

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148532.11 + phase: 0
(308 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
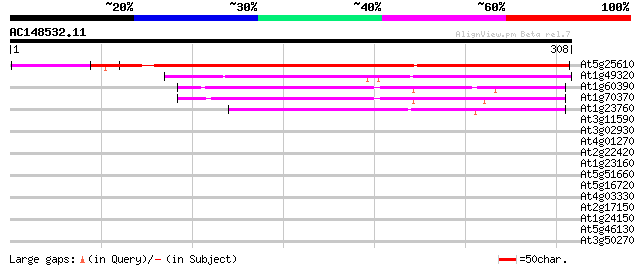
Score E
Sequences producing significant alignments: (bits) Value
At5g25610 dehydration-induced protein RD22 265 2e-71
At1g49320 unknown protein 153 1e-37
At1g60390 128 3e-30
At1g70370 unknown protein 126 2e-29
At1g23760 putative polygalacuronase isoenzyme 1 beta subunit 115 4e-26
At3g11590 unknown protein 30 1.7
At3g02930 unknown protein 30 2.2
At4g01270 putative RING zinc finger protein 28 4.8
At2g22420 putative peroxidase 28 4.8
At1g23160 GH3-like auxin-regulated protein 28 4.8
At5g51660 cleavage and polyadenylation specificity factor subunit 28 6.3
At5g16720 putative protein 28 6.3
At4g03330 SYR1-like syntaxin 28 6.3
At2g17150 unknown protein 28 6.3
At1g24150 unknown protein 28 6.3
At5g46130 putative protein 28 8.2
At3g50270 anthranilate N-hydroxycinnamoyl/benzoyltransferase - l... 28 8.2
>At5g25610 dehydration-induced protein RD22
Length = 392
Score = 265 bits (678), Expect = 2e-71
Identities = 140/273 (51%), Positives = 180/273 (65%), Gaps = 17/273 (6%)
Query: 45 DVGVRKG-------YEGGGTYLNDEK-IIPLIYFYPIPIPLNESQIQLDDKQNVTLFFLK 96
DVGV KG ++G Y+ + P +Y Y + QL D N LFFL+
Sbjct: 126 DVGVGKGGVTVHTRHKGRPIYVGVKPGANPFVYNYAA------KETQLHDDPNAALFFLE 179
Query: 97 KDLHHGTKLNLQFKETTSNNNGTKFLPREVANSIPFSSNKMENILNKFSIKEGSKEAEIV 156
KDL G ++N++F T FLPR A ++PF S K L +FS++ GS+EAE++
Sbjct: 180 KDLVRGKEMNVRFNAEDGYGGKTAFLPRGEAETVPFGSEKFSETLKRFSVEAGSEEAEMM 239
Query: 157 KRTISECEANGIKGEEKLCVTSLESMVDFTISKLGN-NVEAVSTEVDKNSNGLQQYVIAK 215
K+TI ECEA + GEEK C TSLESMVDF++SKLG +V AVSTEV K + +Q+Y IA
Sbjct: 240 KKTIEECEARKVSGEEKYCATSLESMVDFSVSKLGKYHVRAVSTEVAKKNAPMQKYKIAA 299
Query: 216 -GVKKLGEKNKTIVCHKENYPYAVFYCHKTDSTEVYSVPLEGVDGNMVKTIAVCHTDTSE 274
GVKKL + +K++VCHK+ YP+AVFYCHK T VY+VPLEG +G K +AVCH +TS
Sbjct: 300 AGVKKLSD-DKSVVCHKQKYPFAVFYCHKAMMTTVYAVPLEGENGMRAKAVAVCHKNTSA 358
Query: 275 WNPKHLAFYVLKVQPGTVPICHILPQDHVVWVS 307
WNP HLAF VLKV+PGTVP+CH LP+ HVVW S
Sbjct: 359 WNPNHLAFKVLKVKPGTVPVCHFLPETHVVWFS 391
Score = 44.7 bits (104), Expect = 6e-05
Identities = 22/59 (37%), Positives = 30/59 (50%)
Query: 2 LPPQLYWQSMLPNSPMPKAFTNLLHPAGYWSKEKARDASNGGLDVGVRKGYEGGGTYLN 60
L P+ YW + LPN+P+P + NLL K GG++V KG G GT +N
Sbjct: 23 LTPERYWSTALPNTPIPNSLHNLLTFDFTDEKSTNVQVGKGGVNVNTHKGKTGSGTAVN 81
>At1g49320 unknown protein
Length = 280
