

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
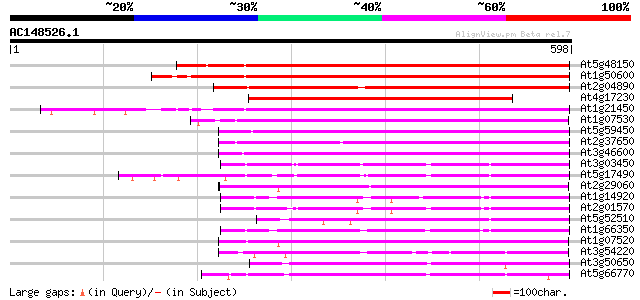
Query= AC148526.1 + phase: 0
(598 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At5g48150 SCARECROW gene regulator-like 467 e-132
At1g50600 scarecrow-like 5 (SCL5) 439 e-123
At2g04890 putative SCARECROW gene regulator 435 e-122
At4g17230 scarecrow-like 13 (SCL13) 391 e-109
At1g21450 scarecrow-like 1 (SCL1) 355 4e-98
At1g07530 transcription factor scarecrow-like 14, putative 211 7e-55
At5g59450 putative SCARECROW transcriptional regulatory protein ... 206 4e-53
At2g37650 scarecrow-like 9 (SCL9) 205 5e-53
At3g46600 scarecrow-like protein 203 2e-52
At3g03450 RGA1-like protein 200 2e-51
At5g17490 RGA-like protein 192 6e-49
At2g29060 putative SCARECROW gene regulator 192 6e-49
At1g14920 signal response protein (GAI) 186 3e-47
At2g01570 putative RGA1, giberellin repsonse modulation protein 182 4e-46
At5g52510 scarecrow-like 8 (SCL8) 180 2e-45
At1g66350 gibberellin regulatory protein like 180 2e-45
At1g07520 transcription factor scarecrow-like 14, putative 175 6e-44
At3g54220 SCARECROW1 167 1e-41
At3g50650 scarecrow-like 7 (SCL7) 164 1e-40
At5g66770 SCARECROW gene regulator 161 1e-39
>At5g48150 SCARECROW gene regulator-like
Length = 490
Score = 467 bits (1202), Expect = e-132
Identities = 235/420 (55%), Positives = 302/420 (70%), Gaps = 5/420 (1%)
Query: 179 KESEISLLGPSS-DIVDSCQSNLNGSLHQGTSQYDWSQFEEIIPKLDLKEELIRCAQFVF 237
+E E ++GP S D++ C + + + Q + W E I + DL+ +L+ CA+ +
Sbjct: 74 REIETVMMGPDSLDLLVDCTDSFDSTASQEIN--GWRSTLEAISRRDLRADLVSCAKAMS 131
Query: 238 DGDFQKAIGFMNKVLGKMVSVAGSPIQRLGAYMLEGLRARVESSGSAIYKAL-KCEEPTS 296
+ D A M K L +MVSV+G PIQRLGAY+LEGL A++ SSGS+IYKAL +C EP S
Sbjct: 132 ENDLMMAHSMMEK-LRQMVSVSGEPIQRLGAYLLEGLVAQLASSGSSIYKALNRCPEPAS 190
Query: 297 IELMSAMHILYQICPYFQFAYISSNAVICEEMQNESRIHIIDFQIAQGSQWMLLLHALKH 356
EL+S MHILY++CPYF+F Y+S+N I E M+ E+R+HIIDFQI QGSQW+ L+ A
Sbjct: 191 TELLSYMHILYEVCPYFKFGYMSANGAIAEAMKEENRVHIIDFQIGQGSQWVTLIQAFAA 250
Query: 357 KPGGPPFIRVTGIDDSQSFHARGGKLDIVGKKLEDCAKTCKVPFEFNSVKMYGCEVQLED 416
+PGGPP IR+TGIDD S +ARGG L IVG +L AK VPFEFNSV + EV+ ++
Sbjct: 251 RPGGPPRIRITGIDDMTSAYARGGGLSIVGNRLAKLAKQFNVPFEFNSVSVSVSEVKPKN 310
Query: 417 FEVQHDEVLVVNFPFALHHIPDESVSMENHRDRLLRLVKILSPKVVLFVEQESNTNTSPF 476
V+ E L VNF F LHH+PDESVS ENHRDRLLR+VK LSPKVV VEQESNTNT+ F
Sbjct: 311 LGVRPGEALAVNFAFVLHHMPDESVSTENHRDRLLRMVKSLSPKVVTLVEQESNTNTAAF 370
Query: 477 LPRFAETLNYYTAMFESIDVALPRDDKKRINAEQHCVARDIVNIIACEGDERFERHELFG 536
PRF ET+NYY AMFESIDV LPRD K+RIN EQHC+ARD+VNIIACEG +R ERHEL G
Sbjct: 371 FPRFMETMNYYAAMFESIDVTLPRDHKQRINVEQHCLARDVVNIIACEGADRVERHELLG 430
Query: 537 KWKARFSMAGFVPLLLSPSVIDSVRTLLKDFNKDYRIEQTDVAINLAWKSKVMCTSSAWR 596
KW++RF MAGF P LSP V ++++LL++++ YR+E+ D A+ L W + + S AW+
Sbjct: 431 KWRSRFGMAGFTPYPLSPLVNSTIKSLLRNYSDKYRLEERDGALYLGWMHRDLVASCAWK 490
>At1g50600 scarecrow-like 5 (SCL5)
Length = 597
Score = 439 bits (1130), Expect = e-123
Identities = 230/447 (51%), Positives = 313/447 (69%), Gaps = 11/447 (2%)
Query: 152 NSNGSLHQGTSQD-NEFHIDEMISEQGFKESEISLLGPSSDIVDSCQSNLNGSLHQ-GTS 209
N+N L ++ + NE + M+ K+ E +++ P VD+ +N G Q G
Sbjct: 160 NNNSPLSGSSATNTNETELSLML-----KDLETAMMEPD---VDNSYNNQGGFGQQHGVV 211
Query: 210 QYDWSQFEEIIPKLDLKEELIRCAQFVFDGDFQKAIGFMNKVLGKMVSVAGSPIQRLGAY 269
+ E+I + DLK L CA+ V + D + +++ L +MVSV+G P+QRLGAY
Sbjct: 212 SSAMYRSMEMISRGDLKGVLYECAKAVENYDLEMTDWLISQ-LQQMVSVSGEPVQRLGAY 270
Query: 270 MLEGLRARVESSGSAIYKALKCEEPTSIELMSAMHILYQICPYFQFAYISSNAVICEEMQ 329
MLEGL AR+ SSGS+IYKAL+C++PT EL++ MHILY+ CPYF+F Y S+N I E ++
Sbjct: 271 MLEGLVARLASSGSSIYKALRCKDPTGPELLTYMHILYEACPYFKFGYESANGAIAEAVK 330
Query: 330 NESRIHIIDFQIAQGSQWMLLLHALKHKPGGPPFIRVTGIDDSQSFHARGGKLDIVGKKL 389
NES +HIIDFQI+QG QW+ L+ AL +PGGPP +R+TGIDD +S AR G L++VG++L
Sbjct: 331 NESFVHIIDFQISQGGQWVSLIRALGARPGGPPNVRITGIDDPRSSFARQGGLELVGQRL 390
Query: 390 EDCAKTCKVPFEFNSVKMYGCEVQLEDFEVQHDEVLVVNFPFALHHIPDESVSMENHRDR 449
A+ C VPFEF+ + EV++E V++ E L VNFP LHH+PDESV++ENHRDR
Sbjct: 391 GKLAEMCGVPFEFHGAALCCTEVEIEKLGVRNGEALAVNFPLVLHHMPDESVTVENHRDR 450
Query: 450 LLRLVKILSPKVVLFVEQESNTNTSPFLPRFAETLNYYTAMFESIDVALPRDDKKRINAE 509
LLRLVK LSP VV VEQE+NTNT+PFLPRF ET+N+Y A+FESIDV L RD K+RIN E
