

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148532.6 + phase: 0
(243 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
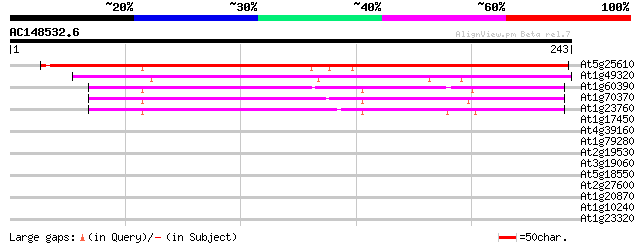
Score E
Sequences producing significant alignments: (bits) Value
At5g25610 dehydration-induced protein RD22 244 4e-65
At1g49320 unknown protein 150 5e-37
At1g60390 132 1e-31
At1g70370 unknown protein 131 4e-31
At1g23760 putative polygalacuronase isoenzyme 1 beta subunit 121 4e-28
At1g17450 hypothetical protein 31 0.52
At4g39160 putative protein 30 1.5
At1g79280 hypothetical protein 29 2.0
At2g19530 unknown protein 28 3.4
At3g19060 hypothetical protein 28 4.4
At5g18550 zinc finger -like protein 28 5.8
At2g27600 putative AAA-type ATPase 27 7.5
At1g20870 unknown protein 27 7.5
At1g10240 unknown protein 27 7.5
At1g23320 putative alliinase 27 9.8
>At5g25610 dehydration-induced protein RD22
Length = 392
Score = 244 bits (622), Expect = 4e-65
Identities = 128/237 (54%), Positives = 157/237 (66%), Gaps = 9/237 (3%)
Query: 14 FIYWPYDASVTQLHDYQNLDLFFFEKDLHNGTKLNMQFTKTGA-----TFLPREVANSIP 68
F+Y Y A TQLHD N LFF EKDL G ++N++F FLPR A ++P
Sbjct: 156 FVY-NYAAKETQLHDDPNAALFFLEKDLVRGKEMNVRFNAEDGYGGKTAFLPRGEAETVP 214
Query: 69 FSSNKVENILNYFSIKQGSAESENLKQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKLG 128
F S K L FS++ GS E+E +K+ I CE + GEEK C TSLESMVDF+ SKLG
Sbjct: 215 FGSEKFSETLKRFSVEAGSEEAEMMKKTIEECEARKVSGEEKYCATSLESMVDFSVSKLG 274
Query: 129 N-NVEAVSTE-ANKESDKQQY-IIAKGVKKLSENKIVVCHLLSYPYAVFYCHKLRATKVY 185
+V AVSTE A K + Q+Y I A GVKKLS++K VVCH YP+AVFYCHK T VY
Sbjct: 275 KYHVRAVSTEVAKKNAPMQKYKIAAAGVKKLSDDKSVVCHKQKYPFAVFYCHKAMMTTVY 334
Query: 186 YLPMEGIDGTKVKTAAICHNDTSQWSPKHLAFHVLKVQPGTVPVCHFLQHGHVIWFS 242
+P+EG +G + K A+CH +TS W+P HLAF VLKV+PGTVPVCHFL HV+WFS
Sbjct: 335 AVPLEGENGMRAKAVAVCHKNTSAWNPNHLAFKVLKVKPGTVPVCHFLPETHVVWFS 391
>At1g49320 unknown protein
Length = 280
Score = 150 bits (380), Expect = 5e-37
Identities = 82/230 (35%), Positives = 131/230 (56%), Gaps = 14/230 (6%)
Query: 28 DYQNLDLFFFEKDLHNGTKLNMQFTKTGATFLP----REVANSIPFSSNKVENILNYFSI 83
D +L ++F DL GTKL + F K LP R+ A+ IPF+ +K++ +L++FSI
Sbjct: 51 DDPSLYMYFTLNDLKLGTKLLIYFYKNDLQKLPPLLTRQQADLIPFTKSKLDFLLDHFSI 110
Query: 84 KQGSAESENLKQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKLGNNVE--------AVS 135
+ S + + +K+ +G+C+ IEGE K C TSLES++D +G NV+ V
Sbjct: 111 TKDSPQGKAIKETLGHCDAKAIEGEHKFCGTSLESLIDLVKKTMGYNVDLKVMTTKVMVP 170
Query: 136 TEANKESDKQQYIIAKGVKKLSENKIVVCHLLSYPYAVFYCHKLR-ATKVYYLPMEGIDG 194
+ + Y + K+L K++ CH + YPYAV+YCH + ++V+ + + DG
Sbjct: 171 AQNSISYALHNYTFVEAPKELVGIKMLGCHRMPYPYAVYYCHGHKGGSRVFEVNLVTDDG 230
