

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
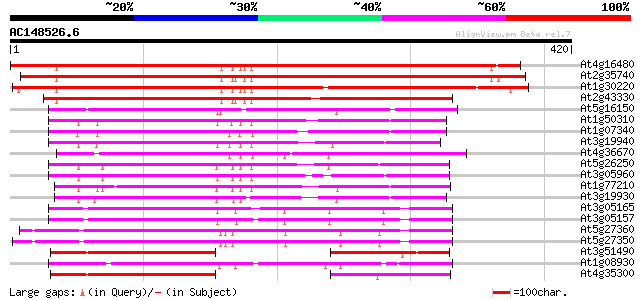
Query= AC148526.6 + phase: 0 /pseudo
(420 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At4g16480 membrane transporter like protein 448 e-126
At2g35740 putative sugar transporter 418 e-117
At1g30220 unknown protein 374 e-104
At2g43330 membrane transporter like protein 262 3e-70
At5g16150 sugar transporter like protein 132 3e-31
At1g50310 hexose transporter, putative 131 6e-31
At1g07340 hexose transporter like protein 130 2e-30
At3g19940 putative monosaccharide transport protein, STP4 129 3e-30
At4g36670 sugar transporter like protein 122 4e-28
At5g26250 hexose transporter - like protein 122 5e-28
At3g05960 putative hexose transporter 117 1e-26
At1g77210 unknown protein 117 2e-26
At3g19930 monosaccharide transport protein, STP4 108 4e-24
At3g05165 sugar transporter like protein 105 5e-23
At3g05157 sugar transporter like protein 101 7e-22
At5g27360 sugar-porter family protein 2 (SFP2) 100 2e-21
At5g27350 sugar transporter like protein 98 7e-21
At3g51490 sugar transporter-like protein 97 2e-20
At1g08930 ERD6 protein 95 8e-20
At4g35300 sugar transporter like protein 93 3e-19
>At4g16480 membrane transporter like protein
Length = 582
Score = 448 bits (1153), Expect = e-126
Identities = 241/420 (57%), Positives = 303/420 (71%), Gaps = 39/420 (9%)
Query: 1 MAEGGHPLADKTEFTECWRRTAESPYLMRLALS---GGLLFGYDTGVISGASLYIRDEFE 57
M EGG ADKTEFTECWR T ++PY+MRLALS GGLLFGYDTGVISGA L+I+++F+
Sbjct: 1 MVEGGIAKADKTEFTECWRTTWKTPYIMRLALSAGIGGLLFGYDTGVISGALLFIKEDFD 60
Query: 58 QVDKKPWLQETIVSMASAGAIIGAAFGGYMNDKMGRKKTILMADVVFVAGALVMAAAPAP 117
+VDKK WLQ TIVSMA AGAI+GAA GG++NDK GR+ +IL+ADV+F+ GA+VMA APAP
Sbjct: 61 EVDKKTWLQSTIVSMAVAGAIVGAAVGGWINDKFGRRMSILIADVLFLIGAIVMAFAPAP 120
Query: 118 WVIIIGRLLVGLGVGAASMTAPLYISEASPAKIRGALVCR--------------LHQWFT 163
WVII+GR+ VG GVG ASMT+PLYISEASPA+IRGALV ++ F
Sbjct: 121 WVIIVGRIFVGFGVGMASMTSPLYISEASPARIRGALVSTNGLLITGGQFFSYLINLAFV 180
Query: 164 HN----RWSISL--LPH--QFGIY--------QVYRQSKEEEAKQILSKIYRPGEVEEEM 207
H RW + + +P QF + +YR+ + E++ IL +IY EVE EM
Sbjct: 181 HTPGTWRWMLGVAGVPAIVQFVLMLSLPESPRWLYRKDRIAESRAILERIYPADEVEAEM 240
Query: 208 KAMHESIEAEKAEDGLIGHSLAQKLKGAWSNDVVRRGLYAGITVQVVQQIVGINTIMYYS 267
+A+ S+EAEKA++ +IG S + KLKGA+ N VVRRGL AGITVQV QQ VGINT+MYYS
Sbjct: 241 EALKLSVEAEKADEAIIGDSFSAKLKGAFGNPVVRRGLAAGITVQVAQQFVGINTVMYYS 300
Query: 268 PTIVQFAGIASNSTAFALSLVTSGLNAVGTIVSMVLIDRFGRRKLMLISLIGIFVSLVTL 327
P+IVQFAG ASN TA ALSL+TSGLNA+G+IVSM+ +DR+GRRKLM+IS+ GI L+ L
Sbjct: 301 PSIVQFAGYASNKTAMALSLITSGLNALGSIVSMMFVDRYGRRKLMIISMFGIIACLIIL 360
Query: 328 SVTFNQAAHHAPSLSIQDSLSFGGNSTCKAYT-----TAPNHLSWNCMQCLHEDCAFCAN 382
+ F+QAA HAP + +S +F N+TC AY AP WNCM+CL +C FCA+
Sbjct: 361 ATVFSQAAIHAPKIDAFESRTFAPNATCSAYAPLAAENAPPS-RWNCMKCLRSECGFCAS 419
>At2g35740 putative sugar transporter
Length = 580
Score = 418 bits (1075), Expect = e-117
Identities = 225/415 (54%), Positives = 289/415 (69%), Gaps = 37/415 (8%)
Query: 9 ADKTEFTECWRRTAESPYLMRLALS---GGLLFGYDTGVISGASLYIRDEFEQVDKKPWL 65
+++ TE W T E+PY+MRLALS GGLLFGY+TGVI+GA LYI++EF +VD K WL
Sbjct: 8 SEQINITEVWTTTWETPYIMRLALSAGIGGLLFGYNTGVIAGALLYIKEEFGEVDNKTWL 67
Query: 66 QETIVSMASAGAIIGAAFGGYMNDKMGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRL 125
QE IVSM AGAI+GAA GG+ NDK GR+ ++L+ADV+F+ GALVM A APWVII+GRL
Sbjct: 68 QEIIVSMTVAGAIVGAAIGGWYNDKFGRRMSVLIADVLFLLGALVMVIAHAPWVIILGRL 127
Query: 126 LVGLGVGAASMTAPLYISEASPAKIRGALVCR--------------LHQWFTHN----RW 167
LVG GVG ASMT+PLYISE SPA+IRGALV ++ F H RW
Sbjct: 128 LVGFGVGMASMTSPLYISEMSPARIRGALVSTNGLLITGGQFLSYLINLAFVHTPGTWRW 187
Query: 168 --SISLLPH--QFGIY--------QVYRQSKEEEAKQILSKIYRPGEVEEEMKAMHESIE 215
