

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148482.2 + phase: 0
(530 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
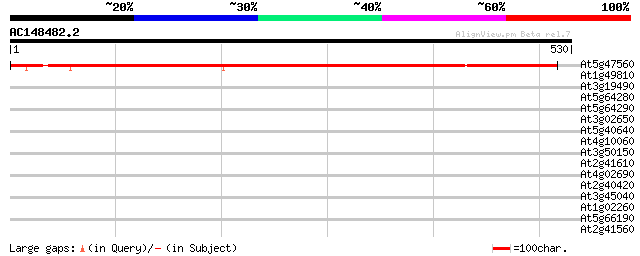
Score E
Sequences producing significant alignments: (bits) Value
At5g47560 sodium-dicarboxylate cotransporter-like 683 0.0
At1g49810 Na+/H+ antiporter, putative 35 0.079
At3g19490 unknown protein 34 0.18
At5g64280 2-oxoglutarate/malate translocator 34 0.23
At5g64290 2-oxoglutarate/malate translocator 33 0.51
At3g02650 hypothetical protein 32 0.87
At5g40640 unknown protein 31 1.9
At4g10060 putative protein 30 2.5
At3g50150 putative protein 30 2.5
At2g41610 hypothetical protein 30 3.3
At4g02690 putative glutamate-/aspartate-binding peptide 29 5.7
At2g40420 unknown protein 29 5.7
At3g45040 unknown protein 29 7.4
At1g02260 unknown protein 29 7.4
At5g66190 ferredoxin-NADP+ reductase 28 9.6
At2g41560 putative Ca2+-ATPase 28 9.6
>At5g47560 sodium-dicarboxylate cotransporter-like
Length = 540
Score = 683 bits (1762), Expect = 0.0
Identities = 337/527 (63%), Positives = 418/527 (78%), Gaps = 14/527 (2%)
Query: 1 MAGEDSLNATLLPL---QEPIQ-ETPNNNSITIFSILNLQNFYIILGPLLSIIICLLVKL 56
+AG D L + LLP+ EP + +T TIF+ +N YI LGPLL ++CL V L
Sbjct: 8 VAGSDDLKSPLLPVVHNDEPFERQTVGQQLRTIFTP---KNCYIALGPLLCAVVCLCVDL 64
Query: 57 --DAPTTSTKMLGVIAWVFTWWITNAVPLPVTSMCPLFLFPLFGIASADTVAHSYMDDVI 114
D TT+ MLGV+ W+F WW+T AVP+P+TSM PLFLFPLFGI++AD VA+SYMDDVI
Sbjct: 65 GGDETTTARNMLGVLVWMFAWWLTEAVPMPITSMTPLFLFPLFGISAADDVANSYMDDVI 124
Query: 115 TLVLGSFILALAVERYNVHRRLALNVTLVFCGDPLNPSMLLLGLCATTFFVSMWLHNVAT 174
+LVLGSFILALAVE YN+HRRLALN+TLVFC +PLN +LLLG+CATT FVSMW+HNVA
Sbjct: 125 SLVLGSFILALAVEHYNIHRRLALNITLVFCVEPLNAPLLLLGICATTAFVSMWMHNVAA 184
Query: 175 AVMMMPVATGILHRLPPAEEQPELMN----KFSRAVILTVVYATPIGGISTLTGTGVNLI 230
AVMMMPVATGIL RLP + E+++ KFSRAV+L V+Y+ +GG+STLTGTGVNLI
Sbjct: 185 AVMMMPVATGILQRLPSSSSTTEVVHPAVGKFSRAVVLGVIYSAAVGGMSTLTGTGVNLI 244
Query: 231 IIGMWKSLAPEAKPISFNSWFFFGFPVAVLLLLCFWCILCLIYVRKGSSRALSDYLDRAL 290
++GMWKS PEA PISF+ WFFFGFP+A+ + + WC+LC++Y KG+ +ALS YL ++
Sbjct: 245 LVGMWKSYFPEADPISFSQWFFFGFPLALCIFVVLWCVLCVMYCPKGAGQALSPYLHKSH 304
Query: 291 LKRDLEALGPMSFAEKMVLCVFGLLIVLWMTRRISDDLPGWGVLFHGLVGDGSISVLAAV 350
L+R+L+ LGPM+FAEKMVL VFG L+VLWMTR I+DD+PGWG +F G GDG++SV+ A
Sbjct: 305 LRRELDLLGPMNFAEKMVLAVFGGLVVLWMTRNITDDIPGWGRIFAGRAGDGTVSVMMAT 364
Query: 351 LLFIIPNMKQNGEKLMDWNECKKLPWNLILLLGAGFALADGVQSSGLADVMSKALEFLKD 410
LLFIIP+ + GEKLMDWN+CKKLPWN++LLLGAGFA+ADGV++SGLA+V+SK L FL+