Sbjct: 451 LLRLVKHLSPNVVTLVEQEANTNTAPFLPRFVETMNHYLAVFESIDVKLARDHKERINVE 510
Query: 510 QHCVARDIVNIIACEGDERFERHELFGKWKARFSMAGFVPLLLSPSVIDSVRTLLKDFNK 569
QHC+AR++VN+IACEG ER ERHE GKW++RF MAGF P LS V +++ LL+ +++
Sbjct: 511 QHCLAREVVNLIACEGVEREERHEPLGKWRSRFHMAGFKPYPLSSYVNATIKGLLESYSE 570
Query: 570 DYRIEQTDVAINLAWKSKVMCTSSAWR 596
Y +E+ D A+ L WK++ + TS AWR
Sbjct: 571 KYTLEERDGALYLGWKNQPLITSCAWR 597
>At2g04890 putative SCARECROW gene regulator
Length = 413
Score = 435 bits (1119), Expect = e-122
Identities = 218/379 (57%), Positives = 278/379 (72%), Gaps = 8/379 (2%)
Query: 218 EIIPKLDLKEELIRCAQFVFDGDFQKAIGFMNKVLGKMVSVAGSPIQRLGAYMLEGLRAR 277
E I + DLK L+ CA+ V + + A M ++ G MVS++G PIQRLGAYMLEGL AR
Sbjct: 43 EAISRGDLKLVLVACAKAVSENNLLMARWCMGELRG-MVSISGEPIQRLGAYMLEGLVAR 101
Query: 278 VESSGSAIYKALKCEEPTSIELMSAMHILYQICPYFQFAYISSNAVICEEMQNESRIHII 337
+ +SGS+IYK+L+ EP S E +S +++L+++CPYF+F Y+S+N I E M++E RIHII
Sbjct: 102 LAASGSSIYKSLQSREPESYEFLSYVYVLHEVCPYFKFGYMSANGAIAEAMKDEERIHII 161
Query: 338 DFQIAQGSQWMLLLHALKHKPGGPPFIRVTGIDDSQSFHARGGKLDIVGKKLEDCAKTCK 397
DFQI QGSQW+ L+ A +PGG P IR+TG+ D G L V K+LE AK
Sbjct: 162 DFQIGQGSQWIALIQAFAARPGGAPNIRITGVGD-------GSVLVTVKKRLEKLAKKFD 214
Query: 398 VPFEFNSVKMYGCEVQLEDFEVQHDEVLVVNFPFALHHIPDESVSMENHRDRLLRLVKIL 457
VPF FN+V CEV++E+ +V+ E L VNF + LHH+PDESVSMENHRDRLLR+VK L
Sbjct: 215 VPFRFNAVSRPSCEVEVENLDVRDGEALGVNFAYMLHHLPDESVSMENHRDRLLRMVKSL 274
Query: 458 SPKVVLFVEQESNTNTSPFLPRFAETLNYYTAMFESIDVALPRDDKKRINAEQHCVARDI 517
SPKVV VEQE NTNTSPFLPRF ETL+YYTAMFESIDV LPR+ K+RIN EQHC+ARD+
Sbjct: 275 SPKVVTLVEQECNTNTSPFLPRFLETLSYYTAMFESIDVMLPRNHKERINIEQHCMARDV 334
Query: 518 VNIIACEGDERFERHELFGKWKARFSMAGFVPLLLSPSVIDSVRTLLKDFNKDYRIEQTD 577
VNIIACEG ER ERHEL GKWK+RFSMAGF P LS + ++R LL+D++ Y IE+ D
Sbjct: 335 VNIIACEGAERIERHELLGKWKSRFSMAGFEPYPLSSIISATIRALLRDYSNGYAIEERD 394
Query: 578 VAINLAWKSKVMCTSSAWR 596
A+ L W +++ +S AW+
Sbjct: 395 GALYLGWMDRILVSSCAWK 413
>At4g17230 scarecrow-like 13 (SCL13)
Length = 287
Score = 391 bits (1004), Expect = e-109
Identities = 190/282 (67%), Positives = 227/282 (80%)
Query: 255 MVSVAGSPIQRLGAYMLEGLRARVESSGSAIYKALKCEEPTSIELMSAMHILYQICPYFQ 314
MVSV+GSPIQRLG YM EGLRAR+E SGS IYK+LKC EPT ELMS M +LY+ICPY++
Sbjct: 1 MVSVSGSPIQRLGTYMAEGLRARLEGSGSNIYKSLKCNEPTGRELMSYMSVLYEICPYWK 60
Query: 315 FAYISSNAVICEEMQNESRIHIIDFQIAQGSQWMLLLHALKHKPGGPPFIRVTGIDDSQS 374
FAY ++N I E + E+R+HIIDFQIAQGSQ+M L+ L PGGPP +RVTG+DDSQS
Sbjct: 61 FAYTTANVEILEAIAGETRVHIIDFQIAQGSQYMFLIQELAKHPGGPPLLRVTGVDDSQS 120
Query: 375 FHARGGKLDIVGKKLEDCAKTCKVPFEFNSVKMYGCEVQLEDFEVQHDEVLVVNFPFALH 434
+ARGG L +VG++L A++C VPFEF+ M GC+VQ E ++ +VVNFP+ LH
Sbjct: 121 TYARGGGLSLVGERLATLAQSCGVPFEFHDAIMSGCKVQREHLGLEPGFAVVVNFPYVLH 180
Query: 435 HIPDESVSMENHRDRLLRLVKILSPKVVLFVEQESNTNTSPFLPRFAETLNYYTAMFESI 494
H+PDESVS+ENHRDRLL L+K LSPK+V VEQESNTNTSPFL RF ETL+YYTAMFESI
Sbjct: 181 HMPDESVSVENHRDRLLHLIKSLSPKLVTLVEQESNTNTSPFLSRFVETLDYYTAMFESI 240
Query: 495 DVALPRDDKKRINAEQHCVARDIVNIIACEGDERFERHELFG 536
D A PRDDK+RI+AEQHCVARDIVN+IACE ER ERHE+ G
Sbjct: 241 DAARPRDDKQRISAEQHCVARDIVNMIACEESERVERHEVLG 282
>At1g21450 scarecrow-like 1 (SCL1)
Length = 593
Score = 355 bits (911), Expect = 4e-98
Identities = 212/591 (35%), Positives = 323/591 (53%), Gaps = 59/591 (9%)
Query: 34 TLQTNSNNGG---------RRQRTNVSLETSNWESNWEKQFCTLESSQVANDVIAFDSPS 84
TL N NN G R + + S ++EK F + + I +
Sbjct: 34 TLNENGNNNGVSSAQIFDPDRSKNPCLTDDSYPSQSYEKYFLDSPTDEFVQHPIGSGASV 93
Query: 85 SASGD----PFNMRSTF--SPQISRYCSSDNTTNVSPLGVYSSA------------DDDS 126
S+ G P+ R S + S +T++ LG Y + DD+
Sbjct: 94 SSFGSLDSFPYQSRPVLGCSMEFQLPLDSTSTSSTRLLGDYQAVSYSPSMDVVEEFDDEQ 153
Query: 127 FEFKMKEIELSLVGTDIEVFGSCLSNSNGSLHQGTSQDNEFHIDEMISEQGFKESEISLL 186
K++E+E +L+G + + DN ID S Q ESE
Sbjct: 154 MRSKIQELERALLGDEDDKM--------------VGIDNLMEIDSEWSYQN--ESEQHQD 197
Query: 187 GPSSDIVDSCQSNLNGSLHQGTSQYDWSQFEEIIPKLDLKEELIRCAQFVFDGDFQKAIG 246
P S SN + S +E++ + K+ LI CA+ + +G ++A+
Sbjct: 198 SPKES--SSADSNSHVSS------------KEVVSQATPKQILISCARALSEGKLEEALS 243
Query: 247 FMNKVLGKMVSVAGSPIQRLGAYMLEGLRARVESSGSAIYKALKCEEPTSIELMSAMHIL 306
+N+ L ++VS+ G P QR+ AYM+EGL AR+ +SG IY+ALKC+EP S E ++AM +L
Sbjct: 244 MVNE-LRQIVSIQGDPSQRIAAYMVEGLAARMAASGKFIYRALKCKEPPSDERLAAMQVL 302
Query: 307 YQICPYFQFAYISSNAVICEEMQNESRIHIIDFQIAQGSQWMLLLHALKHKPGGPPFIRV 366
