

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148483.6 + phase: 0
(125 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
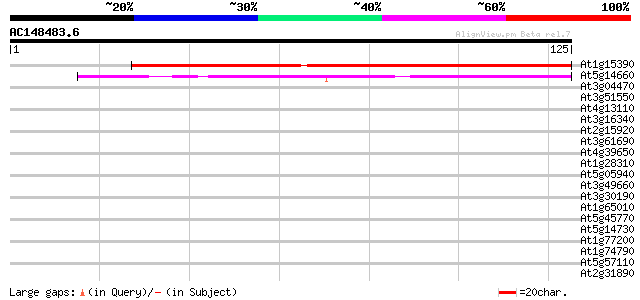
Score E
Sequences producing significant alignments: (bits) Value
At1g15390 peptide deformylase-like protein 90 3e-19
At5g14660 unknown protein 42 6e-05
At3g04470 unknown protein 30 0.31
At3g51550 receptor protein kinase like protein 29 0.53
At4g13110 putative protein 28 1.2
At3g16340 putative ABC transporter 28 1.5
At2g15920 putative retroelement pol polyprotein 28 1.5
At3g61690 unknown protein 27 2.0
At4g39650 putative gamma-glutamyltransferase 27 2.6
At1g28310 zinc finger like protein OBP3 27 2.6
At5g05940 unklnown protein 27 3.4
At3g49660 putative WD-40 repeat - protein 27 3.4
At3g30190 hypothetical protein 27 3.4
At1g65010 hypothetical protein 27 3.4
At5g45770 predicted GPI-anchored protein 26 4.5
At5g14730 unknown protein 26 4.5
At1g77200 hypothetical protein 26 4.5
At1g74790 predicted GPI-anchored protein 26 4.5
At5g57110 Ca2+-transporting ATPase-like protein (emb|CAB79748.1) 26 5.9
At2g31890 unknown protein 26 5.9
>At1g15390 peptide deformylase-like protein
Length = 269
Score = 89.7 bits (221), Expect = 3e-19
Identities = 53/98 (54%), Positives = 64/98 (65%), Gaps = 1/98 (1%)
Query: 28 SSSSCNNKIQLSSTKFSKFGSTLSSPSSETALLRKTVNKLPYIVKAGDPVIHEPAREVDH 87
SS N K+ T S ST + +K V+ LP IV +GDPV+HE AREVD
Sbjct: 42 SSHLLNRKLYNLPTSSSSSLSTKAGWLLGLGEKKKKVD-LPEIVASGDPVLHEKAREVDP 100
Query: 88 SEIKSDKIQNIIDDMILVMRKAPGVGVAAPQIGIPLRV 125
EI S++IQ IIDDMI VMR APGVG+AAPQIG+PLR+
Sbjct: 101 GEIGSERIQKIIDDMIKVMRLAPGVGLAAPQIGVPLRI 138
>At5g14660 unknown protein
Length = 273
Score = 42.4 bits (98), Expect = 6e-05
Identities = 34/128 (26%), Positives = 62/128 (47%), Gaps = 28/128 (21%)
Query: 16 MPVLSNGVVSLSSSSSCNNKIQLSSTKFSKFGSTLSSPSSETALLRKTVNKLPY------ 69
+PVLS +LS+ +L ST F ST++ S T+ +R V ++
Sbjct: 18 LPVLSRRATTLSAGYG-----RLKSTV--TFCSTVNRTSPLTSSVRAEVKRVSRKDDKVA 70
Query: 70 ------------IVKAGDPVIHEPAREVDHSEIKSDKIQNIIDDMILVMRKAPGVGVAAP 117
IV+ DP++ + +D I + ++N++D M VM K G+G++AP
Sbjct: 71 SATDVQFETPLKIVEYPDPILRAKNKRID---IFDENLKNLVDAMFDVMYKTDGIGLSAP 127
Query: 118 QIGIPLRV 125
Q+G+ +++
Sbjct: 128 QVGLNVQL 135
>At3g04470 unknown protein
Length = 640
Score = 30.0 bits (66), Expect = 0.31
Identities = 19/70 (27%), Positives = 36/70 (51%), Gaps = 3/70 (4%)