Score = 153 bits (386), Expect = 1e-37
Identities = 87/232 (37%), Positives = 132/232 (56%), Gaps = 11/232 (4%)
Query: 86 DKQNVTLFFLKKDLHHGTKLNLQFKETTSNNNGTKFLPREVANSIPFSSNKMENILNKFS 145
D ++ ++F DL GTKL + F + L R+ A+ IPF+ +K++ +L+ FS
Sbjct: 51 DDPSLYMYFTLNDLKLGTKLLIYFYKNDLQKL-PPLLTRQQADLIPFTKSKLDFLLDHFS 109
Query: 146 IKEGSKEAEIVKRTISECEANGIKGEEKLCVTSLESMVDFTISKLGNNVE--AVSTEV-- 201
I + S + + +K T+ C+A I+GE K C TSLES++D +G NV+ ++T+V
Sbjct: 110 ITKDSPQGKAIKETLGHCDAKAIEGEHKFCGTSLESLIDLVKKTMGYNVDLKVMTTKVMV 169
Query: 202 ---DKNSNGLQQYVIAKGVKKLGEKNKTIVCHKENYPYAVFYCH-KTDSTEVYSVPLEGV 257
+ S L Y + K+L K + CH+ YPYAV+YCH + V+ V L
Sbjct: 170 PAQNSISYALHNYTFVEAPKEL-VGIKMLGCHRMPYPYAVYYCHGHKGGSRVFEVNLVTD 228
Query: 258 DGNM-VKTIAVCHTDTSEWNPKHLAFYVLKVQPGTVPICHILPQDHVVWVSK 308
DG V AVCH DTS W+ H+AF VLK++P + P+CH P D++VWV+K
Sbjct: 229 DGRQRVVGPAVCHMDTSTWDADHVAFKVLKMEPRSAPVCHFFPLDNIVWVTK 280
>At1g60390
Length = 624
Score = 128 bits (322), Expect = 3e-30
Identities = 75/219 (34%), Positives = 108/219 (49%), Gaps = 13/219 (5%)
Query: 93 FFLKKDLHHGTKLNLQFKETTSNNNGTKFLPREVANSIPFSSNKMENILNKFSIKEGSKE 152
FF + L GT + Q + FLPR + ++PFSS+ + I F E S
Sbjct: 409 FFREAMLKEGTLM--QMPDIKDKMPKRTFLPRNIVKNLPFSSSTIGEIWRVFGAGENSSM 466
Query: 153 AEIVKRTISECEANGIKGEEKLCVTSLESMVDFTISKLGNNVEAVSTEVDKNSNGLQQYV 212
A I+ +SECE GE K CV S E M+DF S LG V +TE N G ++ V
Sbjct: 467 AGIISSAVSECERPASHGETKRCVGSAEDMIDFATSVLGRGVVVRTTE---NVVGSKKKV 523
Query: 213 IAKGVKKL--GEKNKTIVCHKENYPYAVFYCHKTDSTEVYSVPLEGVDGNMVKTI----A 266
+ V + G+ + + CH+ YPY ++YCH VY L +D ++ I A
Sbjct: 524 VIGKVNGINGGDVTRAVSCHQSLYPYLLYYCHSVPRVRVYETDL--LDPKSLEKINHGVA 581
Query: 267 VCHTDTSEWNPKHLAFYVLKVQPGTVPICHILPQDHVVW 305
+CH DTS W+P H AF L PG + +CH + ++ + W
Sbjct: 582 ICHIDTSAWSPSHGAFLALGSGPGQIEVCHWIFENDMTW 620
>At1g70370 unknown protein
Length = 626
Score = 126 bits (316), Expect = 2e-29
Identities = 72/217 (33%), Positives = 107/217 (49%), Gaps = 9/217 (4%)
Query: 93 FFLKKDLHHGTKLNLQFKETTSNNNGTKFLPREVANSIPFSSNKMENILNKFSIKEGSKE 152
FF + L GT + + + FLPR + +PFS++K+ I F E S
Sbjct: 411 FFRESSLKEGTVIPMP--DIKDKMPKRSFLPRSIITKLPFSTSKLGEIKRIFHAVENSTM 468
Query: 153 AEIVKRTISECEANGIKGEEKLCVTSLESMVDFTISKLGNNVEAVSTEVDKNSNGLQQYV 212
I+ ++ECE GE K CV S E M+DF S LG +V +TE N G ++ V
Sbjct: 469 GGIITDAVTECERPPSVGETKRCVGSAEDMIDFATSVLGRSVVLRTTE---NVAGSKEKV 525
Query: 213 IAKGVKKL--GEKNKTIVCHKENYPYAVFYCHKTDSTEVYSVPLEGVDG--NMVKTIAVC 268
+ V + G+ K + CH+ YPY ++YCH VY L ++ + IA+C
