

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148469.7 - phase: 0
(540 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
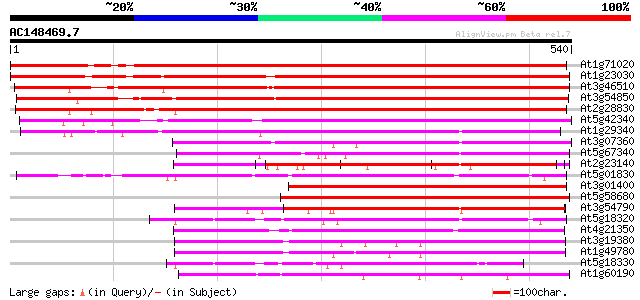
Score E
Sequences producing significant alignments: (bits) Value
At1g71020 unknown protein 634 0.0
At1g23030 unknown protein 633 0.0
At3g46510 arm repeat containing protein homolog 452 e-127
At3g54850 putative protein 440 e-124
At2g28830 unknown protein 416 e-116
At5g42340 arm repeat containing protein 365 e-101
At1g29340 arm repeat-containing protein, putative 279 2e-75
At3g07360 unknown protein 255 4e-68
At5g67340 unknown proteins 251 7e-67
At2g23140 hypothetical protein 236 2e-62
At5g01830 unknown protein 234 9e-62
At3g01400 unknown protein 228 5e-60
At5g58680 unknown protein 221 1e-57
At3g54790 unknown protein 210 2e-54
At5g18320 putative protein 203 2e-52
At4g21350 putative protein 175 7e-44
At3g19380 unknown protein 164 1e-40
At1g49780 unknown protein 159 4e-39
At5g18330 putative protein 157 1e-38
At1g60190 155 7e-38
>At1g71020 unknown protein
Length = 628
Score = 634 bits (1636), Expect = 0.0
Identities = 354/538 (65%), Positives = 411/538 (75%), Gaps = 18/538 (3%)
Query: 1 DGAAKKIIFQFQRVTWKLEKLLSSLPYDDLDISEEVKEQVDLVRNQLRRATDKYGFMISK 60
DGAAK+I FQFQ VTWKLEK L L YD DIS+EV+EQV+L R QLRRA +YG + SK
Sbjct: 99 DGAAKRISFQFQCVTWKLEKALGDLTYDRYDISDEVREQVELARLQLRRAMQRYGSLNSK 158
Query: 61 MPSFDSSQPLAQEISQVLGQSVSGLHKQHSCPENLSELGSIPKSNEGKSCNPFGAGSRLE 120
S S+P+ ++ S + + K S PE + L S+E K +P
Sbjct: 159 KFSSGLSEPMEKDASS----NRKVIEKLESIPETVHSL-----SDEKKFESP-------P 202
Query: 121 RTRSIHASSEVSFSIKTAPESQEISGSGNLPEVKKPDAIVIPEDFLCPISLELMRDPVIV 180
+S S S E E + + N + +K D + IPEDFLCPISLELM+DP IV
Sbjct: 203 PWKSSSVSLAFFLSKDGDDERLEKAVTENSDDSQKSDNLTIPEDFLCPISLELMKDPAIV 262
Query: 181 ATGQTYERSYIQRWIDCGNTTCPKTQQKLQHLTLTPNYVLRSLVSQWCIEHNIEQPTGLT 240
+TGQTYERS+IQRWIDCGN +CPKTQQKL++ TLTPNYVLRSL+SQWC +HNIEQP G
Sbjct: 263 STGQTYERSFIQRWIDCGNLSCPKTQQKLENFTLTPNYVLRSLISQWCTKHNIEQPGGYM 322
Query: 241 NGKIKKSDGSFRDVTGDIAAIETLVRKLSCRSVEESRAAVAEIRSLSKRSTDNRILIAEA 300
NG+ K SDGSFRD++GD++AI LV KLS +S+E+ R AV+EIRSLSKRSTDNRILIAEA
Sbjct: 323 NGRTKNSDGSFRDLSGDMSAIRALVCKLSSQSIEDRRTAVSEIRSLSKRSTDNRILIAEA 382
Query: 301 GAIPVLVSLLTSE-DVMTQENAVTSILNLSIYENNKGLIMLAGAIPSIVQVLRAGTMEAR 359
GAIPVLV LLTS+ D TQENAVT ILNLSIYE+NK LIMLAGA+ SIV VLRAG+MEAR
Sbjct: 383 GAIPVLVKLLTSDGDTETQENAVTCILNLSIYEHNKELIMLAGAVTSIVLVLRAGSMEAR 442
Query: 360 ENAAATLFSLSLADENKIIIGASGAISALVDLLQNGSPRGKKDAATALFNLCIYQGNKGR 419
ENAAATLFSLSLADENKIIIGASGAI ALVDLLQ GS RGKKDAATALFNLCIYQGNKGR
Sbjct: 443 ENAAATLFSLSLADENKIIIGASGAIMALVDLLQYGSVRGKKDAATALFNLCIYQGNKGR 502