+++CP F+F ++++N I E ++ E +HIIDF I QG+Q+M L+ ++ PG P +R+
Sbjct: 303 FEVCPCFKFGFLAANGAILEAIKGEEEVHIIDFDINQGNQYMTLIRSIAELPGKRPRLRL 362
Query: 367 TGIDDSQSFHARGGKLDIVGKKLEDCAKTCKVPFEFNSVKMYGCEVQLEDFEVQHDEVLV 426
TGIDD +S G L I+G +LE A+ V F+F ++ V + E L+
Sbjct: 363 TGIDDPESVQRSIGGLRIIGLRLEQLAEDNGVSFKFKAMPSKTSIVSPSTLGCKPGETLI 422
Query: 427 VNFPFALHHIPDESVSMENHRDRLLRLVKILSPKVVLFVEQESNTNTSPFLPRFAETLNY 486
VNF F LHH+PDESV+ N RD LL +VK L+PK+V VEQ+ NTNTSPF PRF E Y
Sbjct: 423 VNFAFQLHHMPDESVTTVNQRDELLHMVKSLNPKLVTVVEQDVNTNTSPFFPRFIEAYEY 482
Query: 487 YTAMFESIDVALPRDDKKRINAEQHCVARDIVNIIACEGDERFERHELFGKWKARFSMAG 546
Y+A+FES+D+ LPR+ ++R+N E+ C+ARDIVNI+ACEG+ER ER+E GKW+AR MAG
Sbjct: 483 YSAVFESLDMTLPRESQERMNVERQCLARDIVNIVACEGEERIERYEAAGKWRARMMMAG 542
Query: 547 FVPLLLSPSVIDSVRTLLK-DFNKDYRIEQTDVAINLAWKSKVMCTSSAWR 596
F P +S V ++++ L+K + Y++++ ++ W+ K + +SAWR
Sbjct: 543 FNPKPMSAKVTNNIQNLIKQQYCNKYKLKEEMGELHFCWEEKSLIVASAWR 593
>At1g07530 transcription factor scarecrow-like 14, putative
Length = 769
Score = 211 bits (538), Expect = 7e-55
Identities = 128/408 (31%), Positives = 220/408 (53%), Gaps = 10/408 (2%)
Query: 193 VDSCQSN---LNGSLHQGTSQYDWSQFEEIIPKLDLKEELIRCAQFVFDGDFQKAIGFMN 249
V + QSN + G TS + S+ E DL+ L+ CAQ V D ++ M
Sbjct: 362 VVTAQSNGAKIRGKKSTSTSHSNDSKKETA----DLRTLLVLCAQAVSVDD-RRTANEML 416
Query: 250 KVLGKMVSVAGSPIQRLGAYMLEGLRARVESSGSAIYKALKCEEPTSIELMSAMHILYQI 309
+ + + S G+ +RL Y L AR+ +G+ IY AL ++ ++ +++ A +
Sbjct: 417 RQIREHSSPLGNGSERLAHYFANSLEARLAGTGTQIYTALSSKKTSAADMLKAYQTYMSV 476
Query: 310 CPYFQFAYISSNAVICEEMQNESRIHIIDFQIAQGSQWMLLLHALK-HKPGGPPFIRVTG 368
CP+ + A I +N + N + IHIIDF I+ G QW L+H L +PGG P +R+TG
Sbjct: 477 CPFKKAAIIFANHSMMRFTANANTIHIIDFGISYGFQWPALIHRLSLSRPGGSPKLRITG 536
Query: 369 IDDSQSFHARGGKLDIVGKKLEDCAKTCKVPFEFNSVKMYGCEVQLEDFEVQHDEVLVVN 428
I+ Q + G +L + VPFE+N++ +Q+ED +++ E +VVN
Sbjct: 537 IELPQRGFRPAEGVQETGHRLARYCQRHNVPFEYNAIAQKWETIQVEDLKLRQGEYVVVN 596
Query: 429 FPFALHHIPDESVSMENHRDRLLRLVKILSPKVVLFVEQESNTNTSPFLPRFAETLNYYT 488
F ++ DE+V + + RD +L+L++ ++P V + N N F+ RF E L +Y+
Sbjct: 597 SLFRFRNLLDETVLVNSPRDAVLKLIRKINPNVFIPAILSGNYNAPFFVTRFREALFHYS 656
Query: 489 AMFESIDVALPRDDKKRINAEQHCVARDIVNIIACEGDERFERHELFGKWKARFSMAGFV 548
A+F+ D L R+D+ R+ E+ R+IVN++ACEG ER ER E + +W+AR AGF
Sbjct: 657 AVFDMCDSKLAREDEMRLMYEKEFYGREIVNVVACEGTERVERPETYKQWQARLIRAGFR 716
Query: 549 PLLLSPSVIDSVRTLLKD-FNKDYRIEQTDVAINLAWKSKVMCTSSAW 595
L L ++ +++ +++ ++K++ ++Q + WK +++ SS W
Sbjct: 717 QLPLEKELMQNLKLKIENGYDKNFDVDQNGNWLLQGWKGRIVYASSLW 764
>At5g59450 putative SCARECROW transcriptional regulatory protein
(F2O15.13)
Length = 610
Score = 206 bits (523), Expect = 4e-53
Identities = 114/378 (30%), Positives = 200/378 (52%), Gaps = 5/378 (1%)
Query: 223 LDLKEELIRCAQFVFDGDFQKAIGFMNKVLGKMVSVAGSPIQRLGAYMLEGLRARVESSG 282
+DL+ L +CAQ V D ++A + ++ S G QRL Y E L AR+ +
Sbjct: 222 VDLRSLLTQCAQAVASFDQRRATDKLKEIRAHSSS-NGDGTQRLAFYFAEALEARITGNI 280
Query: 283 SA-IYKALKCEEPTSIELMSAMHILYQICPYFQFAYISSNAVICEEMQNESRIHIIDFQI 341
S + + ++++ A + CP + Y ++N I E +++HI+DF +
Sbjct: 281 SPPVSNPFPSSTTSMVDILKAYKLFVHTCPIYVTDYFAANKSIYELAMKATKLHIVDFGV 340
Query: 342 AQGSQWMLLLHALKHKPGGPPFIRVTGIDDSQSFHARGGKLDIVGKKLEDCAKTCKVPFE 401
G QW LL AL +PGGPP +RVTGI+ Q+ +++ G++L+ VPFE
Sbjct: 341 LYGFQWPCLLRALSKRPGGPPMLRVTGIELPQAGFRPSDRVEETGRRLKRFCDQFNVPFE 400
Query: 402 FNSVKMYGCEVQLEDFEVQHDEVLVVNFPFALHHIPDESVSMENHRDRLLRLVKILSPKV 461
FN + + L++ + E VVN L + PDE+VS+++ RD +L+L + ++P +
Sbjct: 401 FNFIAKKWETITLDELMINPGETTVVNCIHRLQYTPDETVSLDSPRDTVLKLFRDINPDL 460
Query: 462 VLFVEQESNTNTSPFLPRFAETLNYYTAMFESIDVALPRDD--KKRINAEQHCVARDIVN 519
+F E N+ F+ RF E L +Y+++F+ D + +D K R E+ + RD ++
Sbjct: 461 FVFAEINGMYNSPFFMTRFREALFHYSSLFDMFDTTIHAEDEYKNRSLLERELLVRDAMS 520
Query: 520 IIACEGDERFERHELFGKWKARFSMAGFVPLLLSPSVIDSVRTLL-KDFNKDYRIEQTDV 578
+I+CEG ERF R E + +W+ R AGF P +S ++ + ++ K +++D+ I+ +
Sbjct: 521 VISCEGAERFARPETYKQWRVRILRAGFKPATISKQIMKEAKEIVRKRYHRDFVIDSDNN 580
Query: 579 AINLAWKSKVMCTSSAWR 596
+ WK +V+ S W+
Sbjct: 581 WMLQGWKGRVIYAFSCWK 598
>At2g37650 scarecrow-like 9 (SCL9)
Length = 718
Score = 205 bits (522), Expect = 5e-53
Identities = 116/375 (30%), Positives = 202/375 (52%), Gaps = 4/375 (1%)