Sbjct: 526 VIGKVNGINGGKLTKAVSCHQSLYPYLLYYCHSVPKVRVYEADLLELNSKKKINHGIAIC 585
Query: 269 HTDTSEWNPKHLAFYVLKVQPGTVPICHILPQDHVVW 305
H DTS W P H AF L +PG + +CH + ++ + W
Sbjct: 586 HMDTSSWGPSHGAFLALGSKPGRIEVCHWIFENDMNW 622
>At1g23760 putative polygalacuronase isoenzyme 1 beta subunit
Length = 622
Score = 115 bits (287), Expect = 4e-26
Identities = 62/187 (33%), Positives = 90/187 (47%), Gaps = 3/187 (1%)
Query: 121 FLPREVANSIPFSSNKMENILNKFSIKEGSKEAEIVKRTISECEANGIKGEEKLCVTSLE 180
FLPR + + +PFS++K+ I F + S I+ + ECE E K CV S E
Sbjct: 433 FLPRSIVSKLPFSTSKIAEIKRVFHANDNSTMEGIITDAVRECERPPTVSETKRCVGSAE 492
Query: 181 SMVDFTISKLGNNVEAVSTEVDKNSNGLQQYVIAKGVKKLGEKNKTIVCHKENYPYAVFY 240
M+DF S LG +V +TE S G+ G K++ CH+ YPY ++Y
Sbjct: 493 DMIDFATSVLGRSVVLRTTESVAGSKEKVMIGKVNGING-GRVTKSVSCHQSLYPYLLYY 551
Query: 241 CHKTDSTEVYSVPL--EGVDGNMVKTIAVCHTDTSEWNPKHLAFYVLKVQPGTVPICHIL 298
CH VY L + IA+CH DTS W H AF +L +PG + +CH +
Sbjct: 552 CHSVPKVRVYESDLLDPKSKAKINHGIAICHMDTSAWGANHGAFMLLGSRPGQIEVCHWI 611
Query: 299 PQDHVVW 305
++ + W
Sbjct: 612 FENDMNW 618
>At3g11590 unknown protein
Length = 622
Score = 30.0 bits (66), Expect = 1.7
Identities = 25/89 (28%), Positives = 37/89 (41%), Gaps = 2/89 (2%)
Query: 146 IKEGSKEAEIVKRTISECEANGIKGEEKLCVTSL--ESMVDFTISKLGNNVEAVSTEVDK 203
I E E E +KR + + K E L + E V +S+ + +E + VDK
Sbjct: 374 ISEDKAEVEELKRESFKVKEEVEKEREMLQLADALREERVQMKLSEAKHQLEEKNAAVDK 433
Query: 204 NSNGLQQYVIAKGVKKLGEKNKTIVCHKE 232
N LQ Y+ AK K+ + H E
Sbjct: 434 LRNQLQTYLKAKRCKEKTREPPQTQLHNE 462
>At3g02930 unknown protein
Length = 806
Score = 29.6 bits (65), Expect = 2.2
Identities = 18/93 (19%), Positives = 39/93 (41%), Gaps = 5/93 (5%)
Query: 105 LNLQFKETTSNNNGTKFLPREVANSIPFS-----SNKMENILNKFSIKEGSKEAEIVKRT 159
+N++ +E T + P + + F + + + +K S KE ++ +V ++
Sbjct: 703 MNMKLEEDTEKKEKKERSPEDETVEVEFKMWESCQIEKKEVFHKESAKEEEEDLNVVDQS 762
Query: 160 ISECEANGIKGEEKLCVTSLESMVDFTISKLGN 192
NG+ GE++L + K+GN
Sbjct: 763 QKTSPVNGLTGEDELLKEKEKKKKKTLFGKVGN 795
>At4g01270 putative RING zinc finger protein
Length = 506
Score = 28.5 bits (62), Expect = 4.8
Identities = 22/77 (28%), Positives = 33/77 (42%), Gaps = 6/77 (7%)
Query: 133 SSNKMENILNKFS-IKEGSKEAEIVKRTISECEANGIKGEEKLCVT-----SLESMVDFT 186
SS K+E L K +K+ +E E++ I +K T ++ESM F
Sbjct: 249 SSEKLEKALEKIEKLKKRMRELELITEERENRALRDINVSKKCSYTEVSEPAIESMSSFR 308
Query: 187 ISKLGNNVEAVSTEVDK 203
+ N VE +ST K
Sbjct: 309 MLSSDNKVEKISTPPGK 325
>At2g22420 putative peroxidase
