

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148532.15 + phase: 1 /partial
(178 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
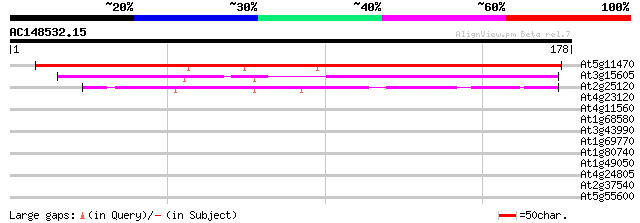
At5g11470 putative protein 173 5e-44
At3g15605 hypothetical protein, 3' partial 105 2e-23
At2g25120 hypothetical protein 44 6e-05
At4g23120 putative protein 39 0.001
At4g11560 unknown protein 32 0.14
At1g68580 unknown protein 32 0.14
At3g43990 putative protein 30 0.68
At1g69770 putative chromomethylase 30 0.89
At1g80740 chromomethylase (CMT1) 29 1.2
At1g49050 unknown protein 29 1.2
At4g24805 unknown protein 28 3.4
At2g37540 putative oxidoreductase 28 3.4
At5g55600 unknown protein 26 9.9
>At5g11470 putative protein
Length = 691
Score = 173 bits (438), Expect = 5e-44
Identities = 90/174 (51%), Positives = 119/174 (67%), Gaps = 7/174 (4%)
Query: 9 SPESETLEFKWGQKRGKGGKKRDIQFYQSFTLRGVDYSLFDNVYVKN----DFAEPRIGK 64
S SE LEFKWG+K+G GGKK+D+QFY+SFT G +Y L+D V V N D EP IG
Sbjct: 4 SVASEGLEFKWGKKKGVGGKKKDVQFYESFTYDGDEYRLYDCVLVGNASEPDSTEPFIGM 63
Query: 65 IIKIWETPT--LERKIKVQWFFRPIEVSKCLTWI-KIYFNELFFACGDGDGLATIHPLES 121
IIKIWE + +K+K+ WFF+P E++ L + + NE+F A G+G GLA + LE+
Sbjct: 64 IIKIWEHANKHIPKKVKLLWFFKPSEIAPYLEGVPNVLANEVFLASGEGLGLANTNQLEA 123
Query: 122 IAGKCNIVCISKDSRNPQLSDKVTWSAAFVFYRYFDVGQRKIVEEVDDKSIGIE 175
I GKC+++CISKD RNPQ SD+ SA FVF R FDVG K+V+ +DDK G++
Sbjct: 124 IGGKCSVLCISKDKRNPQPSDEKFNSADFVFCRAFDVGSCKVVDTIDDKIAGVD 177
>At3g15605 hypothetical protein, 3' partial
Length = 450
Score = 105 bits (261), Expect = 2e-23
Identities = 63/163 (38%), Positives = 87/163 (52%), Gaps = 24/163 (14%)
Query: 16 EFKWGQKRGKGGKKRDIQFYQSFTLRGVDYSLFDNVYVK-NDFAEPRIGKIIKIWETPTL 74
+FKWG KRG G K ++FY+SFTL G++Y LFD Y + +E IGK++ T
Sbjct: 56 DFKWGAKRGVGRKDNKVRFYESFTLEGIEYRLFDCAYFYVHGQSETSIGKLVTF--TRHQ 113
Query: 75 ER---KIKVQWFFRPIEVSKCLTWIKIYFNELFFACGDGDGLATIHPLESIAGKCNIVCI 131
+R + + QW +ELF ACGD G++ I+ +E+I GKCN+VC
Sbjct: 114 QRFLGEYEPQW------------------DELFLACGDEKGVSNINDVETIMGKCNVVCT 155
Query: 132 SKDSRNPQLSDKVTWSAAFVFYRYFDVGQRKIVEEVDDKSIGI 174
S D RNP+ K A ++F R FD R I E+ D GI
Sbjct: 156 SDDRRNPRPGTKELRRAKYIFSRTFDTRLRIISEDFADAIAGI 198
>At2g25120 hypothetical protein
Length = 380
Score = 43.5 bits (101), Expect = 6e-05
Identities = 43/156 (27%), Positives = 71/156 (44%), Gaps = 17/156 (10%)
Query: 24 GKGGKKRDIQFYQSFTLRGVDYSLFDNV-YVKNDFAEPRIGKIIKIWETPTLER--KIKV 80
GKG KK+ +++FT RG Y+L D+V V +D IIK P E+ K+ V
Sbjct: 77 GKGKKKKS--HFKTFTFRGNQYALEDSVQLVPDDPNSKPYCAIIKDIYIPNKEKYVKLAV 134
Query: 81 QWFFRPIEVSK--CLTWIKIYFNELFFACGDGDGLATIHPLESIAGKCNIVCISKDSRNP 138
WF+RP +V K W LF++ + A ES+ KC + + ++ + P
