

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148532.13 + phase: 0
(303 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
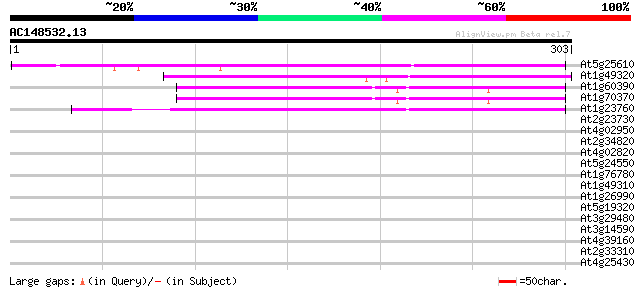
At5g25610 dehydration-induced protein RD22 308 3e-84
At1g49320 unknown protein 156 1e-38
At1g60390 132 2e-31
At1g70370 unknown protein 132 2e-31
At1g23760 putative polygalacuronase isoenzyme 1 beta subunit 122 2e-28
At2g23730 hypothetical protein 31 0.72
At4g02950 putative protein 30 1.2
At2g34820 putative bHLH transcription factor (bHLH053) 30 1.2
At4g02820 Unknown protein (At4g02820) 30 2.1
At5g24550 beta-glucosidase 29 2.8
At1g76780 putative heat shock protein 29 3.6
At1g49310 unknown protein 29 3.6
At1g26990 polyprotein, putative 29 3.6
At5g19320 RAN GTPase activating protein 2 28 4.7
At3g29480 hypothetical protein 28 4.7
At3g14590 hypothetical protein 28 4.7
At4g39160 putative protein 28 6.1
At2g33310 auxin regulated protein (IAA13) 28 6.1
At4g25430 hypothetical protein 28 8.0
>At5g25610 dehydration-induced protein RD22
Length = 392
Score = 308 bits (788), Expect = 3e-84
Identities = 171/370 (46%), Positives = 220/370 (59%), Gaps = 74/370 (20%)
Query: 2 LPPQLYWKSMLPNSPMPKAITNLLHPAGYWSEEKGTWVDVGKGGVDVGVRKGYY------ 55
L P+ YW + LPN+P+P ++ NLL +++EK T V VGKGGV+V KG
Sbjct: 23 LTPERYWSTALPNTPIPNSLHNLL--TFDFTDEKSTNVQVGKGGVNVNTHKGKTGSGTAV 80
Query: 56 ---------------EGGGTDVNVGVGR-------------------------------- 68
GGGT V+VG G+
Sbjct: 81 NVGKGGVRVDTGKGKPGGGTHVSVGSGKGHGGGVAVHTGKPGKRTDVGVGKGGVTVHTRH 140
Query: 69 -------------SPFIYNYAASETQLHDKPNVALFFLEKDLHHGTKLNLQFSKTTS--N 113
+PF+YNYAA ETQLHD PN ALFFLEKDL G ++N++F+
Sbjct: 141 KGRPIYVGVKPGANPFVYNYAAKETQLHDDPNAALFFLEKDLVRGKEMNVRFNAEDGYGG 200
Query: 114 AATFLPRQVANSIPFSSNKMEYIINKLNIKKGSKGVQIVKNTISECEEQGIKGEEKVCVT 173
FLPR A ++PF S K + + +++ GS+ +++K TI ECE + + GEEK C T
Sbjct: 201 KTAFLPRGEAETVPFGSEKFSETLKRFSVEAGSEEAEMMKKTIEECEARKVSGEEKYCAT 260
Query: 174 SLESMVDFTTSKLGK-NVEAVSTEV-NKESNLQQYTIAS-GVKKLGEKNKAVVCHKENYP 230
SLESMVDF+ SKLGK +V AVSTEV K + +Q+Y IA+ GVKKL + +K+VVCHK+ YP
Sbjct: 261 SLESMVDFSVSKLGKYHVRAVSTEVAKKNAPMQKYKIAAAGVKKLSD-DKSVVCHKQKYP 319
Query: 231 YAVFYCHKTDTTKAYSVPLEGADGSRVKAIAVCHTDTSEWNPKHLAFQVLKVQPGTVPVC 290
+AVFYCHK T Y+VPLEG +G R KA+AVCH +TS WNP HLAF+VLKV+PGTVPVC