Query: 223 LDLKEELIRCAQFVFDGDFQKAIGFMNKVLGKMVSVAGSPIQRLGAYMLEGLRARVESSG 282
+DL+ LI CAQ V D ++ G + K + + G QRL GL AR+ +G
Sbjct: 342 VDLRSLLIHCAQAVAADD-RRCAGQLLKQIRLHSTPFGDGNQRLAHCFANGLEARLAGTG 400
Query: 283 SAIYKALKCEEPTSIELMSAMHILYQICPYFQFAYISSNAVICEEMQNESRIHIIDFQIA 342
S IYK + + ++ ++ A + CP+ + +Y +N I + + N R+H+IDF I
Sbjct: 401 SQIYKGIVSKPRSAAAVLKAHQLFLACCPFRKLSYFITNKTIRDLVGNSQRVHVIDFGIL 460
Query: 343 QGSQWMLLLHALKHKPGGPPFIRVTGIDDSQSFHARGGKLDIVGKKLEDCAKTCKVPFEF 402
G QW L+H + G P +R+TGI+ Q +++ G++L AK VPFE+
Sbjct: 461 YGFQWPTLIH--RFSMYGSPKVRITGIEFPQPGFRPAQRVEETGQRLAAYAKLFGVPFEY 518
Query: 403 NSVKMYGCEVQLEDFEVQHDEVLVVNFPFALHHIPDESVSMENHRDRLLRLVKILSPKVV 462
++ +QLED ++ DE+ VVN + ++ DESV +E+ RD +L L+ ++P +
Sbjct: 519 KAIAKKWDAIQLEDLDIDRDEITVVNCLYRAENLHDESVKVESCRDTVLNLIGKINPDLF 578
Query: 463 LFVEQESNTNTSPFLPRFAETLNYYTAMFESIDVALPRDDKKRINAEQHCVARDIVNIIA 522
+F N F+ RF E L +++++F+ ++ +PR+D++R+ E R+ +N+IA
Sbjct: 579 VFGIVNGAYNAPFFVTRFREALFHFSSIFDMLETIVPREDEERMFLEMEVFGREALNVIA 638
Query: 523 CEGDERFERHELFGKWKARFSMAGFVPLLLSPSVI-DSVRTLLKDFNKDYRIEQTDVAIN 581
CEG ER ER E + +W R +G V + PS++ S+ + ++KD+ I+Q + +
Sbjct: 639 CEGWERVERPETYKQWHVRAMRSGLVQVPFDPSIMKTSLHKVHTFYHKDFVIDQDNRWLL 698
Query: 582 LAWKSKVMCTSSAWR 596
WK + + S W+
Sbjct: 699 QGWKGRTVMALSVWK 713
>At3g46600 scarecrow-like protein
Length = 583
Score = 203 bits (517), Expect = 2e-52
Identities = 117/375 (31%), Positives = 196/375 (52%), Gaps = 3/375 (0%)
Query: 223 LDLKEELIRCAQFVFDGDFQKAIGFMNKVLGKMVSVAGSPIQRLGAYMLEGLRARVESSG 282
+D++ L++CAQ V D ++A + K + + S G QRLG + E L AR+ +
Sbjct: 207 VDMRNLLMQCAQAVASFDQRRAFEKL-KEIREHSSRHGDATQRLGYHFAEALEARITGTM 265
Query: 283 SAIYKALKCEEPTSIELMSAMHILYQICPYFQFAYISSNAVICEEMQNESRIHIIDFQIA 342
+ A + ++++ A Q CP Y ++N I E + +HIIDF I
Sbjct: 266 TTPISATS-SRTSMVDILKAYKGFVQACPTLIMCYFTANRTINELASKATTLHIIDFGIL 324
Query: 343 QGSQWMLLLHALKHKPGGPPFIRVTGIDDSQSFHARGGKLDIVGKKLEDCAKTCKVPFEF 402
G QW L+ AL + GPP +RVTGI+ QS +++ G++L+ VPFE+
Sbjct: 325 YGFQWPCLIQALSKRDIGPPLLRVTGIELPQSGFRPSERVEETGRRLKRFCDKFNVPFEY 384
Query: 403 NSVKMYGCEVQLEDFEVQHDEVLVVNFPFALHHIPDESVSMENHRDRLLRLVKILSPKVV 462
+ + + L+D + E VVN L + PDE+VS+ + RD L+L + ++P +
Sbjct: 385 SFIAKNWENITLDDLVINSGETTVVNCILRLQYTPDETVSLNSPRDTALKLFRDINPDLF 444
Query: 463 LFVEQESNTNTSPFLPRFAETLNYYTAMFESIDVALPRDDKKRINAEQHCVARDIVNIIA 522
+F E N+ FL RF E L + +++F+ + L DD R E+ + RD +++IA
Sbjct: 445 VFAEINGTYNSPFFLTRFREALFHCSSLFDMYETTLSEDDNCRTLVERELIIRDAMSVIA 504
Query: 523 CEGDERFERHELFGKWKARFSMAGFVPLLLSPSVIDSVRTLLKD-FNKDYRIEQTDVAIN 581
CEG ERF R E + +W+ R AGF P LS ++ + ++K+ ++KD+ I+ + +
Sbjct: 505 CEGSERFARPETYKQWQVRILRAGFRPAKLSKQIVKDGKEIVKERYHKDFVIDNDNHWMF 564
Query: 582 LAWKSKVMCTSSAWR 596
WK +V+ S W+
Sbjct: 565 QGWKGRVLYAVSCWK 579
>At3g03450 RGA1-like protein
Length = 547
Score = 200 bits (508), Expect = 2e-51
Identities = 122/376 (32%), Positives = 195/376 (51%), Gaps = 14/376 (3%)
Query: 225 LKEELIRCAQFVFDGDFQKAIGFMNKVLGKMVSVAGSPIQRLGAYMLEGLRARVESSGSA 284
L L+ CA+ + + A + +V G + + ++ Y + L R+ +A
Sbjct: 180 LVHALVACAEAIHQENLNLADALVKRV-GTLAGSQAGAMGKVATYFAQALARRIYRDYTA 238
Query: 285 IYKALKCEEPTSIELMSAMHILYQICPYFQFAYISSNAVICEEMQNESRIHIIDFQIAQG 344
P+ E++ MH Y+ CPY +FA+ ++N I E + R+H+ID + QG
Sbjct: 239 ETDVCAAVNPSFEEVLE-MHF-YESCPYLKFAHFTANQAILEAVTTARRVHVIDLGLNQG 296
Query: 345 SQWMLLLHALKHKPGGPPFIRVTGIDDSQSFHARGGKLDIVGKKLEDCAKTCKVPFEFNS 404
QW L+ AL +PGGPP R+TGI Q+ L +G KL A+ V FEF
Sbjct: 297 MQWPALMQALALRPGGPPSFRLTGIGPPQT--ENSDSLQQLGWKLAQFAQNMGVEFEFKG 354
Query: 405 VKMYG-CEVQLEDFEVQ-HDEVLVVNFPFALHHIPDESVSMENHRDRLLRLVKILSPKVV 462
+ +++ E FE + E LVVN F LH + S S+E +LL VK + P +V
Sbjct: 355 LAAESLSDLEPEMFETRPESETLVVNSVFELHRLLARSGSIE----KLLNTVKAIKPSIV 410
Query: 463 LFVEQESNTNTSPFLPRFAETLNYYTAMFESIDVALPRDDKKRINAEQHCVARDIVNIIA 522
VEQE+N N FL RF E L+YY+++F+S++ + + R+ +E + + R I+N++A
Sbjct: 411 TVVEQEANHNGIVFLDRFNEALHYYSSLFDSLEDSYSLPSQDRVMSEVY-LGRQILNVVA 469
Query: 523 CEGDERFERHELFGKWKARFSMAGFVPLLLSPSVIDSVRTLLKDF--NKDYRIEQTDVAI 580
EG +R ERHE +W+ R AGF P+ L S LL + YR+E+ D +
Sbjct: 470 AEGSDRVERHETAAQWRIRMKSAGFDPIHLGSSAFKQASMLLSLYATGDGYRVEENDGCL 529
Query: 581 NLAWKSKVMCTSSAWR 596
+ W+++ + T+SAW+
Sbjct: 530 MIGWQTRPLITTSAWK 545
>At5g17490 RGA-like protein
Length = 523
Score = 192 bits (487), Expect = 6e-49