Sbjct: 135 HWFYRPEDVDKKHVGKWESKDSRNLFYSFHRDEVFA-----ESVKHKCVVNFVPENKQIP 189
Query: 139 QLSDKVTWSAAFVFYRYFDVGQRKIVEEVDDKSIGI 174
+ F+ +D ++K V + DK+ +
Sbjct: 190 NRRE----HPCFIVQNVYDFVKKK-VRKFTDKNFDV 220
>At4g23120 putative protein
Length = 360
Score = 39.3 bits (90), Expect = 0.001
Identities = 41/164 (25%), Positives = 70/164 (42%), Gaps = 22/164 (13%)
Query: 24 GKGGKKRDIQFYQSFTLRGVDYSLFDNVYVKNDFAEPRIGKIIKIWETPTLER--KIKVQ 81
GKG KK+ Y++F Y L D+V + + E IIK T E K++VQ
Sbjct: 40 GKGEKKKC--HYKTFQFHANKYGLEDSVLLVPEDGEKPYVAIIKDIYTQRKEGHVKLEVQ 97
Query: 82 WFFRPIEVSKCL--TWIKIYFNELFFACGDGDGLATIHPLESIAGKCNIVCISKDSRNP- 138
W +RP EV K W +LF++ + A ES+ C + + ++ + P
Sbjct: 98 WLYRPEEVEKKYVGNWKSKGSRDLFYSFHRDEVFA-----ESVKDDCIVHFVQENKQIPN 152
Query: 139 ----------QLSDKVTWSAAFVFYRYFDVGQRKIVEEVDDKSI 172
+ D V + + FD+ Q++ ++ +K+I
Sbjct: 153 RRKHPGFIVQHVYDNVKKKLRKLTFNGFDLQQKREIDHFVEKTI 196
>At4g11560 unknown protein
Length = 587
Score = 32.3 bits (72), Expect = 0.14
Identities = 35/149 (23%), Positives = 61/149 (40%), Gaps = 15/149 (10%)
Query: 24 GKGGKKRDIQFYQSFTLRGVDYSLFDNVYV--KNDFAEPRIGKIIKIWETPTLERKIKVQ 81
GKG KR + F G Y L V + ++ +P + I I +T I Q
Sbjct: 112 GKGKGKRT--HFNQFAYDGNTYDLEVPVLLVPEDKSQKPYVAIIKDITQTKDGSMMILGQ 169
Query: 82 WFFRPIEVSK--CLTWIKIYFNELFFACGDGDGLATIHPLESIAGKCNIVCISKDSRNPQ 139
WF+RP E K W ELF++ + P ES+ +C + + + P+
Sbjct: 170 WFYRPEEAEKRGGGNWQSSDTRELFYSFHRDE-----VPAESVMHRCVVYFVPAHKQLPK 224
Query: 140 LSDKVTWSAAFVFYRYFDVGQRKIVEEVD 168
+ + F+ + +D ++K+ + D
Sbjct: 225 RKN----NPGFIVRKVYDTVEKKLWKLTD 249
>At1g68580 unknown protein
Length = 648
Score = 32.3 bits (72), Expect = 0.14
Identities = 23/85 (27%), Positives = 39/85 (45%), Gaps = 2/85 (2%)
Query: 10 PESETLEFKWGQKRGKGGKKRDIQFYQSFTLRGVDYSLFDNVYVKNDFAEPRIGKIIKIW 69
P + F W K+R + YQS+ GV S+ D VYV + + + I ++
Sbjct: 118 PIQQIKTFSWMGFSWTCRKRR--KHYQSYLRNGVRISVNDFVYVLAEQHKRLVAYIEDLY 175
Query: 70 ETPTLERKIKVQWFFRPIEVSKCLT 94
E ++ + V+WF + EV L+
Sbjct: 176 EDSKGKKMVVVRWFHKTEEVGSVLS 200
>At3g43990 putative protein
Length = 380
Score = 30.0 bits (66), Expect = 0.68
Identities = 36/121 (29%), Positives = 54/121 (43%), Gaps = 14/121 (11%)
Query: 24 GKGGKKRDIQFYQSFTLRGVDYSLFDNV--YVKNDFAEPRIGKIIKIW--ETPTLERKIK 79
GKG K+ Y++F G Y L D V Y +++ + I I I+ E L K++
Sbjct: 65 GKGQNKKC--HYETFEFHGKQYRLKDFVLLYPEDNKQKEYIAIIKDIYSQEKDGLV-KME 121
Query: 80 VQWFFR--PIEVSKCLTWIKIYFNELFFACGDGDGLATIHPLESIAGKCNIVCISKDSRN 137
VQWF+R IE W E+FF+ + A ES+ KC + + D +
Sbjct: 122 VQWFYRREDIEEKHFGKWKTENPREIFFSFHCDEVFA-----ESVKYKCLVYFVPDDKQI 176
Query: 138 P 138
P
Sbjct: 177 P 177
>At1g69770 putative chromomethylase
Length = 839
Score = 29.6 bits (65), Expect = 0.89