Query: 195 -TKVKTAAICHNDTSQWSPKHLAFHVLKVQPGTVPVCHFLQHGHVIWFSK 243
+V A+CH DTS W H+AF VLK++P + PVCHF +++W +K
Sbjct: 231 RQRVVGPAVCHMDTSTWDADHVAFKVLKMEPRSAPVCHFFPLDNIVWVTK 280
>At1g60390
Length = 624
Score = 132 bits (333), Expect = 1e-31
Identities = 78/215 (36%), Positives = 107/215 (49%), Gaps = 12/215 (5%)
Query: 35 FFFEKDLHNGTKLNMQFTKTGA---TFLPREVANSIPFSSNKVENILNYFSIKQGSAESE 91
FF E L GT + M K TFLPR + ++PFSS+ + I F + S+ +
Sbjct: 409 FFREAMLKEGTLMQMPDIKDKMPKRTFLPRNIVKNLPFSSSTIGEIWRVFGAGENSSMAG 468
Query: 92 NLKQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKLGNNVEAVSTEANKESDKQQYIIAK 151
+ A+ CE P GE K CV S E M+DF TS LG V V T N K++ +I K
Sbjct: 469 IISSAVSECERPASHGETKRCVGSAEDMIDFATSVLGRGV-VVRTTENVVGSKKKVVIGK 527
Query: 152 --GVKKLSENKIVVCHLLSYPYAVFYCHKLRATKVYYLPMEGIDGTKVKT----AAICHN 205
G+ + V CH YPY ++YCH + +VY + +D ++ AICH
Sbjct: 528 VNGINGGDVTRAVSCHQSLYPYLLYYCHSVPRVRVYETDL--LDPKSLEKINHGVAICHI 585
Query: 206 DTSQWSPKHLAFHVLKVQPGTVPVCHFLQHGHVIW 240
DTS WSP H AF L PG + VCH++ + W
Sbjct: 586 DTSAWSPSHGAFLALGSGPGQIEVCHWIFENDMTW 620
>At1g70370 unknown protein
Length = 626
Score = 131 bits (329), Expect = 4e-31
Identities = 77/213 (36%), Positives = 106/213 (49%), Gaps = 8/213 (3%)
Query: 35 FFFEKDLHNGTKLNMQFTKTGA---TFLPREVANSIPFSSNKVENILNYFSIKQGSAESE 91
FF E L GT + M K +FLPR + +PFS++K+ I F + S
Sbjct: 411 FFRESSLKEGTVIPMPDIKDKMPKRSFLPRSIITKLPFSTSKLGEIKRIFHAVENSTMGG 470
Query: 92 NLKQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKLGNNVEAVSTEANKESDKQQYIIAK 151
+ A+ CE P GE K CV S E M+DF TS LG +V +TE N K++ +I K
Sbjct: 471 IITDAVTECERPPSVGETKRCVGSAEDMIDFATSVLGRSVVLRTTE-NVAGSKEKVVIGK 529
Query: 152 --GVKKLSENKIVVCHLLSYPYAVFYCHKLRATKVYYLPMEGIDGTKV--KTAAICHNDT 207
G+ K V CH YPY ++YCH + +VY + ++ K AICH DT
Sbjct: 530 VNGINGGKLTKAVSCHQSLYPYLLYYCHSVPKVRVYEADLLELNSKKKINHGIAICHMDT 589
Query: 208 SQWSPKHLAFHVLKVQPGTVPVCHFLQHGHVIW 240
S W P H AF L +PG + VCH++ + W
Sbjct: 590 SSWGPSHGAFLALGSKPGRIEVCHWIFENDMNW 622
>At1g23760 putative polygalacuronase isoenzyme 1 beta subunit
Length = 622
Score = 121 bits (303), Expect = 4e-28
Identities = 75/213 (35%), Positives = 106/213 (49%), Gaps = 8/213 (3%)
Query: 35 FFFEKDLHNGTKLNMQFTKTGA---TFLPREVANSIPFSSNKVENILNYFSIKQGSAESE 91
FF E L GT + M K +FLPR + + +PFS++K+ I F S
Sbjct: 407 FFRESMLKEGTLIWMPDIKDKMPKRSFLPRSIVSKLPFSTSKIAEIKRVFHANDNSTMEG 466
Query: 92 NLKQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKLGNNVEAVSTEANKESDKQQYIIAK 151
+ A+ CE P E K CV S E M+DF TS LG +V +TE+ S K++ +I K
Sbjct: 467 IITDAVRECERPPTVSETKRCVGSAEDMIDFATSVLGRSVVLRTTESVAGS-KEKVMIGK 525