+S +P QF + +YR ++ E++ IL +IY VE E+ A+ ES+
Sbjct: 188 MLGVSAIPAIIQFCLMLTLPESPRWLYRNDRKAESRDILERIYPAEMVEAEIAALKESVR 247
Query: 216 AEKAEDGLIGHSLAQKLKGAWSNDVVRRGLYAGITVQVVQQIVGINTIMYYSPTIVQFAG 275
AE A++ +IGH+ + KL+GA SN VVR GL AGITVQV QQ VGINT+MYYSPTI+QFAG
Sbjct: 248 AETADEDIIGHTFSDKLRGALSNPVVRHGLAAGITVQVAQQFVGINTVMYYSPTILQFAG 307
Query: 276 IASNSTAFALSLVTSGLNAVGTIVSMVLIDRFGRRKLMLISLIGIFVSLVTLSVTFNQAA 335
ASN TA AL+L+TSGLNAVG++VSM+ +DR+GRRKLM+IS+ GI LV L+ FN+A+
Sbjct: 308 YASNKTAMALALITSGLNAVGSVVSMMFVDRYGRRKLMIISMFGIITCLVILAAVFNEAS 367
Query: 336 HHAPSLSIQDSLSFGGNSTCKAYT--TAPNH--LSWNCMQCLHEDCAFCANSQNE 386
+HAP + +DS +F N+TC A+ TA +WNCM+CL DC FC+N E
Sbjct: 368 NHAPKIDKRDSRNFAKNATCPAFAPFTASRSPPSNWNCMKCLQYDCGFCSNGAQE 422
>At1g30220 unknown protein
Length = 580
Score = 374 bits (959), Expect = e-104
Identities = 216/424 (50%), Positives = 283/424 (65%), Gaps = 42/424 (9%)
Query: 3 EGG--HPLADKTEFTECWRRTAESPYLMRLALS---GGLLFGYDTGVISGASLYIRDEFE 57
EGG H AD++ F EC+ T ++PY++RLA S GGLLFGYDTGVISGA LYIRD+F+
Sbjct: 2 EGGIIHGGADESAFKECFSLTWKNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFK 61
Query: 58 QVDKKPWLQETIVSMASAGAIIGAAFGGYMNDKMGRKKTILMADVVFVAGALVMAAAPAP 117
VD+ WLQE IVSMA AGAI+GAA GG+ NDK+GR+ ILMAD +F+ GA++MAAAP P
Sbjct: 62 SVDRNTWLQEMIVSMAVAGAIVGAAIGGWANDKLGRRSAILMADFLFLLGAIIMAAAPNP 121
Query: 118 WVIIIGRLLVGLGVGAASMTAPLYISEASPAKIRGALVCR--------------LHQWFT 163
++++GR+ VGLGVG ASMTAPLYISEASPAKIRGALV ++ FT
Sbjct: 122 SLLVVGRVFVGLGVGMASMTAPLYISEASPAKIRGALVSTNGFLITGGQFLSYLINLAFT 181
Query: 164 HN----RWSISL--LPH--QFGIY--------QVYRQSKEEEAKQILSKIYRPGEVEEEM 207
RW + + +P QF + +YR+ +EEEAK IL +IY +VE+E+
Sbjct: 182 DVTGTWRWMLGIAGIPALLQFVLMFTLPESPRWLYRKGREEEAKAILRRIYSAEDVEQEI 241
Query: 208 KAMHESIEAEKAEDGLIGHSLAQKLKGAWSNDVVRRGLYAGITVQVVQQIVGINTIMYYS 267
+A+ +S+E E E+G KL A VRRGL AG+ +QV QQ VGINT+MYYS
Sbjct: 242 RALKDSVETEILEEGSSEKINMIKLCKA---KTVRRGLIAGVGLQVFQQFVGINTVMYYS 298
Query: 268 PTIVQFAGIASNSTAFALSLVTSGLNAVGTIVSMVLIDRFGRRKLMLISLIGIFVSLVTL 327
PTIVQ AG ASN TA LSLVT+GLNA G+I+S+ IDR GR+KL++ISL G+ +SL L
Sbjct: 299 PTIVQLAGFASNRTALLLSLVTAGLNAFGSIISIYFIDRIGRKKLLIISLFGVIISLGIL 358
Query: 328 SVTFNQAAHHAPSLSIQDSLSFGGNSTCKAYTTAPNHLSWNCMQCL---HEDCAFCANSQ 384
+ F +AA HAP++S ++ F N +C Y +A N +W+CM CL C +C++
Sbjct: 359 TGVFYEAATHAPAISSLETQRF-NNISCPDYKSAMNTNAWDCMTCLKASSPSCGYCSSPI 417
Query: 385 NEEH 388
+EH
Sbjct: 418 GKEH 421
>At2g43330 membrane transporter like protein
Length = 509
Score = 262 bits (669), Expect = 3e-70
Identities = 157/339 (46%), Positives = 213/339 (62%), Gaps = 39/339 (11%)
Query: 26 YLMRLALS---GGLLFGYDTGVISGASLYIRDEFEQVDKKPWLQETIVSMASAGAIIGAA 82
Y++ L ++ GGLLFGYDTGVISGA LYI+D+FE V + +LQETIVSMA GA+IGAA
Sbjct: 30 YILGLTVTAGIGGLLFGYDTGVISGALLYIKDDFEVVKQSSFLQETIVSMALVGAMIGAA 89
Query: 83 FGGYMNDKMGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVGAASMTAPLYI 142
GG++ND GRKK L ADVVF AGA+VMAAAP P+V+I GRLLVGLGVG AS+TAP+YI
Sbjct: 90 AGGWINDYYGRKKATLFADVVFAAGAIVMAAAPDPYVLISGRLLVGLGVGVASVTAPVYI 149
Query: 143 SEASPAKIRGALVCR--------------LHQWFTHN----RW--SISLLPH--QFGIY- 179
+EASP+++RG LV ++ FT RW +S +P QF +
Sbjct: 150 AEASPSEVRGGLVSTNVLMITGGQFLSYLVNSAFTQVPGTWRWMLGVSGVPAVIQFILML 209
Query: 180 -------QVYRQSKEEEAKQILSKIYRPGEVEEEMKAMHESIEAEKAEDGLIGHSLAQKL 232
++ ++++ EA Q+L++ Y +E+E+ + + E EK +G+
Sbjct: 210 FMPESPRWLFMKNRKAEAIQVLARTYDISRLEDEIDHLSAAEEEEKQRKRTVGY------ 263
Query: 233 KGAWSNDVVRRGLYAGITVQVVQQIVGINTIMYYSPTIVQFAGIASNSTAFALSLVTSGL 292
+ + +R AG +Q QQ GINT+MYYSPTIVQ AG SN A LSL+ + +
Sbjct: 264 LDVFRSKELRLAFLAGAGLQAFQQFTGINTVMYYSPTIVQMAGFHSNQLALFLSLIVAAM 323
Query: 293 NAVGTIVSMVLIDRFGRRKLMLISLIGIFVSLVTLSVTF 331