Sbjct: 365 LLFIIPSNIKKGEKLMDWNKCKKLPWNIVLLLGAGFAIADGVRTSGLAEVLSKGLVFLET 424
Query: 411 VPYLAIVPAVSLLCSIITEFITSNDATATLLVPLLYHIAINMHVHPLLLMIPGGIATEFA 470
PY AI P V L+ + ITEF TSN+AT TLLVPLL IA NM +HPLLLM+PG I +FA
Sbjct: 425 APYWAIAPTVCLIAATITEF-TSNNATTTLLVPLLIEIAKNMGIHPLLLMVPGAIGAQFA 483
Query: 471 FWLPTSTPSNVVGFATGHIEIKDMLKVGVPLKVAGVSVLALLMPTLG 517
F LPT TPSNVVGF TGHIEIKDM+K G+PLK+AG L++LMPTLG
Sbjct: 484 FLLPTGTPSNVVGFTTGHIEIKDMIKTGLPLKIAGTIFLSILMPTLG 530
>At1g49810 Na+/H+ antiporter, putative
Length = 420
Score = 35.4 bits (80), Expect = 0.079
Identities = 33/120 (27%), Positives = 57/120 (47%), Gaps = 18/120 (15%)
Query: 106 AHSYMDDVITLVLGSFILALAVERYNVHRRLALNVTLVFCGDPLNPSMLLLGLCATTFFV 165
A S + ++ +LG+ + VE + H+ L + C P +LL + TFF+
Sbjct: 53 ATSEVSQIVFYMLGAMTI---VEIIDAHQGFKL---VTDCITSRKPKILLWVIGFATFFL 106
Query: 166 SMWLHNVATAVMMMPVATGILHRLPPAEEQPELMNKFSRAVILTVVYA----TPIGGIST 221
S L N+ + ++M+ +L RL P E +L+ AV++ A TPIG ++T
Sbjct: 107 SSVLDNLTSTIVMV----SLLRRLIPPSEYRKLLG----AVVVIAANAGGAWTPIGDVTT 158
>At3g19490 unknown protein
Length = 576
Score = 34.3 bits (77), Expect = 0.18
Identities = 40/191 (20%), Positives = 79/191 (40%), Gaps = 34/191 (17%)
Query: 76 WITNAVPLPVTSMCPLFLFPLFGIASADTVAHSYMDDVITLVLGSFILALAVERYNVHRR 135
W+ ++ P T + L L A + + +++ +LG+ + V+ + +
Sbjct: 176 WVVRSIGAPSTEIAVLDL----------QHATAEVSEIVFFLLGAMTIVEIVDAHQGFKL 225
Query: 136 LALNVTLVFCGDPLNPSMLLLGLCATTFFVSMWLHNVATAVMMMPVATGILHRLPPAEEQ 195
+ N+T P LL + TFF+S L N+ + ++M+ ++ +L P E
Sbjct: 226 VTDNITT------RKPKTLLWVVGFVTFFLSSILDNLTSTIVMV----SLIRKLVPQSEY 275
Query: 196 PELMNKFSRAVILTVVYA----TPIGGISTLTGTGVNLIIIGMWKSLAPEAKPISFNSWF 251
+L+ V++ A TPIG ++T ++ I S P K + S
Sbjct: 276 RKLLG----GVVVIAANAGGAWTPIGDVTT------TMLWIHGQISTLPTMKDLFLPSVV 325
Query: 252 FFGFPVAVLLL 262
P+A++ L
Sbjct: 326 SLAVPLALMSL 336
>At5g64280 2-oxoglutarate/malate translocator
Length = 549
Score = 33.9 bits (76), Expect = 0.23
Identities = 88/459 (19%), Positives = 169/459 (36%), Gaps = 31/459 (6%)
Query: 46 LSIIICLLVKLDAPTTST--KMLGVIAWVFTWWITNAVPLPVTSMCPLFLFPLFGIASAD 103
+ +I+ L+ TS ++L + + + + +P+ + L +
Sbjct: 91 IGLIVRFLIPRPEQVTSQGWQLLSIFLFTISGLVLGPLPVGAWAFIGLTASIVTKTLPFS 150
Query: 104 TVAHSYMDDVITLVLGSFILALAVERYNVHRRLALNVTLVFCGDPLNPSMLLLGLCATTF 163
T ++ +++I L+ SF A + + R+A + + G L C T
Sbjct: 151 TAFAAFTNELIWLIAISFFFARGFIKTGLGDRIA-TYFVKWLGKSTLGLSYGLAFCETLM 209
Query: 164 FVSMWLHNVATAVMMMPVATGILHRLPPAEEQPELMNKFSRAVILTVVYATPIGGISTLT 223
+ M + +PV + P K +I T + + G LT
Sbjct: 210 GLIMPSTMARAGGVFLPVIKSLAISAGSYPGDPS-SRKLGSFLIQTQLQCSGASGAILLT 268