Identities = 148/510 (29%), Positives = 243/510 (47%), Gaps = 47/510 (9%)
Query: 117 GVYSSADDDSFEF------KMKEIELSLVGTDIEVFGSCLSNS--------NGSLHQGTS 162
G DD+ EF K++ +++ V +E LSN N ++H S
Sbjct: 24 GCGGGGDDNMDEFLAVLGYKVRSSDMADVAQKLEQLEMVLSNDIASSSNAFNDTVHYNPS 83
Query: 163 QDNEFHIDEMISEQGF----KESEISLLGPSSDIVDSCQSNLNGSLHQGTSQYDWSQFEE 218
D M+S+ + + I L P +D + C SN N + + S E
Sbjct: 84 -DLSGWAQSMLSDLNYYPDLDPNRICDLRPITDDDECCSSNSNSNKRIRLGPWCDSVTSE 142
Query: 219 IIPKLDLKEE--------LIRCAQFVFDGDFQKAIGFMNKVLGKMVSVAGSPIQRLGAYM 270
+ L EE L+ CA+ V + A + +V G + + + ++ Y
Sbjct: 143 STRSVVLIEETGVRLVQALVACAEAVQLENLSLADALVKRV-GLLAASQAGAMGKVATYF 201
Query: 271 LEGLRARVESSGSAIYKALKCEEPTSIELMSAMHILYQICPYFQFAYISSNAVICEEMQN 330
E L R+ I+ + +P+ E++ Y CPY +FA+ ++N I E +
Sbjct: 202 AEALARRIYR----IHPSAAAIDPSFEEILQMN--FYDSCPYLKFAHFTANQAILEAVTT 255
Query: 331 ESRIHIIDFQIAQGSQWMLLLHALKHKPGGPPFIRVTGIDDSQSFHARGGKLDIVGKKLE 390
+H+ID + QG QW L+ AL +PGGPP R+TG+ + + R G + +G KL
Sbjct: 256 SRVVHVIDLGLNQGMQWPALMQALALRPGGPPSFRLTGVGNPSN---REG-IQELGWKLA 311
Query: 391 DCAKTCKVPFEFNSVKMYG-CEVQLEDFEVQ-HDEVLVVNFPFALHHIPDESVSMENHRD 448
A+ V F+FN + +++ + FE + E LVVN F LH + + S+E
Sbjct: 312 QLAQAIGVEFKFNGLTTERLSDLEPDMFETRTESETLVVNSVFELHPVLSQPGSIE---- 367
Query: 449 RLLRLVKILSPKVVLFVEQESNTNTSPFLPRFAETLNYYTAMFESIDVALPRDDKKRINA 508
+LL VK + P +V VEQE+N N FL RF E L+YY+++F+S++ + + R+ +
Sbjct: 368 KLLATVKAVKPGLVTVVEQEANHNGDVFLDRFNEALHYYSSLFDSLEDGVVIPSQDRVMS 427
Query: 509 EQHCVARDIVNIIACEGDERFERHELFGKWKARFSMAGFVPLLLSPSVIDSVRTL--LKD 566
E + + R I+N++A EG +R ERHE +W+ R AGF P+ L L L
Sbjct: 428 EVY-LGRQILNLVATEGSDRIERHETLAQWRKRMGSAGFDPVNLGSDAFKQASLLLALSG 486
Query: 567 FNKDYRIEQTDVAINLAWKSKVMCTSSAWR 596
YR+E+ D ++ LAW++K + +SAW+
Sbjct: 487 GGDGYRVEENDGSLMLAWQTKPLIAASAWK 516
>At2g29060 putative SCARECROW gene regulator
Length = 1336
Score = 192 bits (487), Expect = 6e-49
Identities = 112/377 (29%), Positives = 200/377 (52%), Gaps = 7/377 (1%)
Query: 224 DLKEELIRCAQFVFDGDFQKAIGFMNKVLGKMVSVAGSPIQRLGAYMLEGLRARVESSGS 283
DL+ L+ CAQ V D + A ++++ S G +RL Y L AR+ G+
Sbjct: 317 DLRTMLVSCAQAVSINDRRTADELLSRIRQHSSSY-GDGTERLAHYFANSLEARLAGIGT 375
Query: 284 AIYKALKCEEPTSIELMSAMHILYQICPYFQFAYISSNAVICE--EMQNESRIHIIDFQI 341
+Y AL ++ ++ +++ A +CP+ + A I +N I N IHIIDF I
Sbjct: 376 QVYTALSSKKTSTSDMLKAYQTYISVCPFKKIAIIFANHSIMRLASSANAKTIHIIDFGI 435
Query: 342 AQGSQWMLLLHALKHKPGGPPFIRVTGIDDSQ-SFHARGGKLDIVGKKLEDCAKTCKVPF 400
+ G QW L+H L + G +R+TGI+ Q F G ++ G++L + +PF
Sbjct: 436 SDGFQWPSLIHRLAWRRGSSCKLRITGIELPQRGFRPAEGVIE-TGRRLAKYCQKFNIPF 494
Query: 401 EFNSVKMYGCEVQLEDFEVQHDEVLVVNFPFALHHIPDESVSMENHRDRLLRLVKILSPK 460
E+N++ ++LED +++ E + VN F ++ DE+V++ + RD +L+L++ + P
Sbjct: 495 EYNAIAQKWESIKLEDLKLKEGEFVAVNSLFRFRNLLDETVAVHSPRDTVLKLIRKIKPD 554
Query: 461 VVLFVEQESNTNTSPFLPRFAETLNYYTAMFESIDVALPRDDKKRINAEQHCVARDIVNI 520
V + + N F+ RF E L +Y+++F+ D L R+D R+ E+ R+I+N+
Sbjct: 555 VFIPGILSGSYNAPFFVTRFREVLFHYSSLFDMCDTNLTREDPMRVMFEKEFYGREIMNV 614
Query: 521 IACEGDERFERHELFGKWKARFSMAGFVPLLLSPSVIDSVRTLLKD--FNKDYRIEQTDV 578
+ACEG ER ER E + +W+AR AGF + L ++ ++ +++ K++ ++Q
Sbjct: 615 VACEGTERVERPESYKQWQARAMRAGFRQIPLEKELVQKLKLMVESGYKPKEFDVDQDCH 674
Query: 579 AINLAWKSKVMCTSSAW 595
+ WK +++ SS W
Sbjct: 675 WLLQGWKGRIVYGSSIW 691
Score = 186 bits (473), Expect = 2e-47
Identities = 111/382 (29%), Positives = 198/382 (51%), Gaps = 10/382 (2%)
Query: 223 LDLKEELIRCAQFVFDGDFQKAIGFMNKVLGKMVSVAGSPIQRLGAYMLEGLRARVESSG 282
+D + L CAQ + GD A+ F+ ++ + S G QRL L AR++ S
Sbjct: 953 VDFRTLLTHCAQAISTGDKTTALEFLLQIR-QQSSPLGDAGQRLAHCFANALEARLQGST 1011
Query: 283 SAI----YKALKCE-EPTSIELMSAMHILYQICPYFQFAYISSNAVICEEMQNESRIHII 337
+ Y AL + T+ + + A + P+ Y S +I + ++ +HI+
Sbjct: 1012 GPMIQTYYNALTSSLKDTAADTIRAYRVYLSSSPFVTLMYFFSIWMILDVAKDAPVLHIV 1071
Query: 338 DFQIAQGSQWMLLLHALKHKPGGPPFIRVTGIDDSQSFHARGGKLDIVGKKLEDCAKTCK 397
DF I G QW + + ++ + P +R+TGI+ Q +++ G++L + K
Sbjct: 1072 DFGILYGFQWPMFIQSISDRKDVPRKLRITGIELPQCGFRPAERIEETGRRLAEYCKRFN 1131
Query: 398 VPFEFNSVKMYGCE-VQLEDFEVQHDEVLVVNFPFALHHIPDESVSMEN-HRDRLLRLVK 455
VPFE+ ++ E +++ED +++ +EVL VN L ++ DE+ S EN RD +L+L++
Sbjct: 1132 VPFEYKAIASQNWETIRIEDLDIRPNEVLAVNAGLRLKNLQDETGSEENCPRDAVLKLIR 1191
Query: 456 ILSPKVVLFVEQESNTNTSPFLPRFAETLNYYTAMFESIDVALPRDDKKRINAEQHCVAR 515