Query: 152 --GVKKLSENKIVVCHLLSYPYAVFYCHKLRATKVYYLP-MEGIDGTKVKTA-AICHNDT 207
G+ K V CH YPY ++YCH + +VY ++ K+ AICH DT
Sbjct: 526 VNGINGGRVTKSVSCHQSLYPYLLYYCHSVPKVRVYESDLLDPKSKAKINHGIAICHMDT 585
Query: 208 SQWSPKHLAFHVLKVQPGTVPVCHFLQHGHVIW 240
S W H AF +L +PG + VCH++ + W
Sbjct: 586 SAWGANHGAFMLLGSRPGQIEVCHWIFENDMNW 618
>At1g17450 hypothetical protein
Length = 1783
Score = 31.2 bits (69), Expect = 0.52
Identities = 33/145 (22%), Positives = 60/145 (40%), Gaps = 12/145 (8%)
Query: 15 IYWPYDASVTQLHDYQNLDLFFFEKDLHNGTKLNM---QFTKTGATFLPREVANSIPFSS 71
+Y P DAS+ QL + LDL F + G + + Q T + + R V + +
Sbjct: 60 VYEPSDASIQQLEEALRLDLRIFANEKLRGNFVGLYDAQSNNTTISAIQRRVLERLAVA- 118
Query: 72 NKVENILNYFSIKQGSAESENLKQAIGYCETPTIEGEEKSCVTSLESMVD-----FTTSK 126
+ + K+ E N + + E+ + ++ + V + E VD TTS
Sbjct: 119 -RANGVAQNLLAKEFGIEGRNFFYIVKHLESRGLVVKQPAIVRTKE--VDGEGDSKTTSC 175
Query: 127 LGNNVEAVSTEANKESDKQQYIIAK 151
+ N+ +S A +Q++ I K
Sbjct: 176 ISTNMIYLSRYAKPLGSQQRFEICK 200
>At4g39160 putative protein
Length = 545
Score = 29.6 bits (65), Expect = 1.5
Identities = 15/54 (27%), Positives = 26/54 (47%)
Query: 91 ENLKQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKLGNNVEAVSTEANKESDK 144
E +Q + C T+ GEE++C+ + SK G + A S + K S++
Sbjct: 213 EQEEQGVSPCINNTVTGEEENCMGNTVEEQSKRESKTGKSKRATSRKRKKTSEE 266
>At1g79280 hypothetical protein
Length = 2111
Score = 29.3 bits (64), Expect = 2.0
Identities = 26/76 (34%), Positives = 40/76 (52%), Gaps = 9/76 (11%)
Query: 82 SIKQGSAESENL----KQAIGYCETP-TIEGEEKSCVTSLESMVDFTTSKLGNNVEAVST 136
S+ QG SE K+A G P T EGE + +++ ++D TT+ G+N E T
Sbjct: 1788 SLTQGETSSEIAPPASKKAKGSESHPDTSEGENLAKEPAIDELMDATTTTDGDNEE---T 1844
Query: 137 EA-NKESDKQQYIIAK 151
EA N E ++Y+ A+
Sbjct: 1845 EAENAEEKTEEYVEAQ 1860
>At2g19530 unknown protein
Length = 202
Score = 28.5 bits (62), Expect = 3.4
Identities = 18/61 (29%), Positives = 32/61 (51%), Gaps = 4/61 (6%)
Query: 84 KQGSAESENLKQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKLGNNVEAVSTEANKESD 143
K+G+ E++ K+A ET E EE + L+ + +FT + + + V+TE K +
Sbjct: 76 KKGAEETDKAKEA----ETAIPEKEETKLIPELDPLFEFTDATDQSMFQTVATEHVKVAR 131
Query: 144 K 144
K
Sbjct: 132 K 132
>At3g19060 hypothetical protein
Length = 1647
Score = 28.1 bits (61), Expect = 4.4
Identities = 17/60 (28%), Positives = 28/60 (46%)
Query: 101 ETPTIEGEEKSCVTSLESMVDFTTSKLGNNVEAVSTEANKESDKQQYIIAKGVKKLSENK 160
E +I +E + LE + + + +GN VE ++T K D+ + K LSE K
Sbjct: 731 ELESINVKENEKLAYLEKLFESSLMGIGNLVEELATVVRKLQDESSVALTGMAKDLSELK 790
>At5g18550 zinc finger -like protein
Length = 456
Score = 27.7 bits (60), Expect = 5.8
Identities = 16/43 (37%), Positives = 22/43 (50%), Gaps = 2/43 (4%)