Length = 329
Score = 28.5 bits (62), Expect = 4.8
Identities = 24/106 (22%), Positives = 44/106 (40%), Gaps = 15/106 (14%)
Query: 118 GTKFLPREVANSIPFSSNKMENILNKFSIKEGSKEAEIVKRTISECEANGIKGEEKLCVT 177
G PR + + P + + + + K IKE A +++ +C NG C
Sbjct: 20 GETLRPRFYSETCPEAESIVRREMKKAMIKEARSVASVMRFQFHDCFVNG-------CDA 72
Query: 178 SLESMVDFTISKLGNNVEAVSTEVDKNSNGLQQYVIAKGVKKLGEK 223
SL ++D T + LG + N + L+ + + +K+ EK
Sbjct: 73 SL--LLDDTPNMLGEKLSL------SNIDSLRSFEVVDDIKEALEK 110
>At1g23160 GH3-like auxin-regulated protein
Length = 578
Score = 28.5 bits (62), Expect = 4.8
Identities = 28/119 (23%), Positives = 53/119 (44%), Gaps = 19/119 (15%)
Query: 169 KGEEKLCVTSLESMVDFTISKLGNNVEAVSTEVDKNSNGLQQYVIAKGVKKLGEKNKTIV 228
K +K+ V LE + ++++ +NV ++ N+N LQ++ + K+ +KN +V
Sbjct: 10 KSSDKMKV--LEDLTS-NVTQIQDNVLEEILTLNANTNYLQKFFLGSFDKESFKKNVPVV 66
Query: 229 CHKENYPYAVFYCHKTDSTEVYSVPLEGVDGNMVKTIAVCHTDTS-------EWNPKHL 280
+++ PY + S + + P+ G V T TS WN K+L
Sbjct: 67 TYEDVKPYIERVVNGEPSNVISARPITGF---------VLSTGTSGGAQKMMPWNEKYL 116
>At5g51660 cleavage and polyadenylation specificity factor subunit
Length = 1448
Score = 28.1 bits (61), Expect = 6.3
Identities = 12/32 (37%), Positives = 19/32 (58%)
Query: 57 TYLNDEKIIPLIYFYPIPIPLNESQIQLDDKQ 88
TY ++ + PLI YP+ PLN+ L D++
Sbjct: 1034 TYYAEKNLYPLIVSYPVSKPLNQVLSSLVDQE 1065
>At5g16720 putative protein
Length = 600
Score = 28.1 bits (61), Expect = 6.3
Identities = 20/88 (22%), Positives = 40/88 (44%), Gaps = 2/88 (2%)
Query: 115 NNNGTKFLPREVANSIPFSSNKMENILNKFSIKEGSKEAEIVKRTISECEANGIKGEEKL 174
+ +G++ LP + S K+E I S+ E +E E + C ++ KG++
Sbjct: 511 SEDGSQGLPESDEKNFGSDSEKLEIIKQVDSVYERLQELETDGEFLKNCMSSAKKGDKGT 570
Query: 175 CVTS--LESMVDFTISKLGNNVEAVSTE 200
+ L+ + D +L N +E +T+
Sbjct: 571 DILKDILQHLRDLRTIELTNTIENQTTQ 598
>At4g03330 SYR1-like syntaxin
Length = 305
Score = 28.1 bits (61), Expect = 6.3
Identities = 25/126 (19%), Positives = 52/126 (40%), Gaps = 2/126 (1%)
Query: 105 LNLQFKETTSNNNGTKFLPREVANSIPFSSNKMENILNKFSIKEGSKEAEIVKRTISECE 164
++ FK T N+ + E N S N E S+KE K + + + + +
Sbjct: 5 ISSSFKRYTDLNHQVQLDDIESQNVSLDSGNLDEFFGYVESVKEDMKAVDEIHKRLQDAN 64
Query: 165 ANGIKGEEKLCVTSLESMVDFTISKLGNNVEAVSTEVD--KNSNGLQQYVIAKGVKKLGE 222
+ V L + +D +++++ V+ + T++ + SN Q+ V G +
Sbjct: 65 EESKTVHDSKAVKKLRARMDSSVTEVLKRVKMIKTKLVALEKSNAAQRKVAGCGPGSSAD 124
Query: 223 KNKTIV 228
+ +T V
Sbjct: 125 RTRTSV 130
>At2g17150 unknown protein
Length = 909