NA GT+V + ID GR+KL L SL G+ +SL+ LSV+F
Sbjct: 324 NAAGTVVGIYFIDHCGRKKLALSSLFGVIISLLILSVSF 362
>At5g16150 sugar transporter like protein
Length = 546
Score = 132 bits (333), Expect = 3e-31
Identities = 97/330 (29%), Positives = 163/330 (49%), Gaps = 31/330 (9%)
Query: 30 LALSGGLLFGYDTGVISGASLYIRDEFEQVDKKPWLQETIVSMASAGAIIGAAFGGYMND 89
+A G +LFGY GV++GA Y+ + + + LQ IVS AGA +G+ GG + D
Sbjct: 111 VACLGAILFGYHLGVVNGALEYLAKDLG-IAENTVLQGWIVSSLLAGATVGSFTGGALAD 169
Query: 90 KMGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVGAASMTAPLYISEASPAK 149
K GR +T + + GA + A A + +I+GRLL G+G+G +S PLYISE SP +
Sbjct: 170 KFGRTRTFQLDAIPLAIGAFLCATAQSVQTMIVGRLLAGIGIGISSAIVPLYISEISPTE 229
Query: 150 IRGAL-------VC-------------RLHQWFTHNRWSISLLPHQFGIYQVYRQSKEEE 189
IRGAL +C + + + ++++P + + E
Sbjct: 230 IRGALGSVNQLFICIGILAALIAGLPLAANPLWWRTMFGVAVIP---SVLLAIGMAFSPE 286
Query: 190 AKQILSKIYRPGEVEEEMKAMH-ESIEAEKAEDGLIGHSLAQKLKGAWSNDVVRR---GL 245
+ + L + + E E+ +K ++ + E D + + + W + R +
Sbjct: 287 SPRWLVQQGKVSEAEKAIKTLYGKERVVELVRDLSASGQGSSEPEAGWFDLFSSRYWKVV 346
Query: 246 YAGITVQVVQQIVGINTIMYYSPTIVQFAGIASNSTAFALSLVTSGLNAVGTIVSMVLID 305
G + + QQ+ GIN ++YYS ++ + AGI S+ A AL N GT V+ L+D
Sbjct: 347 SVGAALFLFQQLAGINAVVYYSTSVFRSAGIQSDVAASAL---VGASNVFGTAVASSLMD 403
Query: 306 RFGRRKLMLISLIGIFVSLVTLSVTFNQAA 335
+ GR+ L+L S G+ +S++ LS++F A
Sbjct: 404 KMGRKSLLLTSFGGMALSMLLLSLSFTWKA 433
Score = 30.4 bits (67), Expect = 1.9
Identities = 44/169 (26%), Positives = 74/169 (43%), Gaps = 24/169 (14%)
Query: 65 LQETIVSMASAGA--IIGAAFGGYMNDKMGRKKTILMADVVFVAGALVMA--------AA 114
+Q + + A GA + G A + DKMGRK +L + L+++ AA
Sbjct: 377 IQSDVAASALVGASNVFGTAVASSLMDKMGRKSLLLTSFGGMALSMLLLSLSFTWKALAA 436
Query: 115 PAPWVIIIGRLL--VGLGVGAASMTAPLYISEASPAKIRG---ALVCRLHQW---FTHNR 166
+ + ++G +L + +GA + A L + E ++IR AL +H W F
Sbjct: 437 YSGTLAVVGTVLYVLSFSLGAGPVPA-LLLPEIFASRIRAKAVALSLGMH-WISNFVIGL 494
Query: 167 WSISLLPHQFGIYQVYRQSKEEEAKQILSKIYRPGEVEEEMKAMHESIE 215
+ +S++ +FGI VY +L+ +Y G V E E IE
Sbjct: 495 YFLSVVT-KFGISSVYLGF---AGVCVLAVLYIAGNVVETKGRSLEEIE 539
>At1g50310 hexose transporter, putative
Length = 517
Score = 131 bits (330), Expect = 6e-31
Identities = 101/346 (29%), Positives = 164/346 (47%), Gaps = 57/346 (16%)
Query: 30 LALSGGLLFGYDTGVISGASL---YIRDEFEQVDKKPW--------------LQETIVSM 72
+A GGLLFGYD G+ G + ++ F +VDK+ L + S
Sbjct: 31 VAAMGGLLFGYDLGISGGVTSMEEFLSKFFPEVDKQMHEARRETAYCKFDNQLLQLFTSS 90
Query: 73 ASAGAIIGAAFGGYMNDKMGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVG 132
A+ + + K GRK ++ + V F+ G+L A A ++I+GRLL+G+GVG
Sbjct: 91 LYLAALASSFVASAVTRKYGRKISMFVGGVAFLIGSLFNAFATNVAMLIVGRLLLGVGVG 150
Query: 133 AASMTAPLYISEASPAKIRGAL-------------VCRLHQWFTH----NRWSISL---- 171
A+ + P+Y+SE +PAKIRGAL + L + T N W +SL
Sbjct: 151 FANQSTPVYLSEMAPAKIRGALNIGFQMAITIGILIANLINYGTSQMAKNGWRVSLGLAA 210
Query: 172 LPHQFGIY----------QVYRQSKEEEAKQILSKIYRPGEVEEEMKAMHESIEAEKAED 221
+P + + + K E+A+++L KI V+EE + + ++ EA K D
Sbjct: 211 VPAVIMVIGSFVLPDTPNSMLERGKYEQAREMLQKIRGADNVDEEFQDLCDACEAAKKVD 270
Query: 222 GLIGHSLAQKLKGAWSNDVVRRGLYAGITVQVVQQIVGINTIMYYSPTIVQFAGIASNST 281
K + R L + QQI GIN IM+Y+P + + G A +++
Sbjct: 271 N--------PWKNIFQQAKYRPALVFCSAIPFFQQITGINVIMFYAPVLFKTLGFADDAS 322
Query: 282 AFALSLVTSGLNAVGTIVSMVLIDRFGRRKLMLISLIGIFVSLVTL 327
+ +++T +N V T+VS+ +DR+GRR L L I + VS + +
Sbjct: 323 LIS-AVITGAVNVVSTLVSIYAVDRYGRRILFLEGGIQMIVSQIVV 367
>At1g07340 hexose transporter like protein
Length = 383
Score = 130 bits (326), Expect = 2e-30
Identities = 93/344 (27%), Positives = 161/344 (46%), Gaps = 55/344 (15%)
Query: 30 LALSGGLLFGYDTGVISGAS---LYIRDEFEQVDKKPW-------------LQETIVSMA 73
+A GGL+FGYD G+ G + ++ D F V +K L + S