Sbjct: 320 FAVFYCHKAMMTTVYAVPLEGENGMRAKAVAVCHKNTSAWNPNHLAFKVLKVKPGTVPVC 379
Query: 291 HLLPEDHVVW 300
H LPE HVVW
Sbjct: 380 HFLPETHVVW 389
>At1g49320 unknown protein
Length = 280
Score = 156 bits (395), Expect = 1e-38
Identities = 87/231 (37%), Positives = 132/231 (56%), Gaps = 12/231 (5%)
Query: 84 DKPNVALFFLEKDLHHGTKLNLQFSKTT-SNAATFLPRQVANSIPFSSNKMEYIINKLNI 142
D P++ ++F DL GTKL + F K L RQ A+ IPF+ +K++++++ +I
Sbjct: 51 DDPSLYMYFTLNDLKLGTKLLIYFYKNDLQKLPPLLTRQQADLIPFTKSKLDFLLDHFSI 110
Query: 143 KKGSKGVQIVKNTISECEEQGIKGEEKVCVTSLESMVDFTTSKLGKNVE--AVSTEVNKE 200
K S + +K T+ C+ + I+GE K C TSLES++D +G NV+ ++T+V
Sbjct: 111 TKDSPQGKAIKETLGHCDAKAIEGEHKFCGTSLESLIDLVKKTMGYNVDLKVMTTKVMVP 170
Query: 201 SN------LQQYTIASGVKKLGEKNKAVVCHKENYPYAVFYCH-KTDTTKAYSVPLEGAD 253
+ L YT K+L K + CH+ YPYAV+YCH ++ + V L D
Sbjct: 171 AQNSISYALHNYTFVEAPKEL-VGIKMLGCHRMPYPYAVYYCHGHKGGSRVFEVNLVTDD 229
Query: 254 G-SRVKAIAVCHTDTSEWNPKHLAFQVLKVQPGTVPVCHLLPEDHVVWIRK 303
G RV AVCH DTS W+ H+AF+VLK++P + PVCH P D++VW+ K
Sbjct: 230 GRQRVVGPAVCHMDTSTWDADHVAFKVLKMEPRSAPVCHFFPLDNIVWVTK 280
>At1g60390
Length = 624
Score = 132 bits (333), Expect = 2e-31
Identities = 75/214 (35%), Positives = 107/214 (49%), Gaps = 6/214 (2%)
Query: 91 FFLEKDLHHGTKLNLQFSKTTSNAATFLPRQVANSIPFSSNKMEYIINKLNIKKGSKGVQ 150
FF E L GT + + K TFLPR + ++PFSS+ + I + S
Sbjct: 409 FFREAMLKEGTLMQMPDIKDKMPKRTFLPRNIVKNLPFSSSTIGEIWRVFGAGENSSMAG 468
Query: 151 IVKNTISECEEQGIKGEEKVCVTSLESMVDFTTSKLGKNVEAVSTEVNKESNLQQYTI-- 208
I+ + +SECE GE K CV S E M+DF TS LG+ V +TE N + ++ I
Sbjct: 469 IISSAVSECERPASHGETKRCVGSAEDMIDFATSVLGRGVVVRTTE-NVVGSKKKVVIGK 527
Query: 209 ASGVKKLGEKNKAVVCHKENYPYAVFYCHKTDTTKAYSVPLEGADGSRV--KAIAVCHTD 266
+G+ G+ +AV CH+ YPY ++YCH + Y L +A+CH D
Sbjct: 528 VNGING-GDVTRAVSCHQSLYPYLLYYCHSVPRVRVYETDLLDPKSLEKINHGVAICHID 586
Query: 267 TSEWNPKHLAFQVLKVQPGTVPVCHLLPEDHVVW 300
TS W+P H AF L PG + VCH + E+ + W
Sbjct: 587 TSAWSPSHGAFLALGSGPGQIEVCHWIFENDMTW 620
>At1g70370 unknown protein
Length = 626
Score = 132 bits (332), Expect = 2e-31
Identities = 75/214 (35%), Positives = 111/214 (51%), Gaps = 6/214 (2%)
Query: 91 FFLEKDLHHGTKLNLQFSKTTSNAATFLPRQVANSIPFSSNKMEYIINKLNIKKGSKGVQ 150
FF E L GT + + K +FLPR + +PFS++K+ I + + S
Sbjct: 411 FFRESSLKEGTVIPMPDIKDKMPKRSFLPRSIITKLPFSTSKLGEIKRIFHAVENSTMGG 470
Query: 151 IVKNTISECEEQGIKGEEKVCVTSLESMVDFTTSKLGKNVEAVSTEVNKESNLQQYTI-- 208