++P V + + N F+ RF E + +Y+A+F+ D LPRD+K+RI E+ R
Sbjct: 1192 NMNPDVFIHAIVNGSFNAPFFISRFKEAVYHYSALFDMFDSTLPRDNKERIRFEREFYGR 1251
Query: 516 DIVNIIACEGDERFERHELFGKWKARFSMAGFVPLLLSPSVIDSVRTLLKD--FNKDYRI 573
+ +N+IACE +R ER E + +W+ R AGF + P +++ R LK ++KD+ +
Sbjct: 1252 EAMNVIACEEADRVERPETYRQWQVRMVRAGFKQKTIKPELVELFRGKLKKWRYHKDFVV 1311
Query: 574 EQTDVAINLAWKSKVMCTSSAW 595
++ + WK + + SS W
Sbjct: 1312 DENSKWLLQGWKGRTLYASSCW 1333
>At1g14920 signal response protein (GAI)
Length = 533
Score = 186 bits (472), Expect = 3e-47
Identities = 117/383 (30%), Positives = 192/383 (49%), Gaps = 33/383 (8%)
Query: 225 LKEELIRCAQFVFDGDFQKAIGFMNKVLGKMVSVAGSPIQRLGAYMLEGLRARVESSGSA 284
L L+ CA+ V + A + ++ VS G+ ++++ Y E L R
Sbjct: 169 LVHALLACAEAVQKENLTVAEALVKQIGFLAVSQIGA-MRKVATYFAEALARR------- 220
Query: 285 IYKALKCEEPTSIELMSAMHI-LYQICPYFQFAYISSNAVICEEMQNESRIHIIDFQIAQ 343
IY+ + P L + + Y+ CPY +FA+ ++N I E Q + R+H+IDF ++Q
Sbjct: 221 IYRLSPSQSPIDHSLSDTLQMHFYETCPYLKFAHFTANQAILEAFQGKKRVHVIDFSMSQ 280
Query: 344 GSQWMLLLHALKHKPGGPPFIRVTGI-----DDSQSFHARGGKLDIVGKKLEDCAKTCKV 398
G QW L+ AL +PGGPP R+TGI D+ H VG KL A+ V
Sbjct: 281 GLQWPALMQALALRPGGPPVFRLTGIGPPAPDNFDYLHE-------VGCKLAHLAEAIHV 333
Query: 399 PFEFNSV---KMYGCEVQLEDFEVQHDEVLVVNFPFALHHIPDESVSMENHRDRLLRLVK 455
FE+ + + + + E + VN F LH + + D++L +V
Sbjct: 334 EFEYRGFVANTLADLDASMLELRPSEIESVAVNSVFELHKL----LGRPGAIDKVLGVVN 389
Query: 456 ILSPKVVLFVEQESNTNTSPFLPRFAETLNYYTAMFESIDVALPRDDKKRINAEQHCVAR 515
+ P++ VEQESN N+ FL RF E+L+YY+ +F+S++ DK + +E + + +
Sbjct: 390 QIKPEIFTVVEQESNHNSPIFLDRFTESLHYYSTLFDSLEGVPSGQDK--VMSEVY-LGK 446
Query: 516 DIVNIIACEGDERFERHELFGKWKARFSMAGFVPLLLSPSVIDSVRTLLKDFN--KDYRI 573
I N++AC+G +R ERHE +W+ RF AGF + + LL FN + YR+
Sbjct: 447 QICNVVACDGPDRVERHETLSQWRNRFGSAGFAAAHIGSNAFKQASMLLALFNGGEGYRV 506
Query: 574 EQTDVAINLAWKSKVMCTSSAWR 596
E++D + L W ++ + +SAW+
Sbjct: 507 EESDGCLMLGWHTRPLIATSAWK 529
>At2g01570 putative RGA1, giberellin repsonse modulation protein
Length = 587
Score = 182 bits (463), Expect = 4e-46
Identities = 118/382 (30%), Positives = 189/382 (48%), Gaps = 31/382 (8%)
Query: 225 LKEELIRCAQFVFDGDFQKAIGFMNKVLGKMVSVAGSPIQRLGAYMLEGLRARVESSGSA 284
L L+ CA+ + + A + ++ VS AG+ ++++ Y E L R+
Sbjct: 221 LVHALMACAEAIQQNNLTLAEALVKQIGCLAVSQAGA-MRKVATYFAEALARRIYRLSPP 279
Query: 285 IYKALKCEEPTSIELMSAMHILYQICPYFQFAYISSNAVICEEMQNESRIHIIDFQIAQG 344
+ C T MH Y+ CPY +FA+ ++N I E + + R+H+IDF + QG
Sbjct: 280 QNQIDHCLSDTL-----QMHF-YETCPYLKFAHFTANQAILEAFEGKKRVHVIDFSMNQG 333
Query: 345 SQWMLLLHALKHKPGGPPFIRVTGI-----DDSQSFHARGGKLDIVGKKLEDCAKTCKVP 399
QW L+ AL + GGPP R+TGI D+S H VG KL A+ V
Sbjct: 334 LQWPALMQALALREGGPPTFRLTGIGPPAPDNSDHLHE-------VGCKLAQLAEAIHVE 386
Query: 400 FEFNSV---KMYGCEVQLEDFEVQHDEVLVVNFPFALHHIPDESVSMENHRDRLLRLVKI 456
FE+ + + + + E + VN F LH + +E ++L +VK
Sbjct: 387 FEYRGFVANSLADLDASMLELRPSDTEAVAVNSVFELHKLLGRPGGIE----KVLGVVKQ 442
Query: 457 LSPKVVLFVEQESNTNTSPFLPRFAETLNYYTAMFESIDVALPRDDKKRINAEQHCVARD 516
+ P + VEQESN N FL RF E+L+YY+ +F+S++ DK + +E + + +
Sbjct: 443 IKPVIFTVVEQESNHNGPVFLDRFTESLHYYSTLFDSLEGVPNSQDK--VMSEVY-LGKQ 499
Query: 517 IVNIIACEGDERFERHELFGKWKARFSMAGFVPLLLSPSVIDSVRTLLKDFN--KDYRIE 574
I N++ACEG +R ERHE +W RF +G P L + LL FN + YR+E
Sbjct: 500 ICNLVACEGPDRVERHETLSQWGNRFGSSGLAPAHLGSNAFKQASMLLSVFNSGQGYRVE 559
Query: 575 QTDVAINLAWKSKVMCTSSAWR 596
+++ + L W ++ + T+SAW+
Sbjct: 560 ESNGCLMLGWHTRPLITTSAWK 581
>At5g52510 scarecrow-like 8 (SCL8)
Length = 525
Score = 180 bits (456), Expect = 2e-45
Identities = 111/348 (31%), Positives = 179/348 (50%), Gaps = 25/348 (7%)
Query: 264 QRLGAYMLEGLRARVESSGSAIYKALKCEEPTSIELMSAMHILYQICPYFQFAYISSNAV 323
++L +M+ LR+R+ S + +Y E + + +LY++ P F+ + ++N
Sbjct: 188 EKLVDFMVAALRSRIASPVTELYGK---------EHLISTQLLYELSPCFKLGFEAANLA 238
Query: 324 ICEEMQNESR----IHIIDFQIAQGSQWMLLLHALKHKPGGP------PFIRVTGIDDSQ 373
I + N H+IDF I +G Q++ LL L + G P +++T + ++
Sbjct: 239 ILDAADNNDGGMMIPHVIDFDIGEGGQYVNLLRTLSTRRNGKSQSQNSPVVKITAVANNV 298
Query: 374 --SFHARGG--KLDIVGKKLEDCAKTCKVPFEFNSVKMYGC-EVQLEDFEVQHDEVLVVN 428
GG +L VG L + FN V ++ E DE L VN
Sbjct: 299 YGCLVDDGGEERLKAVGDLLSQLGDRLGISVSFNVVTSLRLGDLNRESLGCDPDETLAVN 358
Query: 429 FPFALHHIPDESVSMENHRDRLLRLVKILSPKVVLFVEQESNTNTSPFLPRFAETLNYYT 488