Sbjct: 29 IAAVGGLMFGYDIGISGGVTSMDTFLLDFFPHVYEKKHRVHENNYCKFDDQLLQLFTSSL 88
Query: 74 SAGAIIGAAFGGYMNDKMGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVGA 133
I + Y++ GRK TI++A + F+ GA++ +A ++I GR+L+G G+G
Sbjct: 89 YLAGIFASFISSYVSRAFGRKPTIMLASIFFLVGAILNLSAQELGMLIGGRILLGFGIGF 148
Query: 134 ASMTAPLYISEASPAKIRGALVCRLHQWFT----------------HNRWSIS------- 170
+ T PL+ISE +PA+ RG L T N W S
Sbjct: 149 GNQTVPLFISEIAPARYRGGLNVMFQFLITIGILAASYVNYLTSTLKNGWRYSLGGAAVP 208
Query: 171 ---LLPHQFGIYQ----VYRQSKEEEAKQILSKIYRPGEVEEEMKAMHESIEAEKAEDGL 223
LL F I++ + + K+E+ KQ+L KI ++E E + + E
Sbjct: 209 ALILLIGSFFIHETPASLIERGKDEKGKQVLRKIRGIEDIELEFNEIKYATE-------- 260
Query: 224 IGHSLAQKLKGAWSNDVVRRGLYAGITVQVVQQIVGINTIMYYSPTIVQFAGIASNSTAF 283
+ + K ++ R L G +Q QQ GIN +M+Y+P + Q G N++
Sbjct: 261 VATKVKSPFKELFTKSENRPPLVCGTLLQFFQQFTGINVVMFYAPVLFQTMGSGDNASLI 320
Query: 284 ALSLVTSGLNAVGTIVSMVLIDRFGRRKLMLISLIGIFVSLVTL 327
+ ++VT+G+NA+ T++S++++D GRR L++ + + + +T+
Sbjct: 321 S-TVVTNGVNAIATVISLLVVDFAGRRCLLMEGALQMTATQMTI 363
>At3g19940 putative monosaccharide transport protein, STP4
Length = 514
Score = 129 bits (324), Expect = 3e-30
Identities = 101/344 (29%), Positives = 163/344 (47%), Gaps = 64/344 (18%)
Query: 30 LALSGGLLFGYDTGVISGASL---YIRDEFEQVDKKP--------------WLQETIVSM 72
+A GGLLFGYD G+ G + ++ F QV+ + + + S
Sbjct: 31 VAAMGGLLFGYDLGISGGVTSMEEFLTKFFPQVESQMKKAKHDTAYCKFDNQMLQLFTSS 90
Query: 73 ASAGAIIGAAFGGYMNDKMGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVG 132
A++ + + K GRK ++ + + F+ GAL A A ++IIGRLL+G+GVG
Sbjct: 91 LYLAALVASFMASVITRKHGRKVSMFIGGLAFLIGALFNAFAVNVSMLIIGRLLLGVGVG 150
Query: 133 AASMTAPLYISEASPAKIRGAL-------------VCRLHQWFT----HNRWSISL---- 171
A+ + P+Y+SE +PAKIRGAL V L + T + W +SL
Sbjct: 151 FANQSTPVYLSEMAPAKIRGALNIGFQMAITIGILVANLINYGTSKMAQHGWRVSLGLAA 210
Query: 172 ---LPHQFGIY-------QVYRQSKEEEAKQILSKIYRPGEVEEEMKAMHESIEAEKAED 221
+ G + + + K EEAKQ+L KI V+ E + + +++EA
Sbjct: 211 VPAVVMVIGSFILPDTPNSMLERGKNEEAKQMLKKIRGADNVDHEFQDLIDAVEA----- 265
Query: 222 GLIGHSLAQKLKGAWSNDV---VRRGLYAGITVQVVQQIVGINTIMYYSPTIVQFAGIAS 278
A+K++ W N + R L + QQI GIN IM+Y+P + + G
Sbjct: 266 -------AKKVENPWKNIMESKYRPALIFCSAIPFFQQITGINVIMFYAPVLFKTLGFGD 318
Query: 279 NSTAFALSLVTSGLNAVGTIVSMVLIDRFGRRKLMLISLIGIFV 322
++ A +++T +N + T VS+ +DR+GRR L L I +F+
Sbjct: 319 DA-ALMSAVITGVVNMLSTFVSIYAVDRYGRRLLFLEGGIQMFI 361
>At4g36670 sugar transporter like protein
Length = 493
Score = 122 bits (306), Expect = 4e-28
Identities = 94/349 (26%), Positives = 162/349 (45%), Gaps = 46/349 (13%)
Query: 36 LLFGYDTGVISGASLYIRDEFEQVDKKPWLQETIVSMASAGAIIGAAFGGYMNDKMGRKK 95
++FGYDTGV+SGA ++I ++ + D + E + + + A++G+ G +D +GR+
Sbjct: 29 IIFGYDTGVMSGAMVFIEEDLKTNDVQI---EVLTGILNLCALVGSLLAGRTSDIIGRRY 85
Query: 96 TILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVGAASMTAPLYISEASPAKIRGALV 155
TI++A ++F+ G+++M P V++ GR GLGVG A M AP+Y +E + A RG L
Sbjct: 86 TIVLASILFMLGSILMGWGPNYPVLLSGRCTAGLGVGFALMVAPVYSAEIATASHRGLLA 145
Query: 156 CRLHQWFT------------------HNRWSISL-------LPHQFGIYQ-------VYR 183
H + H W + L L FGI + +
Sbjct: 146 SLPHLCISIGILLGYIVNYFFSKLPMHIGWRLMLGIAAVPSLVLAFGILKMPESPRWLIM 205
Query: 184 QSKEEEAKQILSKIYR-PGEVE---EEMKAMHESIEAEKAEDGLIGHSLAQKLKGAWS-- 237
Q + +E K+IL + P E E +++KA I+ + +D + +G W
Sbjct: 206 QGRLKEGKEILELVSNSPEEAELRFQDIKAA-AGIDPKCVDDVVKMEGKKTHGEGVWKEL 264
Query: 238 ----NDVVRRGLYAGITVQVVQQIVGINTIMYYSPTIVQFAGIASNSTAFALSLVTSGLN 293
VRR L + + Q GI ++ Y P I + AGI + F +++ +
Sbjct: 265 ILRPTPAVRRVLLTALGIHFFQHASGIEAVLLYGPRIFKKAGITTKDKLFLVTIGVGIMK 324
Query: 294 AVGTIVSMVLIDRFGRRKLMLISLIGIFVSLVTLSVTFNQAAHHAPSLS 342
+ +L+D+ GRRKL+L S+ G+ ++L L A + L+
Sbjct: 325 TTFIFTATLLLDKVGRRKLLLTSVGGMVIALTMLGFGLTMAQNAGGKLA 373