I+ + ++ECE GE K CV S E M+DF TS LG++V +TE N + ++ I
Sbjct: 471 IITDAVTECERPPSVGETKRCVGSAEDMIDFATSVLGRSVVLRTTE-NVAGSKEKVVIGK 529
Query: 209 ASGVKKLGEKNKAVVCHKENYPYAVFYCHKTDTTKAYSVPLEGADGSRV--KAIAVCHTD 266
+G+ G+ KAV CH+ YPY ++YCH + Y L + + IA+CH D
Sbjct: 530 VNGING-GKLTKAVSCHQSLYPYLLYYCHSVPKVRVYEADLLELNSKKKINHGIAICHMD 588
Query: 267 TSEWNPKHLAFQVLKVQPGTVPVCHLLPEDHVVW 300
TS W P H AF L +PG + VCH + E+ + W
Sbjct: 589 TSSWGPSHGAFLALGSKPGRIEVCHWIFENDMNW 622
>At1g23760 putative polygalacuronase isoenzyme 1 beta subunit
Length = 622
Score = 122 bits (307), Expect = 2e-28
Identities = 83/270 (30%), Positives = 121/270 (44%), Gaps = 24/270 (8%)
Query: 34 EKGTWVDVGKGGVDVGVRKGYYEGGGTDVNVGVGRSPFIYNYAASETQLHDKPNVALFFL 93
+ T+ D K GV+ GGG VN V FF
Sbjct: 370 QNSTFKDYTKTGVEFAKYNRSSLGGGKTVNKWV--------------------EPGKFFR 409
Query: 94 EKDLHHGTKLNLQFSKTTSNAATFLPRQVANSIPFSSNKMEYIINKLNIKKGSKGVQIVK 153
E L GT + + K +FLPR + + +PFS++K+ I + S I+
Sbjct: 410 ESMLKEGTLIWMPDIKDKMPKRSFLPRSIVSKLPFSTSKIAEIKRVFHANDNSTMEGIIT 469
Query: 154 NTISECEEQGIKGEEKVCVTSLESMVDFTTSKLGKNVEAVSTE-VNKESNLQQYTIASGV 212
+ + ECE E K CV S E M+DF TS LG++V +TE V +G+
Sbjct: 470 DAVRECERPPTVSETKRCVGSAEDMIDFATSVLGRSVVLRTTESVAGSKEKVMIGKVNGI 529
Query: 213 KKLGEKNKAVVCHKENYPYAVFYCHKTDTTKAY-SVPLEGADGSRVK-AIAVCHTDTSEW 270
G K+V CH+ YPY ++YCH + Y S L+ +++ IA+CH DTS W
Sbjct: 530 NG-GRVTKSVSCHQSLYPYLLYYCHSVPKVRVYESDLLDPKSKAKINHGIAICHMDTSAW 588
Query: 271 NPKHLAFQVLKVQPGTVPVCHLLPEDHVVW 300
H AF +L +PG + VCH + E+ + W
Sbjct: 589 GANHGAFMLLGSRPGQIEVCHWIFENDMNW 618
>At2g23730 hypothetical protein
Length = 216
Score = 31.2 bits (69), Expect = 0.72
Identities = 23/86 (26%), Positives = 36/86 (41%), Gaps = 2/86 (2%)
Query: 164 IKGEEKVCVTSLESMVDFTT--SKLGKNVEAVSTEVNKESNLQQYTIASGVKKLGEKNKA 221
+K E + + +++S D T +GKN E S+ + Q IA VK G+
Sbjct: 20 VKSEPEADLNAVKSSTDLVTVTGPIGKNGEGESSPSEPKWLQQDEPIALWVKWRGKWQAG 79
Query: 222 VVCHKENYPYAVFYCHKTDTTKAYSV 247
+ C K ++P T K Y V
Sbjct: 80 IRCAKADWPLTTLRGKPTHDRKKYCV 105
>At4g02950 putative protein
Length = 318
Score = 30.4 bits (67), Expect = 1.2
Identities = 33/146 (22%), Positives = 59/146 (39%), Gaps = 30/146 (20%)
Query: 106 QFSKTTSNAATFLPRQVANSIPFSSNKMEYIINKLNIKKGSKGVQI--VKNTISECEEQG 163
+FSK + + R VA I S++ Y +++L + S V + ++N +Q