Query: 224 GTGVNLIIIGMWKSLAPEAKPISFNSWFFFGFPVAVLLLLCFWCILCLIYVRKGSSRALS 283
NL+ + LA E + N W + +V + C +IY +
Sbjct: 269 SAAQNLLCL----KLAREVGVVISNPWITWFKVASVPAFVSLLCTPLIIYKLYPPELKHT 324
Query: 284 DYLDRALLKRDLEALGPMSFAEKMVLCVFGLLIVLWMTRRISDDLPGWGVLFHGLVGDGS 343
A K+ LE LGP++ E ++L + LW + + G + ++G
Sbjct: 325 PEAPAAAAKK-LERLGPITKNEWIMLGAMAFTVSLW----VFGEAIGIASVVSAMIG--- 376
Query: 344 ISVLAAVLLFIIPNMKQNGEKLMDWNECKKLPWNLILLLGAGFALADGVQSSGLADVMSK 403
L+ +LL + N + L D + L W +L+ AG GV + ++D ++K
Sbjct: 377 ---LSTLLLLGVINW---DDCLSDKSAWDSLTWFAVLIGMAGQLTNLGV-VAWMSDCVAK 429
Query: 404 ALEFLKDVPYLAIVPAVSLLCSIITEFITSNDATATLLVP--LLYHIAINMH--VHPLLL 459
L+ L + + A + +I S A L P L IA + + L L
Sbjct: 430 LLQSL-SLTWPASFIILQACYLLIHYLFASQTGHAGALYPPFLAMQIAAGVPGVLAALCL 488
Query: 460 MIPGGIATEFAFWLPTSTPSNVVGFATGHIEIKDMLKVG 498
++ A + S + + G+++++DM +VG
Sbjct: 489 AFNNNLSGALAHY---SGGPAALYYGAGYVDLRDMFRVG 524
>At5g64290 2-oxoglutarate/malate translocator
Length = 563
Score = 32.7 bits (73), Expect = 0.51
Identities = 61/291 (20%), Positives = 113/291 (37%), Gaps = 47/291 (16%)
Query: 222 LTGTGVNLIIIGMWKSLAPEAKPISFN---SWFFFGFPVAVLLLLCFWCILCLIYVRKGS 278
LT NL+ + LA E + N SWF A++ LLC IL +Y +
Sbjct: 281 LTAAAQNLLCL----KLAEELGVVISNPWVSWFKAASLPAIISLLCTPLILYKLYPPETK 336
Query: 279 SRALSDYLDRALLKRDLEALGPMSFAEKMVLCVFGLLIVLWMTRRISDDLPGWGVLFHGL 338
+ + LK+ +GP++ E +++ L + LW I + G + +
Sbjct: 337 DTPEAPGIAATKLKQ----MGPVTKNEWIMVGTMLLAVTLW----ICGETLGIPSVVAAM 388
Query: 339 VGDGSISVLAAVLLFIIPNMKQNGEKLMDWNEC--KKLPWNLILLLGAGFALADGVQSSG 396
+G + VL +++W++C +K W+ + +A + + G
Sbjct: 389 IGLSILLVLG----------------VLNWDDCLSEKSAWDTLAWFAVLVGMAGQLTNLG 432
Query: 397 ----LADVMSKALEFLKDVPYLAIVPAVSLLCSIITEFITSNDATATLLVPLLYHIAINM 452
++D ++K L+ L PA L FI A+ T V L+ + M
Sbjct: 433 VVTWMSDCVAKVLQSLS-----LSWPAAFGLLQAAYFFIHYLFASQTGHVGALFSAFLAM 487
Query: 453 HVHPLLLMIPGGIATE-----FAFWLPTSTPSNVVGFATGHIEIKDMLKVG 498
H+ + I +A F S+ V + G++++ D+ K+G
Sbjct: 488 HIAAGVPGILAALALAYNTNLFGALTHYSSGQAAVYYGAGYVDLPDVFKIG 538
>At3g02650 hypothetical protein
Length = 1077
Score = 32.0 bits (71), Expect = 0.87
Identities = 25/80 (31%), Positives = 35/80 (43%), Gaps = 7/80 (8%)
Query: 185 ILHRLPPAEEQPELMNKFSRAVILTVVYATPIGGISTLTGTGVNLIIIGMWK-SLAPEAK 243
++ RL +E E+ N S++V LT V G T VN G WK + +
Sbjct: 436 LVSRLKSDQEAAEMWNGMSKSVRLTKV------GFLDKTIEDVNRYYTGRWKVKIGRLVE 489
Query: 244 PISFNSWFFFGFPVAVLLLL 263
+ SW F AVLLL+
Sbjct: 490 VYVYGSWQILAFLAAVLLLI 509
>At5g40640 unknown protein
Length = 586