>At5g26250 hexose transporter - like protein
Length = 507
Score = 122 bits (305), Expect = 5e-28
Identities = 90/350 (25%), Positives = 166/350 (46%), Gaps = 63/350 (18%)
Query: 30 LALSGGLLFGYDTGVISGASL---YIRDEFEQV-DKKPWLQET------------IVSMA 73
+A GGL+FGYD G+ G + ++++ F V ++K E S
Sbjct: 28 IAAVGGLIFGYDIGISGGVTAMDDFLKEFFPSVYERKKHAHENNYCKYDNQFLQLFTSSL 87
Query: 74 SAGAIIGAAFGGYMNDKMGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVGA 133
A++ + F K+GR+ T+ +A + F+ G + A A +++IIGR+L+G GVG
Sbjct: 88 YLAALVASFFASATCSKLGRRPTMQLASIFFLIGVGLAAGAVNIYMLIIGRILLGFGVGF 147
Query: 134 ASMTAPLYISEASPAKIRGA-------------LVCRLHQWFTHN----RWSISL----L 172
+ PL++SE +PA++RG L+ + +FT + W I+L +
Sbjct: 148 GNQAVPLFLSEIAPARLRGGLNIVFQLMVTIGILIANIVNYFTSSIHPYGWRIALGGAGI 207
Query: 173 PHQFGIY----------QVYRQSKEEEAKQILSKIYRPGEVEEEMKAMHESIEAEKAEDG 222
P ++ + ++K +E K+ L KI +V+EE +++ + +
Sbjct: 208 PALILLFGSLLICETPTSLIERNKTKEGKETLKKIRGVEDVDEEYESIVHACD------- 260
Query: 223 LIGHSLAQKLKGAWS---NDVVRRGLYAGITVQVVQQIVGINTIMYYSPTIVQFAGIASN 279
+A+++K ++ R G+ +Q QQ GIN IM+Y+P + Q G N
Sbjct: 261 -----IARQVKDPYTKLMKPASRPPFVIGMLLQFFQQFTGINAIMFYAPVLFQTVGF-GN 314
Query: 280 STAFALSLVTSGLNAVGTIVSMVLIDRFGRRKLMLISLIGIFVSLVTLSV 329
A ++VT +N + T V + L+D+ GRR L+L S + + + + + +
Sbjct: 315 DAALLSAVVTGTINVLSTFVGIFLVDKTGRRFLLLQSSVHMLICQLVIGI 364
>At3g05960 putative hexose transporter
Length = 507
Score = 117 bits (293), Expect = 1e-26
Identities = 93/348 (26%), Positives = 163/348 (46%), Gaps = 59/348 (16%)
Query: 30 LALSGGLLFGYDTGVISGASL---YIRDEFEQV-DKKPWLQET------------IVSMA 73
+A GGL+FGYD G+ G S ++++ F V ++K + E S
Sbjct: 27 IAAVGGLIFGYDIGISGGVSAMDDFLKEFFPAVWERKKHVHENNYCKYDNQFLQLFTSSL 86
Query: 74 SAGAIIGAAFGGYMNDKMGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVGA 133
A++ + K+GR+ T+ A + F+ G + A A ++IIGRL +G GVG
Sbjct: 87 YLAALVASFVASATCSKLGRRPTMQFASIFFLIGVGLTAGAVNLVMLIIGRLFLGFGVGF 146
Query: 134 ASMTAPLYISEASPAKIRGAL-------------VCRLHQWFTHN----RWSISL----L 172
+ PL++SE +PA++RG L + + +FT W I+L +
Sbjct: 147 GNQAVPLFLSEIAPAQLRGGLNIVFQLMVTIGILIANIVNYFTATVHPYGWRIALGGAGI 206
Query: 173 PHQFGIY----------QVYRQSKEEEAKQILSKIYRPGEVEEEMKAM-HESIEAEKAED 221
P ++ + ++K EE K+ L KI ++ +E +++ H A + +D
Sbjct: 207 PAVILLFGSLLIIETPTSLIERNKNEEGKEALRKIRGVDDINDEYESIVHACDIASQVKD 266
Query: 222 GLIGHSLAQKLKGAWSNDVVRRGLYAGITVQVVQQIVGINTIMYYSPTIVQFAGIASNST 281
+ LK A R G+ +Q+ QQ GIN IM+Y+P + Q G S++
Sbjct: 267 -----PYRKLLKPA-----SRPPFIIGMLLQLFQQFTGINAIMFYAPVLFQTVGFGSDA- 315
Query: 282 AFALSLVTSGLNAVGTIVSMVLIDRFGRRKLMLISLIGIFVSLVTLSV 329
A +++T +N + T V + L+DR GRR L+L S + + + + + +
Sbjct: 316 ALLSAVITGSINVLATFVGIYLVDRTGRRFLLLQSSVHMLICQLIIGI 363
>At1g77210 unknown protein
Length = 504
Score = 117 bits (292), Expect = 2e-26
Identities = 101/351 (28%), Positives = 165/351 (46%), Gaps = 69/351 (19%)
Query: 34 GGLLFGYDTGVISGASL---YIRDEFEQVDKKPW--LQET-------------IVSMASA 75
GG LFGYD GV G + ++++ F + K+ L ET S+ A
Sbjct: 36 GGSLFGYDLGVSGGVTSMDDFLKEFFPGIYKRKQMHLNETDYCKYDNQILTLFTSSLYFA 95
Query: 76 GAIIGAAFGG-YMNDKMGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVGAA 134
G I + FG Y+ GR+ +IL+ V F G ++ AAA ++I+GR+ +G+G+G
Sbjct: 96 GLI--STFGASYVTRIYGRRGSILVGSVSFFLGGVINAAAKNILMLILGRIFLGIGIGFG 153
Query: 135 SMTAPLYISEASPAKIRGA-------------LVCRLHQWFTHN----RWSISL------ 171
+ PLY+SE +PAKIRG LV L + T W +SL
Sbjct: 154 NQAVPLYLSEMAPAKIRGTVNQLFQLTTCIGILVANLINYKTEQIHPWGWRLSLGLATVP 213
Query: 172 --LPHQFGIY------QVYRQSKEEEAKQILSKIYRPGEVEEEMKAMHESIEAEKAEDGL 223
L G+ + Q K E+AK +L K+ +E E + + E+ +A +A
Sbjct: 214 AILMFLGGLVLPETPNSLVEQGKLEKAKAVLIKVRGTNNIEAEFQDLVEASDAARA---- 269
Query: 224 IGHSLAQKLKGAWSNDVVRRG----LYAGITVQVVQQIVGINTIMYYSPTIVQFAGIASN 279