F L+ +PDESV EN RD LLR VK L P+VV VEQE N+NT+PFL R +E+ Y
Sbjct: 359 LAFKLYRVPDESVCTENPRDELLRRVKGLKPRVVTLVEQEMNSNTAPFLGRVSESCACYG 418
Query: 489 AMFESIDVALPRDDKKRINAEQHCVARDIVNIIACEGDERFERHELFGKWKARFSMAGFV 548
A+ ES++ +P + R E+ + R +VN +ACEG +R ER E+FGKW+ R SMAGF
Sbjct: 419 ALLESVESTVPSTNSDRAKVEEG-IGRKLVNAVACEGIDRIERCEVFGKWRMRMSMAGFE 477
Query: 549 PLLLSPSVIDSVRTLLKDFNKDYRIEQTDVAINLAWKSKVMCTSSAWR 596
+ LS + +S+++ + + +++ + + W + + +SAWR
Sbjct: 478 LMPLSEKIAESMKSRGNRVHPGFTVKEDNGGVCFGWMGRALTVASAWR 525
>At1g66350 gibberellin regulatory protein like
Length = 511
Score = 180 bits (456), Expect = 2e-45
Identities = 114/377 (30%), Positives = 193/377 (50%), Gaps = 27/377 (7%)
Query: 225 LKEELIRCAQFVFDGDFQKAIGFMNKVLGKMVSVAGSPIQRLGAYMLEGLRARVESSGSA 284
L L+ CA+ V + + A + V G + S ++++ Y EGL R
Sbjct: 152 LVHALLACAEAVQQNNLKLADALVKHV-GLLASSQAGAMRKVATYFAEGLARR------- 203
Query: 285 IYKALKCEEPTSIELMSAMHI-LYQICPYFQFAYISSNAVICEEMQNESRIHIIDFQIAQ 343
IY+ ++ + I Y+ CPY +FA+ ++N I E ++H+ID +
Sbjct: 204 IYRIYPRDDVALSSFSDTLQIHFYESCPYLKFAHFTANQAILEVFATAEKVHVIDLGLNH 263
Query: 344 GSQWMLLLHALKHKPGGPPFIRVTGIDDSQSFHARGGKLDIVGKKLEDCAKTCKVPFEFN 403
G QW L+ AL +P GPP R+TGI S + + VG KL A T V FEF
Sbjct: 264 GLQWPALIQALALRPNGPPDFRLTGIGYSLT------DIQEVGWKLGQLASTIGVNFEFK 317
Query: 404 SVKMYG-CEVQLEDFEVQHD-EVLVVNFPFALHHIPDESVSMENHRDRLLRLVKILSPKV 461
S+ + +++ E +++ E + VN F LH + S+ D+ L +K + P +
Sbjct: 318 SIALNNLSDLKPEMLDIRPGLESVAVNSVFELHRLLAHPGSI----DKFLSTIKSIRPDI 373
Query: 462 VLFVEQESNTNTSPFLPRFAETLNYYTAMFESIDVALPRDDKKRINAEQHCVARDIVNII 521
+ VEQE+N N + FL RF E+L+YY+++F+S++ +D R+ +E + R I+N++
Sbjct: 374 MTVVEQEANHNGTVFLDRFTESLHYYSSLFDSLEGPPSQD---RVMSELF-LGRQILNLV 429
Query: 522 ACEGDERFERHELFGKWKARFSMAGFVPLLLSPSVIDSVRTLLKDF--NKDYRIEQTDVA 579
ACEG++R ERHE +W+ RF + GF P+ + + LL + Y +E+ +
Sbjct: 430 ACEGEDRVERHETLNQWRNRFGLGGFKPVSIGSNAYKQASMLLALYAGADGYNVEENEGC 489
Query: 580 INLAWKSKVMCTSSAWR 596
+ L W+++ + +SAWR
Sbjct: 490 LLLGWQTRPLIATSAWR 506
>At1g07520 transcription factor scarecrow-like 14, putative
Length = 695
Score = 175 bits (444), Expect = 6e-44
Identities = 108/381 (28%), Positives = 195/381 (50%), Gaps = 9/381 (2%)
Query: 223 LDLKEELIRCAQFVFDGDFQKAIGFMNKVLGKMVSVAGSPIQRLGAYMLEGLRARVE-SS 281
+D + L CAQ V GD A + ++ K S G QRL + L AR+E S+
Sbjct: 313 VDFRTLLTLCAQSVSAGDKITADDLLRQIR-KQCSPVGDASQRLAHFFANALEARLEGST 371
Query: 282 GSAI---YKALKCEEPTSIELMSAMHILYQICPYFQFAYISSNAVICEEMQNESRIHIID 338
G+ I Y ++ ++ T+ +++ + + P+ Y SN +I + ++ S +HI+D
Sbjct: 372 GTMIQSYYDSISSKKRTAAQILKSYSVFLSASPFMTLIYFFSNKMILDAAKDASVLHIVD 431
Query: 339 FQIAQGSQWMLLLHALKHKPGGPPFIRVTGIDDSQSFHARGGKLDIVGKKLEDCAKTCKV 398
F I G QW + + L G +R+TGI+ Q ++ G++L + K V
Sbjct: 432 FGILYGFQWPMFIQHLSKSNPGLRKLRITGIEIPQHGLRPTERIQDTGRRLTEYCKRFGV 491
Query: 399 PFEFNSVKMYGCE-VQLEDFEVQHDEVLVVNFPFALHHIPDESVSMEN-HRDRLLRLVKI 456
PFE+N++ E +++E+F+++ +EVL VN ++ D E+ RD L+L++
Sbjct: 492 PFEYNAIASKNWETIKMEEFKIRPNEVLAVNAVLRFKNLRDVIPGEEDCPRDGFLKLIRD 551
Query: 457 LSPKVVLFVEQESNTNTSPFLPRFAETLNYYTAMFESIDVALPRDDKKRINAEQHCVARD 516
++P V L + N F RF E L +Y+A+F+ L +++ +RI+ E R+
Sbjct: 552 MNPNVFLSSTVNGSFNAPFFTTRFKEALFHYSALFDLFGATLSKENPERIHFEGEFYGRE 611
Query: 517 IVNIIACEGDERFERHELFGKWKARFSMAGFVPLLLSPSVIDSVRTLLK--DFNKDYRIE 574
++N+IACEG +R ER E + +W+ R AGF + ++ R +K ++KD+ ++
Sbjct: 612 VMNVIACEGVDRVERPETYKQWQVRMIRAGFKQKPVEAELVQLFREKMKKWGYHKDFVLD 671
Query: 575 QTDVAINLAWKSKVMCTSSAW 595
+ WK +++ +SS W
Sbjct: 672 EDSNWFLQGWKGRILFSSSCW 692
>At3g54220 SCARECROW1
Length = 653
Score = 167 bits (424), Expect = 1e-41
Identities = 116/380 (30%), Positives = 191/380 (49%), Gaps = 25/380 (6%)
Query: 223 LDLKEELIRCAQFVFDGDFQKAIGFMNKVLGKMVSVA---GSPIQRLGAYMLEGLRARVE 279
L L L++CA+ V + ++A NK+L ++ ++ G+ QR+ AY E + AR+
Sbjct: 288 LHLLTLLLQCAEAVSADNLEEA----NKLLLEISQLSTPYGTSAQRVAAYFSEAMSARLL 343
Query: 280 SSGSAIYKALKCE---EPTSIELMSAMHILYQICPYFQFAYISSNAVICEEMQNESRIHI 336
+S IY AL + S++++SA + I P +F++ ++N I E + E +HI
Sbjct: 344 NSCLGIYAALPSRWMPQTHSLKMVSAFQVFNGISPLVKFSHFTANQAIQEAFEKEDSVHI 403
Query: 337 IDFQIAQGSQWMLLLHALKHKPGGPPFIRVTGIDDSQSFHARGGKLDIVGKKLEDCAKTC 396
ID I QG QW L H L +PGGPP +R+TG+ S L GK+L D A
Sbjct: 404 IDLDIMQGLQWPGLFHILASRPGGPPHVRLTGLGTSME------ALQATGKRLSDFADKL 457
Query: 397 KVPFEFNSVKMYGCEVQLEDFEVQHDEVLVVNFPFALHHIPDESVSMENHRDRLLRLVKI 456