Query: 41 TKFSKFGSTLSSPSSETALLRKTVNKLPYIVKAGDPVIHEPAREVDHSEIKSDKIQNIID 100
+K++ + L+ + A LR+ V+ LP + KAG+ + E SE ++D + +ID
Sbjct: 5 SKYTHSPAHLAVVLRDHAALRRIVSDLPRLAKAGEVTTEAESME---SESRADSVSAVID 61
Query: 101 DMILVMRKAP 110
+ R+ P
Sbjct: 62 RRDVPGRETP 71
>At3g51550 receptor protein kinase like protein
Length = 895
Score = 29.3 bits (64), Expect = 0.53
Identities = 18/55 (32%), Positives = 29/55 (52%), Gaps = 6/55 (10%)
Query: 20 SNGVVSLSSSSSCNNKIQLSSTKFSKFGSTLSSPSSETALLRKTVNKLPYIVKAG 74
S+G V++ +S++ N +L+ G SPS++T L R + PYI AG
Sbjct: 211 SSGSVTIDNSTALENVYRLN------VGGNDISPSADTGLYRSWYDDQPYIFGAG 259
>At4g13110 putative protein
Length = 365
Score = 28.1 bits (61), Expect = 1.2
Identities = 24/80 (30%), Positives = 36/80 (45%), Gaps = 14/80 (17%)
Query: 17 PVLSNGVVSLSSSSSCNNKIQLSSTKFS-----------KFGSTLSSPSSETALLRKTVN 65
P+ + S SSS S N I+ STK +FGS L ET+++R+ +
Sbjct: 59 PMHESTTSSSSSSWSFGNLIKTLSTKSESVIGSYRRDLVEFGSELKK---ETSIIRRVAS 115
Query: 66 KLPYIVKAGDPVIHEPAREV 85
+LP ++ G V E V
Sbjct: 116 RLPDSLEIGASVASESLESV 135
>At3g16340 putative ABC transporter
Length = 1416
Score = 27.7 bits (60), Expect = 1.5
Identities = 18/67 (26%), Positives = 28/67 (40%)
Query: 39 SSTKFSKFGSTLSSPSSETALLRKTVNKLPYIVKAGDPVIHEPAREVDHSEIKSDKIQNI 98
S+ FS+ + E AL + KLP + +IH VD +++ D Q
Sbjct: 20 SNNHFSRRSGSTIDDHDEEALKWAALEKLPTFARLRTTIIHPHEDLVDVTKLGVDDRQKF 79
Query: 99 IDDMILV 105
ID + V
Sbjct: 80 IDSIFKV 86
>At2g15920 putative retroelement pol polyprotein
Length = 1100
Score = 27.7 bits (60), Expect = 1.5
Identities = 21/79 (26%), Positives = 39/79 (48%), Gaps = 3/79 (3%)
Query: 23 VVSLSSSSSCNNKIQLSSTKFSKFGSTLSSPSSETALLRKTVNKLPYIVKAGDPVIHEPA 82
VVS SS+ + + L++ + + L S ++ T++ N I A +PV HE
Sbjct: 977 VVSRSSAEAEYRALALATCELVSLHTLLVSLTAATSVPILFSNSTTAIYIAINPVFHERT 1036
Query: 83 REVDHSEIKSDKIQNIIDD 101
+ H+EI + ++ +DD
Sbjct: 1037 K---HTEIDCNTVREKVDD 1052
>At3g61690 unknown protein
Length = 1388
Score = 27.3 bits (59), Expect = 2.0
Identities = 18/78 (23%), Positives = 37/78 (47%), Gaps = 1/78 (1%)
Query: 29 SSSCNNKIQLSSTKFSKFGSTLSSPSSETALLRKTVNKLPYIVKAGDPVIHEPAREVDHS 88
S+S NNK ++ + S PS + + +++ Y + ++PA+EV+ +
Sbjct: 407 SNSLNNKRNQNAIRLGGVHGARSMPSQQNNCGTEITSRVTYQTQKSRGNSYQPAQEVNSN 466
Query: 89 E-IKSDKIQNIIDDMILV 105
+ +DK+Q + LV
Sbjct: 467 QSALNDKLQQTVKPETLV 484
>At4g39650 putative gamma-glutamyltransferase
Length = 574
Score = 26.9 bits (58), Expect = 2.6
Identities = 18/65 (27%), Positives = 29/65 (43%)
