

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148483.13 + phase: 0
(63 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
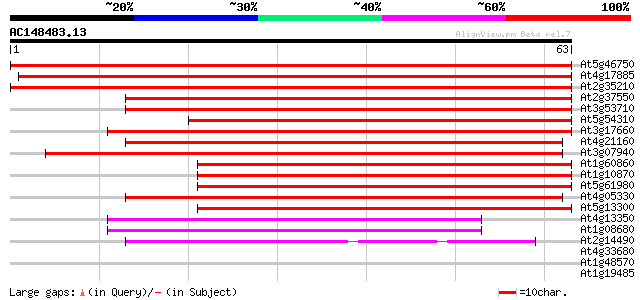
Score E
Sequences producing significant alignments: (bits) Value
At5g46750 zinc finger protein Glo3-like 119 4e-28
At4g17885 unknown protein 116 2e-27
At2g35210 unknown protein 116 2e-27
At2g37550 Asp1 (asp1) 79 3e-16
At3g53710 ARF GAP-like zinc finger-containing protein ZIGA2 79 6e-16
At5g54310 unknown protein 68 7e-13
At3g17660 unknown protein 67 2e-12
At4g21160 unknown protein 64 1e-11
At3g07940 putative GTPase activating protein 64 1e-11
At1g60860 GCN4-complementing like protein 64 2e-11
At1g10870 BRCA1-associated RING domain protein isolog 63 2e-11
At5g61980 GCN4-complementing protein - like 62 7e-11
At4g05330 unknown protein 62 7e-11
At5g13300 putative protein 61 9e-11
At4g13350 unknown protein 44 2e-05
At1g08680 unknown protein 40 3e-04
At2g14490 unknown protein 38 8e-04
At4g33680 unknown protein 25 5.5
At1g48570 unknown protein (At1g48570) 25 7.2
At1g19485 unknown protein 25 9.4
>At5g46750 zinc finger protein Glo3-like
Length = 402
Score = 119 bits (297), Expect = 4e-28
Identities = 54/63 (85%), Positives = 59/63 (92%)
Query: 1 MASDSFTDKNAVFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHIS 60
MA+++ TDKN VFRKLK+KSENK CFDC+AKNPTWASV YGIFLCIDCSAVHRSLGVHIS
Sbjct: 1 MATENLTDKNVVFRKLKSKSENKVCFDCSAKNPTWASVPYGIFLCIDCSAVHRSLGVHIS 60
Query: 61 FVR 63
FVR
Sbjct: 61 FVR 63
>At4g17885 unknown protein
Length = 413
Score = 116 bits (291), Expect = 2e-27
Identities = 52/62 (83%), Positives = 58/62 (92%)
Query: 2 ASDSFTDKNAVFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISF 61
++D+ TDKN VFRKLK+KSENK CFDC+AKNPTWASVTYGIFLCIDCSA HR+LGVHISF
Sbjct: 5 SADNLTDKNIVFRKLKSKSENKVCFDCSAKNPTWASVTYGIFLCIDCSATHRNLGVHISF 64
Query: 62 VR 63
VR
Sbjct: 65 VR 66
>At2g35210 unknown protein
Length = 395
Score = 116 bits (291), Expect = 2e-27
Identities = 53/63 (84%), Positives = 58/63 (91%)
Query: 1 MASDSFTDKNAVFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHIS 60
MAS++ DK +VF+KLK KS+NK CFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHIS
Sbjct: 1 MASENLNDKISVFKKLKAKSDNKICFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHIS 60
Query: 61 FVR 63
FVR
Sbjct: 61 FVR 63
>At2g37550 Asp1 (asp1)
Length = 456
Score = 79.3 bits (194), Expect = 3e-16
Identities = 32/50 (64%), Positives = 41/50 (82%)
Query: 14 RKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVR 63
R L+++ ENK C DC+ KNP WAS++YGIF+C++CS HR LGVHISFVR
Sbjct: 8 RTLQSQPENKVCVDCSQKNPQWASISYGIFMCLECSGKHRGLGVHISFVR 57
>At3g53710 ARF GAP-like zinc finger-containing protein ZIGA2
Length = 459
Score = 78.6 bits (192), Expect = 6e-16
Identities = 33/50 (66%), Positives = 40/50 (80%)