Sbjct: 153 KFSKNQQDRPPLM-RVVAKRIDNGSSRPSYSLDELLAPRDSSTVAVGSIRN-----RDQE 206
Query: 164 IKGEEKVCVTSLESMVDFTTSKLGK----------------------NVEAVSTEVNKES 201
+K E S+E +++FT S K NVE + E+ K
Sbjct: 207 VKNRESSPSDSVEEVINFTDSPAMKMIVMLQPYGYTRMIQVEVTADDNVEELRKELVKMQ 266
Query: 202 NLQQYTIASGVKKLGEKNKAVVCHKE 227
+ + G+ L K+K V H++
Sbjct: 267 ERGELNLPQGMYYLIHKHKQAVLHED 292
>At2g34820 putative bHLH transcription factor (bHLH053)
Length = 291
Score = 30.4 bits (67), Expect = 1.2
Identities = 19/58 (32%), Positives = 28/58 (47%), Gaps = 1/58 (1%)
Query: 143 KKGSKGVQIVKNTISECEE-QGIKGEEKVCVTSLESMVDFTTSKLGKNVEAVSTEVNK 199
KKGS VQ+ + E + Q E+VC+ E + D TT + +S E+NK
Sbjct: 226 KKGSSNVQMETQYLLESQAIQEKLSTEEVCLVPCEMVQDLTTEETICRTPNISREINK 283
>At4g02820 Unknown protein (At4g02820)
Length = 532
Score = 29.6 bits (65), Expect = 2.1
Identities = 20/84 (23%), Positives = 43/84 (50%), Gaps = 3/84 (3%)
Query: 141 NIKKGSKGVQIVKNTISECEEQG-IKGEEKVCVTSLESMVDFTTSKLGKNVEAVSTEVNK 199
++KK + V++VK E EEQG +KG EK + +L + ++L ++ + +
Sbjct: 435 SVKKWTVNVRLVKGACKELEEQGNVKGAEK--LMTLLQKAGYVNTQLYNSLLRTYAKAGE 492
Query: 200 ESNLQQYTIASGVKKLGEKNKAVV 223
+ + + +A +L E+ K ++
Sbjct: 493 MALIVEERMAKDNVELDEETKELI 516
>At5g24550 beta-glucosidase
Length = 531
Score = 29.3 bits (64), Expect = 2.8
Identities = 36/122 (29%), Positives = 54/122 (43%), Gaps = 19/122 (15%)
Query: 47 DVGVRKGYYEGGGTDVNVGVGRSPFIYN---YAASETQLHDKPNVALFFLEKDLHHGTKL 103
D GV Y+ G V G GRSP I++ +A E D +VA+ D +H K
Sbjct: 42 DFGVASSAYQYEGA-VEEG-GRSPSIWDNFTHAFPERTNMDNGDVAV-----DFYHRYKD 94
Query: 104 NLQFSKTTSNAATFLPRQVANSIPFSSNKMEYIINKLNIKKGSKGVQIVKNTISECEEQG 163
+++ K + + + +P S K+ +NK +GVQ KN I E + G
Sbjct: 95 DIKLIKEMNMDSFRFSLSWSRILP--SGKLSDGVNK-------EGVQFYKNLIDELIKNG 145
Query: 164 IK 165
IK
Sbjct: 146 IK 147
>At1g76780 putative heat shock protein
Length = 1871
Score = 28.9 bits (63), Expect = 3.6
Identities = 25/119 (21%), Positives = 47/119 (39%), Gaps = 23/119 (19%)
Query: 119 PRQVANSIPFSSNKMEYIINKLNIKKGSKGVQIVKNTISECEE----------------- 161
P ++ +I + + + N++ G K ++V+ I +CEE
Sbjct: 792 PEKITGTIKQELVSLNSQLRQENVEDGDKTQELVEEKIKDCEEEEGSEESKIKTDDVVRK 851
Query: 162 -QGIKGEE-----KVCVTSLESMVDFTTSKLGKNVEAVSTEVNKESNLQQYTIASGVKK 214
QGIK EE + T + +V+ TT K E + E + E+ G+++
Sbjct: 852 VQGIKEEELYKPKREHGTKITELVEETTGDYEKQEEKETAESDIEAECGSLRKVDGIEE 910