Query: 18 VLSNGVVSLSSSSSCNNKIQLSSTKFSKFGSTLSSPSSETALLRKTVNKLPYIVKAGDPV 77
+ +G LS S N + ++ST FG+ + SPS+ L + + GDP
Sbjct: 365 IKDHGTSHLSIIDSERNAVSMTSTINGYFGAIMLSPSTGIVLNNEMDDFSIPTKSGGDPD 424
Query: 78 IHEPA 82
+ PA
Sbjct: 425 VPPPA 429
>At1g28310 zinc finger like protein OBP3
Length = 311
Score = 26.9 bits (58), Expect = 2.6
Identities = 13/38 (34%), Positives = 19/38 (49%)
Query: 30 SSCNNKIQLSSTKFSKFGSTLSSPSSETALLRKTVNKL 67
SSCNN + + ++FS T SS E+ L + L
Sbjct: 130 SSCNNNLPMIPSRFSDSSKTCSSSGLESEFLSSGFSSL 167
>At5g05940 unklnown protein
Length = 611
Score = 26.6 bits (57), Expect = 3.4
Identities = 13/33 (39%), Positives = 19/33 (57%)
Query: 41 TKFSKFGSTLSSPSSETALLRKTVNKLPYIVKA 73
TK S + LSSP+ + + K V LPY++ A
Sbjct: 470 TKQSDDNNLLSSPAVSSIIAHKKVVPLPYLISA 502
>At3g49660 putative WD-40 repeat - protein
Length = 317
Score = 26.6 bits (57), Expect = 3.4
Identities = 19/69 (27%), Positives = 34/69 (48%), Gaps = 7/69 (10%)
Query: 16 MPVLSNGVVSLSSSSSCNNKIQLSSTKFSKFGSTLSSPSSETALLRKTVNKLPYIVKAGD 75
+P ++ + S + ++ +SS KFS G L+S S++ + T+N + D
Sbjct: 5 IPATASFTPYVHSQTLTSHNRAVSSVKFSSDGRLLASASADKTIRTYTINTI------ND 58
Query: 76 PVIHEPARE 84
P I EP +E
Sbjct: 59 P-IAEPVQE 66
>At3g30190 hypothetical protein
Length = 263
Score = 26.6 bits (57), Expect = 3.4
Identities = 18/47 (38%), Positives = 24/47 (50%)
Query: 12 RAFPMPVLSNGVVSLSSSSSCNNKIQLSSTKFSKFGSTLSSPSSETA 58
R M NG S SSSSS ++ SS+ S S+ SS SS ++
Sbjct: 16 RRHKMTKQCNGSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSTSSSSS 62
>At1g65010 hypothetical protein
Length = 1318
Score = 26.6 bits (57), Expect = 3.4
Identities = 13/75 (17%), Positives = 37/75 (49%)
Query: 28 SSSSCNNKIQLSSTKFSKFGSTLSSPSSETALLRKTVNKLPYIVKAGDPVIHEPAREVDH 87
S SS + ++ + ++ L S+ A ++ + L ++A + E R+V
Sbjct: 302 SKSSASESMESVMKQLAELNHVLHETKSDNAAQKEKIELLEKTIEAQRTDLEEYGRQVCI 361
Query: 88 SEIKSDKIQNIIDDM 102
++ ++ K++N+++ +
Sbjct: 362 AKEEASKLENLVESI 376
>At5g45770 predicted GPI-anchored protein
Length = 425
Score = 26.2 bits (56), Expect = 4.5
Identities = 18/56 (32%), Positives = 28/56 (49%), Gaps = 3/56 (5%)
Query: 17 PVLSNGVVSLSSSSSCNNKI---QLSSTKFSKFGSTLSSPSSETALLRKTVNKLPY 69
P +N S SSS +C++ ++S F+ STLS PS L K++ L +
Sbjct: 48 PCNNNNNQSSSSSITCDDASPYRHITSISFTNCSSTLSLPSKTLKPLSKSLISLSF 103
>At5g14730 unknown protein
Length = 246
Score = 26.2 bits (56), Expect = 4.5
Identities = 17/46 (36%), Positives = 28/46 (59%), Gaps = 1/46 (2%)