Query: 14 RKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVR 63
R L+++ ENK C DC KNP WASV+YGIF+C++CS HR LGVHISFVR
Sbjct: 8 RTLQSQPENKVCVDCAQKNPQWASVSYGIFMCLECSGKHRGLGVHISFVR 57
>At5g54310 unknown protein
Length = 483
Score = 68.2 bits (165), Expect = 7e-13
Identities = 28/43 (65%), Positives = 32/43 (74%)
Query: 21 ENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVR 63
EN+ C DC K P WASV GIF+C+ CS +HRSLGVHIS VR
Sbjct: 27 ENRECADCKTKGPRWASVNLGIFICMQCSGIHRSLGVHISKVR 69
>At3g17660 unknown protein
Length = 117
Score = 67.0 bits (162), Expect = 2e-12
Identities = 28/52 (53%), Positives = 35/52 (66%)
Query: 12 VFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVR 63
+ L +N+ C DC +K P WASV GIF+C+ CS +HRSLGVHIS VR
Sbjct: 18 ILEALLKHPDNRECADCRSKAPRWASVNLGIFICMQCSGIHRSLGVHISQVR 69
>At4g21160 unknown protein
Length = 337
Score = 64.3 bits (155), Expect = 1e-11
Identities = 27/49 (55%), Positives = 34/49 (69%)
Query: 14 RKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFV 62
R L T+S+N+ C DC A +P WAS G+F+C+ C VHRSLG HIS V
Sbjct: 19 RDLLTQSDNRVCADCGAPDPKWASANIGVFICLKCCGVHRSLGSHISKV 67
>At3g07940 putative GTPase activating protein
Length = 385
Score = 63.9 bits (154), Expect = 1e-11
Identities = 29/58 (50%), Positives = 36/58 (62%)
Query: 5 SFTDKNAVFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFV 62
S +D KL + NK C DC + P W S++ G+F+CI CS VHRSLGVHIS V
Sbjct: 42 SSSDPRDRLEKLLKQPGNKYCADCGSPEPKWVSLSLGVFICIKCSGVHRSLGVHISKV 99
>At1g60860 GCN4-complementing like protein
Length = 776
Score = 63.5 bits (153), Expect = 2e-11
Identities = 25/42 (59%), Positives = 33/42 (78%)
Query: 22 NKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVR 63
N +C +CNA +P WAS+ G+ +CI+CS VHR+LGVHIS VR
Sbjct: 479 NNTCAECNAPDPDWASLNLGVLMCIECSGVHRNLGVHISKVR 520
>At1g10870 BRCA1-associated RING domain protein isolog
Length = 531
Score = 63.2 bits (152), Expect = 2e-11
Identities = 26/42 (61%), Positives = 31/42 (72%)
Query: 22 NKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVR 63
N +C +CNA P WAS+ G+ LCI CS VHR+LGVHIS VR
Sbjct: 235 NNACAECNAPEPDWASLNLGVLLCIQCSGVHRNLGVHISKVR 276
>At5g61980 GCN4-complementing protein - like
Length = 850
Score = 61.6 bits (148), Expect = 7e-11
Identities = 24/42 (57%), Positives = 31/42 (73%)
Query: 22 NKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVR 63
N+ C DC A P WAS+ G+ +CI+CS +HR+LGVHIS VR
Sbjct: 532 NERCADCGAPEPDWASLNLGVLICIECSGIHRNLGVHISKVR 573
>At4g05330 unknown protein
Length = 336
Score = 61.6 bits (148), Expect = 7e-11
Identities = 25/49 (51%), Positives = 32/49 (65%)
Query: 14 RKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFV 62
R L + +N+ C DC A +P WAS G+F+C+ C VHRSLG HIS V
Sbjct: 19 RDLLNQPDNRVCADCGASDPKWASANIGVFICLKCCGVHRSLGTHISKV 67
