Positives = 50/122 (40%), Gaps = 14/122 (11%)
Query: 11 VDQIVKLYERCVIACANYPEYWIRYVLCMEASESMDLANNVLARASQVFVKRQPEIHLFC 70
VD+ V+ Y +C+ N+P+ M ++ + N++ AS +F
Sbjct: 341 VDEAVRCYNQCLALQPNHPQ-------AMANLGNIYMEWNMMGPASSLFKATLAVTTGLS 393
Query: 71 ARFK------EQAGDIVGARAAYQLVHTEISPGLLEAIIRHANMEHRLGKLEDAFSLYEQ 124
A F +Q G+ A + Y V I P +A++ N +G++ +A Y
Sbjct: 394 APFNNLAIIYKQQGNYSDAISCYNEV-LRIDPLAADALVNRGNTYKEIGRVTEAIQDYMH 452
Query: 125 AI 126
AI
Sbjct: 453 AI 454
>At5g58880 putative protein
Length = 1088
Score = 29.3 bits (64), Expect = 3.5
Identities = 20/88 (22%), Positives = 48/88 (53%), Gaps = 7/88 (7%)
Query: 93 EISPGLLEAIIRHANMEHRLGKLEDAFSLYEQAIAIEKGKEHSQTLPMLFAQYSRFVYLA 152
++S LLE+++R N + + + E+A+ +E+ K H+ +++ V L
Sbjct: 776 QVSTELLESVVREENGQELVKSAD------EKAMLVEEEKTHNVLEASSSNAHTQLVDLD 829
Query: 153 SGNSEKAREILVGGLENASLSKPLLEAL 180
GN+E + ++++ ++++ S PL E++
Sbjct: 830 YGNAENSSDVILLQVQDSHKS-PLDESV 856
>At1g10270 unknown protein
Length = 913
Score = 28.9 bits (63), Expect = 4.5
Identities = 17/47 (36%), Positives = 18/47 (38%), Gaps = 1/47 (2%)
Query: 288 QSVAGAYPNGPNQWPNYGV-QPQTWPATTQAQGQQWPAGYTQQQVNW 333
QS A P+ W N Q Q W T Q QQW QQ W
Sbjct: 802 QSWANQTPSQQQPWANQTTGQQQGWGNQTTGQQQQWANQTAGQQSGW 848
>At5g28740 putative protein
Length = 917
Score = 28.1 bits (61), Expect = 7.8
Identities = 18/74 (24%), Positives = 34/74 (45%), Gaps = 1/74 (1%)
Query: 57 QVFVKRQPEIHLFCARFKEQAGDIVGARAAYQLVHTEISPGLLEAIIRHANMEHRLGKLE 116
Q+ + R + F +E G + RA Y+ + ++ + I+ +A + E
Sbjct: 508 QMKLHRSLRLWSFYVDLEESLGTLESTRAVYEKI-LDLRIATPQIIMNYAFLLEENKYFE 566
Query: 117 DAFSLYEQAIAIEK 130
DAF +YE+ + I K
Sbjct: 567 DAFKVYERGVKIFK 580
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.321 0.136 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,659,176
Number of Sequences: 26719
Number of extensions: 311489
Number of successful extensions: 855
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 831
Number of HSP's gapped (non-prelim): 23
length of query: 359
length of database: 11,318,596
effective HSP length: 100
effective length of query: 259
effective length of database: 8,646,696
effective search space: 2239494264
effective search space used: 2239494264
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 61 (28.1 bits)
Medicago: description of AC148607.14