>At1g49310 unknown protein
Length = 82
Score = 28.9 bits (63), Expect = 3.6
Identities = 10/20 (50%), Positives = 14/20 (70%)
Query: 7 YWKSMLPNSPMPKAITNLLH 26
YWK M+ N P+P+ I LL+
Sbjct: 35 YWKKMMKNEPLPEPIKELLN 54
>At1g26990 polyprotein, putative
Length = 1436
Score = 28.9 bits (63), Expect = 3.6
Identities = 34/150 (22%), Positives = 62/150 (40%), Gaps = 16/150 (10%)
Query: 134 EYIINKLNIKKGSKGVQI-VKNTISECEEQGI-------KGEE--KVCVTSLESMVDFTT 183
E+ N L++ +K +Q V T EC Q + KG + + + +L+ + +
Sbjct: 481 EFKFNLLSVSVLTKTLQSKVSFTSDECMIQALTKELMLGKGSQVGNLYILNLDKSLVDVS 540
Query: 184 SKLGKNVEAVSTEVNKESNLQQYTIAS-GVKKLGEKNKAVVCHKENYPYAVFYCHKTDTT 242
S GK+V + V ES + + K+ + ++ K+ +CH +
Sbjct: 541 SFPGKSV---CSSVKNESEMWHKRLGHPSFAKIDTLSDVLMLPKQKINKDSSHCHVCHLS 597
Query: 243 KAYSVPLEGADGSRVKAIAVCHTDTSEWNP 272
K +P + + R KA + H DT W P
Sbjct: 598 KQKHLPFKSVNHIREKAFELVHIDT--WGP 625
>At5g19320 RAN GTPase activating protein 2
Length = 545
Score = 28.5 bits (62), Expect = 4.7
Identities = 22/93 (23%), Positives = 44/93 (46%), Gaps = 6/93 (6%)
Query: 66 VGRSPFIYNYAASETQLHDKPNVALFFLEKDLHHGTKLNLQFSKTTSNAATFLPRQVANS 125
V RSP + N+ S T++ K +AL + H KL+L+ + + A L + +++
Sbjct: 294 VKRSPLLENFRCSSTRVGSKGGIALSEALEHCTHMEKLDLRDNMFGTEAGVSLSKTLSS- 352
Query: 126 IPFSSNKMEYIINKLNIKKGSKGVQIVKNTISE 158
+ E ++ LN++ +G + N + E
Sbjct: 353 ---FKHMTELYLSYLNLE--DEGAIAIVNALKE 380
>At3g29480 hypothetical protein
Length = 718
Score = 28.5 bits (62), Expect = 4.7
Identities = 14/46 (30%), Positives = 23/46 (49%)
Query: 53 GYYEGGGTDVNVGVGRSPFIYNYAASETQLHDKPNVALFFLEKDLH 98
GY+ GG + RS N+ +S + + ++A+ FLE D H
Sbjct: 49 GYFSGGTSPEEADSWRSRVERNFGSSRCPVEYRVDLAVHFLEGDAH 94
>At3g14590 hypothetical protein
Length = 705
Score = 28.5 bits (62), Expect = 4.7
Identities = 38/176 (21%), Positives = 70/176 (39%), Gaps = 30/176 (17%)
Query: 63 NVGVGRSPFIYNYAASETQLHDKPNVALFFLEKDLHHGTKLNLQFSKTTSNAATFLPRQV 122
N+ +GR E +L+D P + ++D+ F+ +N +F V
Sbjct: 354 NIKMGRLHLAITVLEDEAKLNDDPFEGVTICKEDMW------ASFASDVTNKGSF-SSVV 406
Query: 123 ANSIPFSSNKMEYIINKLNIKKG----SKGVQIV-----KNTISECEEQGIKGEEKVCVT 173
++ P + ME I + + G G ++ + S C + I+ C
Sbjct: 407 SDKSPRVPDNMEPINIEGQEETGIWVHQPGTEVSQIWEPRKGKSRCLDNKIQ-----CAG 461
Query: 174 SLESMVDFTTSKLGKNVEAVSTEVNKESNLQQYTIASGVKKLG----EKNKAVVCH 225
S+ S T+ N E+ ST+ N+E + ++ G+KK+G + K CH