Identities = 19/63 (30%), Positives = 33/63 (52%), Gaps = 1/63 (1%)
Query: 45 YSLFDNVYVKN-DFAEPRIGKIIKIWETPTLERKIKVQWFFRPIEVSKCLTWIKIYFNEL 103
Y L D+ YV++ + +P I KII+++E + +WF+RP + I I +
Sbjct: 110 YELNDDAYVQSGEGKDPFICKIIEMFEGANGKLYFTARWFYRPSDTVMKEFEILIKKKRV 169
Query: 104 FFA 106
FF+
Sbjct: 170 FFS 172
>At1g80740 chromomethylase (CMT1)
Length = 791
Score = 29.3 bits (64), Expect = 1.2
Identities = 20/70 (28%), Positives = 35/70 (49%), Gaps = 8/70 (11%)
Query: 25 KGGKKRDIQF-------YQSFTLRGVDYSLFDNVYVKNDFAEPR-IGKIIKIWETPTLER 76
KGGKK D + + + GV +L D+VYV + + I K+I+++E
Sbjct: 54 KGGKKEDEEIIKQAKCHFDKALVDGVLINLNDDVYVTGLPGKLKFIAKVIELFEADDGVP 113
Query: 77 KIKVQWFFRP 86
+ +W++RP
Sbjct: 114 YCRFRWYYRP 123
>At1g49050 unknown protein
Length = 583
Score = 29.3 bits (64), Expect = 1.2
Identities = 14/45 (31%), Positives = 25/45 (55%)
Query: 131 ISKDSRNPQLSDKVTWSAAFVFYRYFDVGQRKIVEEVDDKSIGIE 175
+S D RN D + +A+FVF Y + R+ E + ++ +G+E
Sbjct: 111 VSPDERNRDDDDNLRETASFVFPVYHKLRAREFHERILEEDLGLE 155
>At4g24805 unknown protein
Length = 247
Score = 27.7 bits (60), Expect = 3.4
Identities = 15/51 (29%), Positives = 28/51 (54%), Gaps = 2/51 (3%)
Query: 36 QSFTLRGVDYSLFDNVYVKNDFAEPRIGKIIKIWETPTLERKIKV-QWFFR 85
+ +R YS ++ Y+K+ + + K+ K+W T +RK++V FFR
Sbjct: 49 EGIRIRHHGYSSYE-AYIKHQLNKTQNPKLRKVWTTRDWDRKVRVFSTFFR 98
>At2g37540 putative oxidoreductase
Length = 321
Score = 27.7 bits (60), Expect = 3.4
Identities = 16/34 (47%), Positives = 21/34 (61%), Gaps = 5/34 (14%)
Query: 116 IHP-LESIAGK----CNIVCISKDSRNPQLSDKV 144
+HP LE + GK CNIV SK + N L+DK+
Sbjct: 275 LHPDLEGVTGKYFGDCNIVAPSKFATNNSLADKL 308
>At5g55600 unknown protein
Length = 663
Score = 26.2 bits (56), Expect = 9.9
Identities = 18/74 (24%), Positives = 32/74 (42%), Gaps = 2/74 (2%)
Query: 16 EFKWGQKRGKGGKKRDIQFYQSFTLRGVDYSLFDNVYVKNDFAEPRIGKIIKIWETPTLE 75
E W GK+ ++ Y SF G + V+V + + + + ++E
Sbjct: 142 EIMWSGAPWMCGKQ--LKHYPSFCRNGTTIGVQSFVFVLSKGEDRYVAYLEDMYEDKRGL 199
Query: 76 RKIKVQWFFRPIEV 89
+K+KV+WF EV
Sbjct: 200 KKVKVRWFHYTKEV 213
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.321 0.140 0.446
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,595,775
Number of Sequences: 26719
Number of extensions: 200256
Number of successful extensions: 391
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 382
Number of HSP's gapped (non-prelim): 13
length of query: 178
length of database: 11,318,596
effective HSP length: 93
effective length of query: 85
effective length of database: 8,833,729
effective search space: 750866965
effective search space used: 750866965
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 56 (26.2 bits)
Medicago: description of AC148532.15