+K + N + RR + I + QQ+ G+N+I++Y+P + Q G +
Sbjct: 270 --------VKNPFRNLLARRNRPQLVIGAIGLPAFQQLTGMNSILFYAPVMFQSLGFGGS 321
Query: 280 STAFALSLVTSGLNAVGTIVSMVLIDRFGRRKLMLISLIGIFVSLVTLSVT 330
++ + S +T+ V I+SM D+FGRR L+L + + +F +V + VT
Sbjct: 322 ASLIS-STITNAALVVAAIMSMYSADKFGRRFLLLEASVEMFCYMVVVGVT 371
>At3g19930 monosaccharide transport protein, STP4
Length = 514
Score = 108 bits (271), Expect = 4e-24
Identities = 93/347 (26%), Positives = 151/347 (42%), Gaps = 68/347 (19%)
Query: 34 GGLLFGYDTGVISGASL---YIRDEFEQVDKK-------------PWLQETIVSMASAGA 77
GGL+FGYD G+ G + ++ + F V KK L S A
Sbjct: 33 GGLIFGYDLGISGGVTSMEPFLEEFFPYVYKKMKSAHENEYCRFDSQLLTLFTSSLYVAA 92
Query: 78 IIGAAFGGYMNDKMGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVGAASMT 137
++ + F + GRK ++ + F G+ A +++IGR+L+G GVG A+ +
Sbjct: 93 LVSSLFASTITRVFGRKWSMFLGGFTFFIGSAFNGFAQNIAMLLIGRILLGFGVGFANQS 152
Query: 138 APLYISEASPAKIRGA-------------LVCRLHQWFTHNR-----WSISL-------- 171
P+Y+SE +P +RGA +V + +FT W ISL
Sbjct: 153 VPVYLSEMAPPNLRGAFNNGFQVAIIFGIVVATIINYFTAQMKGNIGWRISLGLACVPAV 212
Query: 172 --------LPHQFGIYQVYRQSKEEEAKQILSKIYRPGEVEEEMKAMHESIEAEKAEDGL 223
LP + + EEAK++L I EV+EE + + ++ E K
Sbjct: 213 MIMIGALILPDTPN--SLIERGYTEEAKEMLQSIRGTNEVDEEFQDLIDASEESK----- 265
Query: 224 IGHSLAQKLKGAWSNDVVRR---GLYAGITVQVVQQIVGINTIMYYSPTIVQFAGIASNS 280
++K W N ++ R L + QQ+ GIN I +Y+P + Q G S +
Sbjct: 266 -------QVKHPWKNIMLPRYRPQLIMTCFIPFFQQLTGINVITFYAPVLFQTLGFGSKA 318
Query: 281 TAFALSLVTSGLNAVGTIVSMVLIDRFGRRKLMLISLIGIFVSLVTL 327
+ + ++VT + + T VS+ +DRFGRR L L I + VS + +
Sbjct: 319 SLLS-AMVTGIIELLCTFVSVFTVDRFGRRILFLQGGIQMLVSQIAI 364
>At3g05165 sugar transporter like protein
Length = 467
Score = 105 bits (262), Expect = 5e-23
Identities = 89/328 (27%), Positives = 151/328 (45%), Gaps = 41/328 (12%)
Query: 30 LALSGGLLFGYDTGVISGASLYIRDEFEQVDKKPWLQETIVSMASAGAIIGAAFGGYMND 89
+A+ +G G SGA I E +D S + G +GA F G +
Sbjct: 36 VAVCSAFSYGCAAGYTSGAETAIMKE---LDLSMAQFSAFGSFLNVGGAVGALFSGQLAV 92
Query: 90 KMGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVGAASMTAPLYISEASPAK 149
+GR++T+ D V G L +A A + + +GR+ +G+GVG S P+YI+E +P
Sbjct: 93 ILGRRRTLWACDFFCVFGWLSIAFAKNVFWLDLGRISLGIGVGLISYVVPVYIAEITPKH 152
Query: 150 IRGAL-----------VCRLHQWFTHNRW-------SISLLPHQFGIYQVYRQSKEEEAK 191
+RGA V ++ + T W +I + GI+ + E+
Sbjct: 153 VRGAFTASNQLLQNSGVSLIYFFGTVINWRVMAVIGAIPCILQTIGIFFI------PESP 206
Query: 192 QILSKIYRPGEVE---EEMKAMHESIEAEKAEDGLIGHSLAQKLKGAWSN---DVVRRGL 245
+ L+KI EVE ++ + E AE ++ L + K ++S+ RR L
Sbjct: 207 RWLAKIRLSKEVESSLHRLRGKDTDVSGEAAEIQVMTKMLEEDSKSSFSDMFQKKYRRTL 266
Query: 246 YAGITVQVVQQIVGINTIMYYSPTIVQFAGIAS--NSTAFALSLVTSGLNAVGTIVSMVL 303
GI + ++QQ+ G + I YYS I + AG + S F + ++ L V ++L
Sbjct: 267 VVGIGLMLIQQLSGASGITYYSNAIFRKAGFSERLGSMIFGVFVIPKAL------VGLIL 320
Query: 304 IDRFGRRKLMLISLIGIFVSLVTLSVTF 331
+DR+GRR L+L S +G+ + + + V+F
Sbjct: 321 VDRWGRRPLLLASAVGMSIGSLLIGVSF 348
>At3g05157 sugar transporter like protein
Length = 458
Score = 101 bits (252), Expect = 7e-22
Identities = 87/325 (26%), Positives = 149/325 (45%), Gaps = 35/325 (10%)
Query: 30 LALSGGLLFGYDTGVISGASLYIRDEFEQVDKKPWLQETIVSMASAGAIIGAAFGGYMND 89
+A+ +G G SGA I E +D S + G +GA F G +
Sbjct: 27 VAVCSSFSYGCANGYTSGAETAIMKE---LDLSMAQFSAFGSFLNLGGAVGALFSGQLAV 83
Query: 90 KMGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVGAASMTAPLYISEASPAK 149
+GR++T+ D+ + G L +A A + +GR+ +G+GVG S P+YI+E +P
Sbjct: 84 ILGRRRTLWACDLFCIFGWLSIAFAKNVLWLDLGRISLGIGVGLTSYVVPVYIAEITPKH 143
Query: 150 IRGAL-----------VCRLHQWFTHNRWS----ISLLPHQFGIYQVYRQSKEEEAKQIL 194
+RGA + ++ + T W I LP + +Y E+ + L
Sbjct: 144 VRGAFSASTLLLQNSGISLIYFFGTVINWRVLAVIGALPCFIPVIGIY---FIPESPRWL 200
Query: 195 SKIYRPGEVE---EEMKAMHESIEAEKAEDGLIGHSLAQKLKGAWSN---DVVRRGLYAG 248