Score = 30.8 bits (68), Expect = 1.9
Identities = 26/96 (27%), Positives = 43/96 (44%), Gaps = 16/96 (16%)
Query: 28 TIFSILNLQNFYIILGPLLSIIICLLVKLDAPTTSTKMLGVIAWVFTWWITNAVPLPVTS 87
T++SI + + LGP+L I +CL V LGVI W+ + + + +
Sbjct: 62 TLYSIASAKQ----LGPILKIFLCLCVP----------LGVILWLVVSILGSVLGGAIYG 107
Query: 88 -MCPLF-LFPLFGIASADTVAHSYMDDVITLVLGSF 121
+ P+F F G ++ H + D + V GSF
Sbjct: 108 FLSPIFATFDAVGEGKSNPFFHCFYDGTWSTVQGSF 143
>At4g10060 putative protein
Length = 750
Score = 30.4 bits (67), Expect = 2.5
Identities = 20/56 (35%), Positives = 26/56 (45%), Gaps = 8/56 (14%)
Query: 193 EEQPELMNKFSRA--------VILTVVYATPIGGISTLTGTGVNLIIIGMWKSLAP 240
EE P L N+FS + V L + +GG S LTG N II + + L P
Sbjct: 68 EEAPILTNQFSMSNLGKEEATVTLLFTWENSVGGASGLTGEHFNSTIIHLKRGLVP 123
>At3g50150 putative protein
Length = 509
Score = 30.4 bits (67), Expect = 2.5
Identities = 24/79 (30%), Positives = 32/79 (40%), Gaps = 6/79 (7%)
Query: 185 ILHRLPPAEEQPELMNKFSRAVILTVVYATPIGGISTLTGTGVNLIIIGMWKSLAPEAKP 244
I H L E +L N+ + VI P G + VN W SL +
Sbjct: 417 IEHWLGSDSEVADLFNRLCKEVIFD-----PKDGYLSQLSREVNRYYSRKWNSLKATLRQ 471
Query: 245 ISFNS-WFFFGFPVAVLLL 262
FN+ W +F F AV+LL
Sbjct: 472 KYFNNPWAYFSFSAAVILL 490
>At2g41610 hypothetical protein
Length = 304
Score = 30.0 bits (66), Expect = 3.3
Identities = 38/120 (31%), Positives = 51/120 (41%), Gaps = 8/120 (6%)
Query: 205 AVILTVVYATPIGGISTLTGTGVNLIIIGMWKSLAPEAKPISFNSWFFFGFPVAVLLLLC 264
AVIL Y T + T T +GV I + L K +S+ W F V +L L
Sbjct: 163 AVILP--YYTGFDALVTSTFSGVCKSCICRKEPLIVGGKIVSYRGWSSTTFLVVGVLFLR 220
Query: 265 FWCILCL---IYVRKGSSRALSDYLDRALLKRDLEALGPMSFAEKMVL---CVFGLLIVL 318
C LC I R + + L +L RD L MS E+ +L VFG L++L
Sbjct: 221 IICKLCKEEGINKRVLVVKNVVQGLTLLVLIRDCVYLAVMSPVEEPILFRVFVFGFLLLL 280
>At4g02690 putative glutamate-/aspartate-binding peptide
Length = 248
Score = 29.3 bits (64), Expect = 5.7
Identities = 52/177 (29%), Positives = 70/177 (39%), Gaps = 32/177 (18%)
Query: 181 VATGILHRLP---PA-EEQPELMNKFSR--------------AVILTVVYATPIGGISTL 222
V TG+ R P PA E PEL F R AV TVV PI
Sbjct: 13 VETGVSSRRPLLYPAMHENPELRWGFIRKVYSIIAFQLLATVAVAATVVTVRPIALFFAT 72
Query: 223 TGTGVNL---IIIGMWKSLAP-----EAKPISFNSWFFFGFPVAVLL-LLCFWC----IL 269
TG G+ L III L P + P+++ F +A ++ L C + IL
Sbjct: 73 TGLGLALYIVIIITPLIVLCPLYYYHQKHPVNYLLLGIFTLALAFVVGLTCAFTNGKVIL 132
Query: 270 CLIYVRKGSSRALSDYLDRALLKR-DLEALGPMSFAEKMVLCVFGLLIVLWMTRRIS 325
+ + +L+ Y A K D LGP F VL F L+ +L+ R+S
Sbjct: 133 ESVILTSVVVLSLTLYTFWAARKGYDFNFLGPFLFGALTVLIFFALIQILFPLGRVS 189
>At2g40420 unknown protein
Length = 415