Query: 17 PVLSNGVVSLSSSSSCNNKIQLSSTKFSKFGSTLSSPSSETALLRK 62
P L++ SLSSS S +++ +S +KF F S + SP+ + +L K
Sbjct: 103 PSLTSPSESLSSSDSDDSE-NISPSKFYCFWSPIRSPARDDSLKSK 147
>At1g77200 hypothetical protein
Length = 244
Score = 26.2 bits (56), Expect = 4.5
Identities = 25/77 (32%), Positives = 37/77 (47%), Gaps = 9/77 (11%)
Query: 17 PVLSNGVVSLSSSSSCNNKIQLSSTKFSKFGSTLSSPSSETALLRKTVNKLPYIVKAGDP 76
P S+ VVS ++ + QLSS+ +S S+ SPSSE A T +L IV+
Sbjct: 130 PSSSSLVVSDPTTVIAPAETQLSSSSYSTCTSSSLSPSSEEA--ASTAEELSEIVEL--- 184
Query: 77 VIHEPAREVDHSEIKSD 93
P+ E + E S+
Sbjct: 185 ----PSLETSYDESLSE 197
>At1g74790 predicted GPI-anchored protein
Length = 709
Score = 26.2 bits (56), Expect = 4.5
Identities = 15/46 (32%), Positives = 22/46 (47%)
Query: 20 SNGVVSLSSSSSCNNKIQLSSTKFSKFGSTLSSPSSETALLRKTVN 65
SNGV + S CN ++ + T SSPSS ++ K +N
Sbjct: 634 SNGVYRVVRPSRCNLTCSKENSTARRNPGTSSSPSSSSSSCYKHIN 679
>At5g57110 Ca2+-transporting ATPase-like protein (emb|CAB79748.1)
Length = 1074
Score = 25.8 bits (55), Expect = 5.9
Identities = 12/32 (37%), Positives = 20/32 (62%)
Query: 94 KIQNIIDDMILVMRKAPGVGVAAPQIGIPLRV 125
K+ ++IDD++ V+ A + V A G+PL V
Sbjct: 415 KVGHVIDDVVKVLTVAVTIVVVAVPEGLPLAV 446
>At2g31890 unknown protein
Length = 671
Score = 25.8 bits (55), Expect = 5.9
Identities = 25/72 (34%), Positives = 37/72 (50%), Gaps = 8/72 (11%)
Query: 24 VSLSSSSSCNNKIQLSSTKFSKFGSTLSSPSSETALLRKT--VNKLPYIVKAGDPVIHEP 81
+SLSSSS + + LS+ K+ +F L+ SS L R+T + LP+ V A VI
Sbjct: 32 ISLSSSSFASGILPLSNKKY-RFVGPLAQRSS---LHRRTDSLKHLPFSVNAS--VIGNS 85
Query: 82 AREVDHSEIKSD 93
EV+ + D
Sbjct: 86 EEEVEEEDDDGD 97
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.316 0.132 0.366
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,637,192
Number of Sequences: 26719
Number of extensions: 96307
Number of successful extensions: 399
Number of sequences better than 10.0: 23
Number of HSP's better than 10.0 without gapping: 12
Number of HSP's successfully gapped in prelim test: 11
Number of HSP's that attempted gapping in prelim test: 386
Number of HSP's gapped (non-prelim): 24
length of query: 125
length of database: 11,318,596
effective HSP length: 87
effective length of query: 38
effective length of database: 8,994,043
effective search space: 341773634
effective search space used: 341773634
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 54 (25.4 bits)
Medicago: description of AC148483.6