Score = 28.1 bits (61), Expect = 6.3
Identities = 17/63 (26%), Positives = 27/63 (41%)
Query: 123 PREVANSIPFSSNKMENILNKFSIKEGSKEAEIVKRTISECEANGIKGEEKLCVTSLESM 182
PR ++S SS N N +I ++KR SE + + + EE C+ +S
Sbjct: 724 PRSPSSSCSKSSGSSNNNENTGNILVAEDADAVLKRAHSEAQLHNVNQEETKCLARTQSH 783
Query: 183 VDF 185
F
Sbjct: 784 KTF 786
>At1g24150 unknown protein
Length = 725
Score = 28.1 bits (61), Expect = 6.3
Identities = 18/61 (29%), Positives = 30/61 (48%), Gaps = 4/61 (6%)
Query: 143 KFSIKEGSKEAEIVKRTISECEANGIKGEEKL----CVTSLESMVDFTISKLGNNVEAVS 198
K + +E K E+VKRT +A +KG+ L V +MVD ++ N++ S
Sbjct: 630 KLAKEEEKKVLELVKRTTEYYQAGAVKGKNPLHLFVIVRDFLAMVDKVCVEIARNLQRRS 689
Query: 199 T 199
+
Sbjct: 690 S 690
>At5g46130 putative protein
Length = 376
Score = 27.7 bits (60), Expect = 8.2
Identities = 28/124 (22%), Positives = 54/124 (42%), Gaps = 15/124 (12%)
Query: 35 KARDASNGGLDVGVRKGYEGGGTYLNDEKIIPLIYFYPIPIPLNESQIQLDDKQNVTLFF 94
K +SNG + R G EG + K + + + LN+S + + K+N +L
Sbjct: 43 KIGGSSNGSSSISRRSGSEGENEIIIKSKFLAGEVCEAMSMGLNQSGVNIHMKENTSL-- 100
Query: 95 LKKDLHHGTKLNLQFKETTSNNNGTKF----LPR--EVAN----SIPFSSNKMENILNKF 144
+D+ + +LN+ +K + + LP+ ++ N S+P N + KF
Sbjct: 101 --RDMDN-VRLNIYYKPSDPKSKSVSVHLPPLPKGSQIQNIAMTSLPDLKNDDWVVAIKF 157
Query: 145 SIKE 148
S+ +
Sbjct: 158 SVSQ 161
>At3g50270 anthranilate N-hydroxycinnamoyl/benzoyltransferase - like
protein
Length = 450
Score = 27.7 bits (60), Expect = 8.2
Identities = 16/81 (19%), Positives = 38/81 (46%)
Query: 71 YPIPIPLNESQIQLDDKQNVTLFFLKKDLHHGTKLNLQFKETTSNNNGTKFLPREVANSI 130
YPI +P++E++ ++ ++ K+ + H TK N+ + +N+ EV++
Sbjct: 209 YPIHVPVSEAERSPPSREPSSVPITKEWIFHFTKKNISDLKAKANSEIASSSDLEVSSLQ 268
Query: 131 PFSSNKMENILNKFSIKEGSK 151
S++ +I+ + K
Sbjct: 269 AVSAHMWRSIIRHSGVSREQK 289
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.315 0.134 0.399
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,593,676
Number of Sequences: 26719
Number of extensions: 341799
Number of successful extensions: 655
Number of sequences better than 10.0: 17
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 633
Number of HSP's gapped (non-prelim): 20
length of query: 308
length of database: 11,318,596
effective HSP length: 99
effective length of query: 209
effective length of database: 8,673,415
effective search space: 1812743735
effective search space used: 1812743735
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 60 (27.7 bits)
Medicago: description of AC148532.11