+PFEF + + E V+ E + V++ L H + + H L L++
Sbjct: 458 GLPFEFCPLAEKVGNLDTERLNVRKREAVAVHW---LQHSLYDVTGSDAH---TLWLLQR 511
Query: 457 LSPKVVLFVEQESNTNTSPFLPRFAETLNYYTAMFESIDVALPRDDKKRINAEQHCVARD 516
L+PKVV VEQ+ ++ FL RF E ++YY+A+F+S+ + + ++R EQ ++++
Sbjct: 512 LAPKVVTVVEQDL-SHAGSFLGRFVEAIHYYSALFDSLGASYGEESEERHVVEQQLLSKE 570
Query: 517 IVNIIACEGDERFERHELFGKWKARFSMAGFVPLLLSPSVIDSVRTLLKDFNKD-YRIEQ 575
I N++A G R F W+ + GF + L+ + LL F D Y +
Sbjct: 571 IRNVLAVGGPSR-SGEVKFESWREKMQQCGFKGISLAGNAATQATLLLGMFPSDGYTLVD 629
Query: 576 TDVAINLAWKSKVMCTSSAW 595
+ + L WK + T+SAW
Sbjct: 630 DNGTLKLGWKDLSLLTASAW 649
>At3g50650 scarecrow-like 7 (SCL7)
Length = 542
Score = 164 bits (415), Expect = 1e-40
Identities = 114/351 (32%), Positives = 175/351 (49%), Gaps = 20/351 (5%)
Query: 256 VSVAGSPIQRLGAYMLEGLRAR-VESSGSAIYKALKCEEPTSIELMSAMHILYQICPYFQ 314
VS +G PIQR+G Y E L + ES S+ +L+ + + + L CPY +
Sbjct: 202 VSESGDPIQRVGYYFAEALSHKETESPSSSSSSSLE-------DFILSYKTLNDACPYSK 254
Query: 315 FAYISSNAVICEEMQNESRIHIIDFQIAQGSQWMLLLHALKHKPGGPPF-IRVTGIDDSQ 373
FA++++N I E + IHI+DF I QG QW LL AL + G P IR++GI
Sbjct: 255 FAHLTANQAILEATNQSNNIHIVDFGIFQGIQWSALLQALATRSSGKPTRIRISGIPAPS 314
Query: 374 SFHARGGKLDIVGKKLEDCAKTCKVPFEFNSVKMYGCEVQLEDFEVQHDEVLVVNFPFAL 433
+ G L G +L D A + FEF V + F V DEVLVVNF L
Sbjct: 315 LGDSPGPSLIATGNRLRDFAAILDLNFEFYPVLTPIQLLNGSSFRVDPDEVLVVNFMLEL 374
Query: 434 HHIPDESVSMENHRDRLLRLVKILSPKVVLFVEQESNTNTSPFLPRFAETLNYYTAMFES 493
+ + DE+ + LRL + L+P++V E E + N F R +L +Y+A+FES
Sbjct: 375 YKLLDETATTVG---TALRLARSLNPRIVTLGEYEVSLNRVEFANRVKNSLRFYSAVFES 431
Query: 494 IDVALPRDDKKRINAEQHCVARDIVNIIACEGDE-----RFERHELFGKWKARFSMAGFV 548
++ L RD K+R+ E+ R I++++ + D RF E +W+ AGF
Sbjct: 432 LEPNLDRDSKERLRVERVLFGRRIMDLVRSDDDNNKPGTRFGLMEEKEQWRVLMEKAGFE 491
Query: 549 PLLLSPSVIDSVRTLLKDFNKD--YRIEQTDVA-INLAWKSKVMCTSSAWR 596
P+ S + + LL ++N Y + +++ I+LAW + + T S+WR
Sbjct: 492 PVKPSNYAVSQAKLLLWNYNYSTLYSLVESEPGFISLAWNNVPLLTVSSWR 542
>At5g66770 SCARECROW gene regulator
Length = 584
Score = 161 bits (407), Expect = 1e-39
Identities = 124/402 (30%), Positives = 192/402 (46%), Gaps = 20/402 (4%)
Query: 205 HQGTSQYDWSQFEEIIPKLDLKEELIR----CAQFVFDGDFQKAIGFMNKVLGKMVSVAG 260
H+ ++ D + DL+ L++ CA+ + D D +A + ++ + VS G
Sbjct: 193 HESPTKEDPETNDSEDDDFDLEPPLLKAIYDCAR-ISDSDPNEASKTLLQIR-ESVSELG 250
Query: 261 SPIQRLGAYMLEGLRARVESSGSAIYKALKCEEPTSIELMSAMHILYQICPYFQFAYISS 320
P +R+ Y E L R+ + A + E +L+ + L CPY +FA++++
Sbjct: 251 DPTERVAFYFTEALSNRLSPNSPATSSSSSSTE----DLILSYKTLNDACPYSKFAHLTA 306
Query: 321 NAVICEEMQNESRIHIIDFQIAQGSQWMLLLHALKHKPGGPPF-IRVTGIDDSQSFHARG 379
N I E + ++IHI+DF I QG QW LL AL + G P IRV+GI +
Sbjct: 307 NQAILEATEKSNKIHIVDFGIVQGIQWPALLQALATRTSGKPTQIRVSGIPAPSLGESPE 366
Query: 380 GKLDIVGKKLEDCAKTCKVPFEFNSVKMYGCEVQLEDFEVQHDEVLVVNFPFALHHIPDE 439
L G +L D AK + F+F + + F V DEVL VNF L+ + DE
Sbjct: 367 PSLIATGNRLRDFAKVLDLNFDFIPILTPIHLLNGSSFRVDPDEVLAVNFMLQLYKLLDE 426
Query: 440 SVSMENHRDRLLRLVKILSPKVVLFVEQESNTNTSPFLPRFAETLNYYTAMFESIDVALP 499
+ ++ D LRL K L+P+VV E E + N F R L +Y+A+FES++ L
Sbjct: 427 TPTIV---DTALRLAKSLNPRVVTLGEYEVSLNRVGFANRVKNALQFYSAVFESLEPNLG 483
Query: 500 RDDKKRINAEQHCVARDIVNIIACE--GDERFERHELFGKWKARFSMAGFVPLLLSPSVI 557
RD ++R+ E+ R I +I E G R ER E +W+ AGF + LS +
Sbjct: 484 RDSEERVRVERELFGRRISGLIGPEKTGIHR-ERMEEKEQWRVLMENAGFESVKLSNYAV 542
Query: 558 DSVRTLLKDFNKDYR---IEQTDVAINLAWKSKVMCTSSAWR 596
+ LL ++N +E I+LAW + T S+WR
Sbjct: 543 SQAKILLWNYNYSNLYSIVESKPGFISLAWNDLPLLTLSSWR 584
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.134 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,943,723
Number of Sequences: 26719
Number of extensions: 617811
Number of successful extensions: 1628
Number of sequences better than 10.0: 43
Number of HSP's better than 10.0 without gapping: 34
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 1492
Number of HSP's gapped (non-prelim): 50
length of query: 598
length of database: 11,318,596
effective HSP length: 105
effective length of query: 493
effective length of database: 8,513,101
effective search space: 4196958793
effective search space used: 4196958793
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 63 (28.9 bits)
Medicago: description of AC148526.1