Sbjct: 462 SVRS-----TASTSPNNESSSTDKNQEGKSEMKSVGWGLKKIGLVFHKNGKKEECH 512
>At4g39160 putative protein
Length = 545
Score = 28.1 bits (61), Expect = 6.1
Identities = 16/65 (24%), Positives = 29/65 (44%)
Query: 137 INKLNIKKGSKGVQIVKNTISECEEQGIKGEEKVCVTSLESMVDFTTSKLGKNVEAVSTE 196
+ ++ + S G + + +S C + GEE+ C+ + SK GK+ A S +
Sbjct: 200 VGEMEAQNASGGWEQEEQGVSPCINNTVTGEEENCMGNTVEEQSKRESKTGKSKRATSRK 259
Query: 197 VNKES 201
K S
Sbjct: 260 RKKTS 264
>At2g33310 auxin regulated protein (IAA13)
Length = 246
Score = 28.1 bits (61), Expect = 6.1
Identities = 28/130 (21%), Positives = 53/130 (40%), Gaps = 16/130 (12%)
Query: 37 TWVDVGKGGVDVGVRKGYYEGGGTDVNVGVGRSPFIYNYAASETQLHDKPNVALFFLEKD 96
T +++GKG ++ + G GGGT +G + D P+V
Sbjct: 3 TELEMGKGESELELGLGLSLGGGTAAKIGKSGGGGAWGERGRLLTAKDFPSV-------- 54
Query: 97 LHHGTKLNLQFSKTTSNAATFLPR--QVANSIPFSSNKMEYIINKLNIKKGSKGVQIVKN 154
G+K + + + S+A + PR QV P S++M ++N K + + K
Sbjct: 55 ---GSK---RAADSASHAGSSPPRSSQVVGWPPIGSHRMNSLVNNQATKSAREEEEAGKK 108
Query: 155 TISECEEQGI 164
+ + E + +
Sbjct: 109 KVKDDEPKDV 118
>At4g25430 hypothetical protein
Length = 455
Score = 27.7 bits (60), Expect = 8.0
Identities = 25/78 (32%), Positives = 38/78 (48%), Gaps = 5/78 (6%)
Query: 118 LPRQVANSIPFSSNKME-YIINKLNIKKGSKGVQIVKNTISECEEQGIKGEEKVCVTSLE 176
LP++ ++S + SN + IIN +K G+ N + E EE+ + E+ V +L
Sbjct: 378 LPQEPSSSKFYDSNTISPTIINAGETEKDVPGM----NKLEEEEERVVSEIERQIVDALV 433
Query: 177 SMVDFTTSKLGKNVEAVS 194
TTS G N AVS
Sbjct: 434 QETVETTSLWGLNANAVS 451
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.314 0.132 0.396
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,233,278
Number of Sequences: 26719
Number of extensions: 313485
Number of successful extensions: 644
Number of sequences better than 10.0: 19
Number of HSP's better than 10.0 without gapping: 9
Number of HSP's successfully gapped in prelim test: 10
Number of HSP's that attempted gapping in prelim test: 618
Number of HSP's gapped (non-prelim): 22
length of query: 303
length of database: 11,318,596
effective HSP length: 99
effective length of query: 204
effective length of database: 8,673,415
effective search space: 1769376660
effective search space used: 1769376660
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 60 (27.7 bits)
Medicago: description of AC148532.13