>At5g13300 putative protein
Length = 750
Score = 61.2 bits (147), Expect = 9e-11
Identities = 25/42 (59%), Positives = 30/42 (70%)
Query: 22 NKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVR 63
N C DC A P WAS+ G+ +CI+CS VHR+LGVHIS VR
Sbjct: 436 NDKCADCGAPEPDWASLNLGVLVCIECSGVHRNLGVHISKVR 477
>At4g13350 unknown protein
Length = 602
Score = 43.9 bits (102), Expect = 2e-05
Identities = 16/42 (38%), Positives = 24/42 (57%)
Query: 12 VFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHR 53
+ R L ENK C +CN+ P + T+ F+C +CS +HR
Sbjct: 14 IIRSLLKLPENKRCINCNSLGPQYVCTTFWTFVCTNCSGIHR 55
>At1g08680 unknown protein
Length = 159
Score = 39.7 bits (91), Expect = 3e-04
Identities = 14/42 (33%), Positives = 23/42 (54%)
Query: 12 VFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHR 53
+ R L N+ C +CN+ P + T+ F+C+ CS +HR
Sbjct: 14 IIRGLMKLPPNRRCINCNSLGPQYVCTTFWTFVCMACSGIHR 55
>At2g14490 unknown protein
Length = 119
Score = 38.1 bits (87), Expect = 8e-04
Identities = 21/46 (45%), Positives = 24/46 (51%), Gaps = 2/46 (4%)
Query: 14 RKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHI 59
R L + +N+ C DC A P WA I L C VHRSLG HI
Sbjct: 19 RDLLNQPDNRVCADCGASVPKWAKY-QSIHLLKSC-GVHRSLGTHI 62
>At4g33680 unknown protein
Length = 461
Score = 25.4 bits (54), Expect = 5.5
Identities = 12/33 (36%), Positives = 15/33 (45%)
Query: 7 TDKNAVFRKLKTKSENKSCFDCNAKNPTWASVT 39
T +N F L T F C+ NPT A+ T
Sbjct: 219 TPENGFFPDLSTVGRTDIIFFCSPNNPTGAAAT 251
>At1g48570 unknown protein (At1g48570)
Length = 455
Score = 25.0 bits (53), Expect = 7.2
Identities = 12/33 (36%), Positives = 18/33 (54%), Gaps = 3/33 (9%)
Query: 20 SENKSCFDCNA---KNPTWASVTYGIFLCIDCS 49
+ N+ C +CN + P A V G +LC +CS
Sbjct: 314 ARNERCRECNEVADRRPVAAVVKEGDWLCPECS 346
>At1g19485 unknown protein
Length = 625
Score = 24.6 bits (52), Expect = 9.4
Identities = 10/27 (37%), Positives = 13/27 (48%)
Query: 9 KNAVFRKLKTKSENKSCFDCNAKNPTW 35
KN +RKL T SC+ C + W
Sbjct: 307 KNQKYRKLHTHDIRISCWKCTEQTVLW 333
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.324 0.132 0.419
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,408,528
Number of Sequences: 26719
Number of extensions: 42437
Number of successful extensions: 169
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 19
Number of HSP's successfully gapped in prelim test: 1
Number of HSP's that attempted gapping in prelim test: 149
Number of HSP's gapped (non-prelim): 21
length of query: 63
length of database: 11,318,596
effective HSP length: 39
effective length of query: 24
effective length of database: 10,276,555
effective search space: 246637320
effective search space used: 246637320
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 52 (24.6 bits)
Medicago: description of AC148483.13