+KI EVE ++ + E AE ++ L + K ++ + RR L G
Sbjct: 201 AKIGSVKEVENSLHRLRGKDADVSDEAAEIQVMTKMLEEDSKSSFCDMFQKKYRRTLVVG 260
Query: 249 ITVQVVQQIVGINTIMYYSPTIVQFAGIAS--NSTAFALSLVTSGLNAVGTIVSMVLIDR 306
I + ++QQ+ G + I YYS I + AG + S F + ++ L V ++L+DR
Sbjct: 261 IGLMLIQQLSGASGITYYSNAIFRKAGFSERLGSMIFGVFVIPKAL------VGLILVDR 314
Query: 307 FGRRKLMLISLIGIFVSLVTLSVTF 331
+GRR L+L S +G+ + + + V+F
Sbjct: 315 WGRRPLLLASAVGMSIGSLLIGVSF 339
>At5g27360 sugar-porter family protein 2 (SFP2)
Length = 478
Score = 100 bits (249), Expect = 2e-21
Identities = 91/344 (26%), Positives = 166/344 (47%), Gaps = 30/344 (8%)
Query: 8 LADKTEFTECWRRTAESPYLMRLALSGGLLFGYDTGVISGASLYIRDEFEQVDKKPWLQE 67
L ++ + +EC R TA +A+ G FG G SGA + I + +D
Sbjct: 20 LKNQNDDSEC-RITACVILSTFIAVCGSFSFGVSLGYTSGAEIGI---MKDLDLSIAQFS 75
Query: 68 TIVSMASAGAIIGAAFGGYMNDKMGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLV 127
S+++ GA IGA F G M +GR+KT+ ++D++ + G +A A + GR+
Sbjct: 76 AFASLSTLGAAIGALFSGKMAIILGRRKTMWVSDLLCIIGWFSIAFAKDVMWLNFGRISS 135
Query: 128 GLGVGAASMTAPLYISEASPAKIRGALVC--RLHQ-------WFTHN--RWSI-SLLPHQ 175
G+G+G S P+YI+E SP +RG +L Q +F+ N W I +LL
Sbjct: 136 GIGLGLISYVVPVYIAEISPKHVRGTFTFTNQLLQNSGLAMVYFSGNFLNWRILALLGAL 195
Query: 176 FGIYQVYRQSKEEEAKQILSKIYRPGEVEE---EMKAMHESIEAEKAEDGLIGHSLAQKL 232
QV E+ + L+K+ E+E ++ + I E ++ ++ +
Sbjct: 196 PCFIQVIGLFFVPESPRWLAKVGSDKELENSLLRLRGGNADISREASDIEVMTKMVENDS 255
Query: 233 KGAWSNDVVRRGLY---AGITVQVVQQIVGINTIMYYSPTIVQFAG--IASNSTAFALSL 287
K ++ + R+ Y GI + ++QQ G + ++ Y+ TI++ AG + ST L +
Sbjct: 256 KSSFCDLFQRKYRYTLVVGIGLMLIQQFSGSSAVLSYASTILRKAGFSVTIGSTLLGLFM 315
Query: 288 VTSGLNAVGTIVSMVLIDRFGRRKLMLISLIGIFVSLVTLSVTF 331
+ + + ++L+D++GRR L+L S+ G+ ++ + + V F
Sbjct: 316 IPKAM------IGVILVDKWGRRPLLLTSVSGMCITSMLIGVAF 353
>At5g27350 sugar transporter like protein
Length = 474
Score = 98.2 bits (243), Expect = 7e-21
Identities = 93/349 (26%), Positives = 165/349 (46%), Gaps = 30/349 (8%)
Query: 3 EGGHPLADKTEFTECWRRTAESPYLMRLALSGGLLFGYDTGVISGASLYIRDEFEQVDKK 62
EG L +K + +EC R TA +A+ G FG TG SGA + + +D
Sbjct: 11 EGLLQLKNKNDDSEC-RITACVILSTFVAVCGSFSFGVATGYTSGAETGV---MKDLDLS 66
Query: 63 PWLQETIVSMASAGAIIGAAFGGYMNDKMGRKKTILMADVVFVAGALVMAAAPAPWVIII 122
S A+ GA IGA F G + +GR+ T+ ++D + + G L +A A ++
Sbjct: 67 IAQFSAFGSFATLGAAIGALFCGNLAMVIGRRGTMWVSDFLCITGWLSIAFAKEVVLLNF 126
Query: 123 GRLLVGLGVGAASMTAPLYISEASPAKIRGALVC--RLHQ-------WFTHN--RW-SIS 170
GR++ G+G G S P+YI+E +P +RG +L Q +F N W +++
Sbjct: 127 GRIISGIGFGLTSYVVPVYIAEITPKHVRGTFTFSNQLLQNAGLAMIYFCGNFITWRTLA 186
Query: 171 LLPHQFGIYQVYRQSKEEEAKQILSKIYRPGEVEE---EMKAMHESIEAEKAEDGLIGHS 227
LL QV E+ + L+K+ E+E ++ I E +E ++
Sbjct: 187 LLGALPCFIQVIGLFFVPESPRWLAKVGSDKELENSLFRLRGRDADISREASEIQVMTKM 246
Query: 228 LAQKLKGAWSNDVVRRGLY---AGITVQVVQQIVGINTIMYYSPTIVQFAG--IASNSTA 282
+ K ++S+ R+ Y GI + ++QQ G ++ Y+ TI + AG +A +T
Sbjct: 247 VENDSKSSFSDLFQRKYRYTLVVGIGLMLIQQFSGSAAVISYASTIFRKAGFSVAIGTTM 306
Query: 283 FALSLVTSGLNAVGTIVSMVLIDRFGRRKLMLISLIGIFVSLVTLSVTF 331
+ ++ + + ++L+D++GRR L++ S G+ ++ + L V F
Sbjct: 307 LGIFVIPKAM------IGLILVDKWGRRPLLMTSAFGMSMTCMLLGVAF 349
>At3g51490 sugar transporter-like protein
Length = 729
Score = 96.7 bits (239), Expect = 2e-20
Identities = 47/124 (37%), Positives = 81/124 (64%), Gaps = 1/124 (0%)
Query: 31 ALSGGLLFGYDTGVISGASLYIRDEFEQVDKKPWLQETIVSMASAGAIIGAAFGGYMNDK 90
A G +L G+D I+GA +YI+ EF ++K+P ++ IV+M+ GA + F G ++DK
Sbjct: 11 AAIGNMLQGWDNATIAGAVIYIKKEFH-LEKEPKIEGLIVAMSLIGATLITTFSGPVSDK 69
Query: 91 MGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVGAASMTAPLYISEASPAKI 150
+GR+ ++++ V++ ++VM +P +V++ RLL G G+G A P+YISE +P++I
Sbjct: 70 VGRRSMLILSSVLYFLSSIVMFWSPNVYVLLFARLLDGFGIGLAVTLVPIYISETAPSEI 129