Score = 29.3 bits (64), Expect = 5.7
Identities = 29/119 (24%), Positives = 50/119 (41%), Gaps = 8/119 (6%)
Query: 6 SLNATLLPLQEPIQETPNNNSITIFSILNLQNFYIILGPL-----LSIIICLLVKLDAPT 60
++ A LLP QEP + N+ ++ N+ + G + ++ L+K
Sbjct: 4 AIKAPLLPNQEPSSSSSENHGSFAGAVFNISTSIVGAGIMAIPAAFKVLAGFLMKSSLAG 63
Query: 61 TSTKMLGVIAWVFTWWITNAVPLPVTSMCPLF-LFPLFGIASADTVAHSYMDDVITLVL 118
ST GV+ F + AV + V +M F +F I D ++ + D +I L L
Sbjct: 64 ESTTYAGVMKESF--GKSGAVAVTVVTMVVTFGSMIIFSIIIGDVISGNEKDGIIHLGL 120
>At3g45040 unknown protein
Length = 569
Score = 28.9 bits (63), Expect = 7.4
Identities = 15/59 (25%), Positives = 30/59 (50%)
Query: 251 FFFGFPVAVLLLLCFWCILCLIYVRKGSSRALSDYLDRALLKRDLEALGPMSFAEKMVL 309
F F P+ L L +W +L ++ V + + + S ++R LL++ + + F +VL
Sbjct: 332 FVFSEPLKRLSLCIYWILLIVVSVSRFYNISRSSKVERILLRKYYHLMAVLMFLPALVL 390
>At1g02260 unknown protein
Length = 502
Score = 28.9 bits (63), Expect = 7.4
Identities = 13/54 (24%), Positives = 25/54 (46%)
Query: 208 LTVVYATPIGGISTLTGTGVNLIIIGMWKSLAPEAKPISFNSWFFFGFPVAVLL 261
L + + + + G TL G+ NLI+ + ++F F FG P +++
Sbjct: 440 LLLAWVSTVAGNLTLLGSAANLIVCEQARRAVSHGYTLTFTKHFKFGLPSTLIV 493
>At5g66190 ferredoxin-NADP+ reductase
Length = 360
Score = 28.5 bits (62), Expect = 9.6
Identities = 15/42 (35%), Positives = 20/42 (46%), Gaps = 2/42 (4%)
Query: 223 TGTGVNLIIIGMWKSLAPEAKPISFN--SWFFFGFPVAVLLL 262
TGTG+ +WK E + FN +W F G P + LL
Sbjct: 216 TGTGIAPFRSFLWKMFFEEHEDYKFNGLAWLFLGVPTSSSLL 257
>At2g41560 putative Ca2+-ATPase
Length = 1030
Score = 28.5 bits (62), Expect = 9.6
Identities = 18/58 (31%), Positives = 32/58 (55%), Gaps = 7/58 (12%)
Query: 70 AWVFTWWITNAVPLPVTSMCPLFLFPLFGIASADTVAHSYMDDVITLVLGSFILALAV 127
+WVFTW +T VT + + + G A A TV S+ ++++++GS + +AV
Sbjct: 951 SWVFTWVMT------VTVVFQVIIVEFLG-AFASTVPLSWQHWLLSILIGSLNMIVAV 1001
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.327 0.142 0.440
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,435,077
Number of Sequences: 26719
Number of extensions: 474686
Number of successful extensions: 1402
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 13
Number of HSP's that attempted gapping in prelim test: 1392
Number of HSP's gapped (non-prelim): 18
length of query: 530
length of database: 11,318,596
effective HSP length: 104
effective length of query: 426
effective length of database: 8,539,820
effective search space: 3637963320
effective search space used: 3637963320
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 62 (28.5 bits)
Medicago: description of AC148482.2