Query: 196 KVKTAAICHNDTSQWSPKHLAF-HV-LKVQPGTVPVCHFLQHG 236
K T+ H+ SP+ H+ L ++PG VP HF QHG
Sbjct: 307 KFGTSCRFHHPMEAASPEASTLSHIGLPLRPGAVPCTHFAQHG 349
>At2g27600 putative AAA-type ATPase
Length = 435
Score = 27.3 bits (59), Expect = 7.5
Identities = 23/93 (24%), Positives = 41/93 (43%), Gaps = 11/93 (11%)
Query: 107 GEEKSCVTSLESM----------VDFTTSKLGNNVEAVSTEANKESDKQQYIIAKGVKKL 156
GE + V++L M VD S G E +EA++ + + +GV
Sbjct: 207 GESEKLVSNLFEMARESAPSIIFVDEIDSLCGTRGEGNESEASRRIKTELLVQMQGVGH- 265
Query: 157 SENKIVVCHLLSYPYAVFYCHKLRATKVYYLPM 189
++ K++V + PYA+ + R K Y+P+
Sbjct: 266 NDEKVLVLAATNTPYALDQAIRRRFDKRIYIPL 298
>At1g20870 unknown protein
Length = 463
Score = 27.3 bits (59), Expect = 7.5
Identities = 16/47 (34%), Positives = 23/47 (48%), Gaps = 1/47 (2%)
Query: 53 KTGATFLPREVANS-IPFSSNKVENILNYFSIKQGSAESENLKQAIG 98
K G T + RE + IPFSS E+ + G+A E L ++G
Sbjct: 312 KNGETLMEREKTDKPIPFSSEMKESDAEPSVVTTGTASKETLGSSVG 358
>At1g10240 unknown protein
Length = 680
Score = 27.3 bits (59), Expect = 7.5
Identities = 20/65 (30%), Positives = 29/65 (43%), Gaps = 5/65 (7%)
Query: 133 AVSTEANKESDKQQYIIAKGVKKLSENKIVVCHLLSYP----YAVFYCHKLRA-TKVYYL 187
++ T A ES + KL E ++ H S+ Y V + KL KVY++
Sbjct: 499 SLKTGAPMESHAASVLTPFAFSKLQEQLVLAAHYASFQMDEGYLVRHHTKLDGGRKVYWV 558
Query: 188 PMEGI 192
P EGI
Sbjct: 559 PQEGI 563
>At1g23320 putative alliinase
Length = 388
Score = 26.9 bits (58), Expect = 9.8
Identities = 29/121 (23%), Positives = 50/121 (40%), Gaps = 23/121 (19%)
Query: 13 LFIYWPYDASVTQLHDYQNLDLFFFEKDL-HNGTKLNMQFTKTGATFLPREVANSIPFSS 71
L YWP+ +T+ D+ +L LF F K H G+++ K EVA
Sbjct: 191 LAYYWPHYTPITRRQDH-DLMLFTFSKITGHAGSRIGWALVK------DIEVA------- 236
Query: 72 NKVENILNYFSIKQGSAESENLKQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKLGNNV 131
+ +++Y +I E+ +A T + K+C T ES ++ K+ +
Sbjct: 237 ---KKMVHYLTINSIGVSKESQTRA-----TTILNELTKTCRTQSESFFEYGYEKMKSRW 288
Query: 132 E 132
E
Sbjct: 289 E 289
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.133 0.402
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,354,623
Number of Sequences: 26719
Number of extensions: 209452
Number of successful extensions: 517
Number of sequences better than 10.0: 15
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 497
Number of HSP's gapped (non-prelim): 15
length of query: 243
length of database: 11,318,596
effective HSP length: 97
effective length of query: 146
effective length of database: 8,726,853
effective search space: 1274120538
effective search space used: 1274120538
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 58 (26.9 bits)
Medicago: description of AC148532.6