Query: 151 RGAL 154
RG L
Sbjct: 130 RGLL 133
Score = 55.1 bits (131), Expect = 7e-08
Identities = 32/91 (35%), Positives = 56/91 (61%), Gaps = 3/91 (3%)
Query: 241 VRRGLYAGITVQVVQQIVGINTIMYYSPTIVQFAGIASNSTAFALSLVTSGL--NAVGTI 298
V+R L G+ +Q++QQ GIN +MYY+P I++ G++S T +S ++ L +A+ T+
Sbjct: 508 VKRALMVGVGLQILQQFAGINGVMYYTPQILEETGVSSLLTNLGISAESASLLISALTTL 567
Query: 299 VSMVLIDRFGRRKLMLISLIGIFVSLVTLSV 329
+ + I R LML ++ + +SLVTL +
Sbjct: 568 LMLPCI-LVSMRSLMLSTIPILILSLVTLVI 597
>At1g08930 ERD6 protein
Length = 496
Score = 94.7 bits (234), Expect = 8e-20
Identities = 83/323 (25%), Positives = 151/323 (46%), Gaps = 31/323 (9%)
Query: 30 LALSGGLLFGYDTGVISGASLYIRDEFEQVDKKPWLQETIVSMASAGAIIGAAFGGYMND 89
+A+SG G G SGA I + + + +I+++ G +IGA F G + D
Sbjct: 64 VAVSGSFCTGCGVGFSSGAQAGITKDLSLSVAEYSMFGSILTL---GGLIGAVFSGKVAD 120
Query: 90 KMGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVGAASMTAPLYISEASPAK 149
+GRK+T+L + + G L +A A + GRLL+G+GVG S P+YI+E +P
Sbjct: 121 VLGRKRTMLFCEFFCITGWLCVALAQNAMWLDCGRLLLGIGVGIFSYVIPVYIAEIAPKH 180
Query: 150 IRGALV--------CRLHQWFTHNRW-------SISLLPHQFGIYQVYRQSKEEEAKQIL 194
+RG+ V C + +F + + L+P F ++ ++ E+ + L
Sbjct: 181 VRGSFVFANQLMQNCGISLFFIIGNFIPWRLLTVVGLVPCVFHVFCLF---FIPESPRWL 237
Query: 195 SKIYRPGEVEEEMKAMHESIEAEKAEDGLIGHSLAQKLKGA---WSNDVVRRGLY---AG 248
+K+ R E ++ + S E I ++ G S RR Y G
Sbjct: 238 AKLGRDKECRSSLQRLRGSDVDISREANTIRDTIDMTENGGETKMSELFQRRYAYPLIIG 297
Query: 249 ITVQVVQQIVGINTIMYYSPTIVQFAGIASNSTAFALSLVTSGLNAVGTIVSMVLIDRFG 308
+ + +QQ+ G + + YY+ ++ G S A S++ + + +++ VL+D+ G
Sbjct: 298 VGLMFLQQLCGSSGVTYYASSLFNKGGFPS---AIGTSVIAT-IMVPKAMLATVLVDKMG 353
Query: 309 RRKLMLISLIGIFVSLVTLSVTF 331
RR L++ S + +S + LSV++
Sbjct: 354 RRTLLMASCSAMGLSALLLSVSY 376
>At4g35300 sugar transporter like protein
Length = 739
Score = 92.8 bits (229), Expect = 3e-19
Identities = 48/124 (38%), Positives = 77/124 (61%), Gaps = 1/124 (0%)
Query: 31 ALSGGLLFGYDTGVISGASLYIRDEFEQVDKKPWLQETIVSMASAGAIIGAAFGGYMNDK 90
A G LL G+D I+GA LYI+ EF ++ P ++ IV+M+ GA + G + D
Sbjct: 11 AAVGNLLQGWDNATIAGAVLYIKKEFN-LESNPSVEGLIVAMSLIGATLITTCSGGVADW 69
Query: 91 MGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVGAASMTAPLYISEASPAKI 150
+GR+ ++++ +++ G+LVM +P +V+++GRLL G GVG P+YISE +P +I
Sbjct: 70 LGRRPMLILSSILYFVGSLVMLWSPNVYVLLLGRLLDGFGVGLVVTLVPIYISETAPPEI 129
Query: 151 RGAL 154
RG L
Sbjct: 130 RGLL 133
Score = 58.9 bits (141), Expect = 5e-09
Identities = 33/99 (33%), Positives = 59/99 (59%), Gaps = 9/99 (9%)
Query: 241 VRRGLYAGITVQVVQQIVGINTIMYYSPTIVQFA---------GIASNSTAFALSLVTSG 291
V+R L G+ +Q++QQ GIN ++YY+P I++ A GI+S+S + +S +T+
Sbjct: 514 VKRALVVGVGLQILQQFSGINGVLYYTPQILEQAGVGILLSNMGISSSSASLLISALTTF 573
Query: 292 LNAVGTIVSMVLIDRFGRRKLMLISLIGIFVSLVTLSVT 330
+ V+M L+D GRR L+L ++ + SL+ L ++
Sbjct: 574 VMLPAIAVAMRLMDLSGRRTLLLTTIPILIASLLVLVIS 612
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.321 0.134 0.399
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,582,097
Number of Sequences: 26719
Number of extensions: 344485
Number of successful extensions: 1327
Number of sequences better than 10.0: 89
Number of HSP's better than 10.0 without gapping: 67
Number of HSP's successfully gapped in prelim test: 22
Number of HSP's that attempted gapping in prelim test: 1092
Number of HSP's gapped (non-prelim): 158
length of query: 420
length of database: 11,318,596
effective HSP length: 102
effective length of query: 318
effective length of database: 8,593,258
effective search space: 2732656044
effective search space used: 2732656044
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 61 (28.1 bits)
Medicago: description of AC148526.6