Query: 420 AIRAGIITALLNMLTD-SSKSMVDEALTIMSVLASHQEAKVSIVKASTIPVLIDLLRTGL 478
A+RAGI+ L+ MLTD SS+ M DEALTI+SVLAS+Q AK +I++A+ IP LID L+
Sbjct: 503 AVRAGIVKPLVKMLTDSSSERMADEALTILSVLASNQVAKTAILRANAIPPLIDCLQKDQ 562
Query: 479 PRNKENAAAILLALCKRDTDNLSCISRLGAVIPLSELARTGTERAKRKATSLLEHLRK 536
PRN+ENAAAILL LCKRDT+ L I RLGAV+PL EL+R GTERAKRKA SLLE LRK
Sbjct: 563 PRNRENAAAILLCLCKRDTEKLISIGRLGAVVPLMELSRDGTERAKRKANSLLELLRK 620
>At1g23030 unknown protein
Length = 612
Score = 633 bits (1633), Expect = 0.0
Identities = 350/540 (64%), Positives = 414/540 (75%), Gaps = 22/540 (4%)
Query: 1 DGAAKKIIFQFQRVTWKLEKLLSSLPYDDLDISEEVKEQVDLVRNQLRRATDKYGFMISK 60
DGAAK+I FQFQ VTWKLEK LS+LPYD DIS+EV EQV+L R+QLRRA +YG + S
Sbjct: 94 DGAAKRISFQFQCVTWKLEKALSNLPYDLYDISDEVGEQVELARSQLRRAMQRYGSLNSN 153
Query: 61 MPSFDSSQPLAQEISQVLGQSVSGLHKQHSCPENLSELGSIPKSNEGKSCNPFGAGSRLE 120
S S+P+ ++ G S K E++SE + E +S P L
Sbjct: 154 KFSSALSEPMERD-----GFSNVIKIKAEEKLESVSETLHFGEEEEKQSSPP------LR 202
Query: 121 RTRSIHASSEVSFSIKTAPESQEISGSGNLPEVKKPDAIVIPEDFLCPISLELMRDPVIV 180
R+ SI + +S T + ++ N E KK D + IP DFLCP+SLELM+DPVIV
Sbjct: 203 RSSSISLAYYLSKDADTDRLDKMVNK--NTDESKKSDKLTIPVDFLCPVSLELMKDPVIV 260
Query: 181 ATGQTYERSYIQRWIDCGNTTCPKTQQKLQHLTLTPNYVLRSLVSQWCIEHNIEQPTGLT 240
ATGQTYER+YIQRWIDCGN TCPKTQQKL++ TLTPNYVLRSL+S+WC EHNIEQP G
Sbjct: 261 ATGQTYERAYIQRWIDCGNLTCPKTQQKLENFTLTPNYVLRSLISRWCAEHNIEQPAGYI 320
Query: 241 NGKIKKSDGSFRDVTGDIAAIETLVRKLSCRSVEESRAAVAEIRSLSKRSTDNRILIAEA 300
NG+ K S GD++ I LV++LS RS E+ R AV+EIRSLSKRSTDNRILIAEA
Sbjct: 321 NGRTKNS--------GDMSVIRALVQRLSSRSTEDRRNAVSEIRSLSKRSTDNRILIAEA 372
Query: 301 GAIPVLVSLLTSEDVMTQENAVTSILNLSIYENNKGLIMLAGAIPSIVQVLRAGTMEARE 360
GAIPVLV+LLTSEDV TQENA+T +LNLSIYENNK LIM AGA+ SIVQVLRAGTMEARE
Sbjct: 373 GAIPVLVNLLTSEDVATQENAITCVLNLSIYENNKELIMFAGAVTSIVQVLRAGTMEARE 432
Query: 361 NAAATLFSLSLADENKIIIGASGAISALVDLLQNGSPRGKKDAATALFNLCIYQGNKGRA 420
NAAATLFSLSLADENKIIIG SGAI ALVDLL+NG+PRGKKDAATALFNLCIY GNKGRA
Sbjct: 433 NAAATLFSLSLADENKIIIGGSGAIPALVDLLENGTPRGKKDAATALFNLCIYHGNKGRA 492
Query: 421 IRAGIITALLNMLTDSSK-SMVDEALTIMSVLASHQEAKVSIVKASTIPVLIDLLRTGLP 479
+RAGI+TAL+ ML+DS++ MVDEALTI+SVLA++Q+AK +IVKA+T+P LI +L+T
Sbjct: 493 VRAGIVTALVKMLSDSTRHRMVDEALTILSVLANNQDAKSAIVKANTLPALIGILQTDQT 552
Query: 480 RNKENAAAILLALCKRDTDNLSCISRLGAVIPLSELARTGTERAKRKATSLLEHLRKLQQ 539
RN+ENAAAILL+LCKRDT+ L I RLGAV+PL +L++ GTER KRKA SLLE LRK Q
Sbjct: 553 RNRENAAAILLSLCKRDTEKLITIGRLGAVVPLMDLSKNGTERGKRKAISLLELLRKACQ 612
>At3g46510 arm repeat containing protein homolog
Length = 660
Score = 452 bits (1162), Expect = e-127
Identities = 258/557 (46%), Positives = 360/557 (64%), Gaps = 41/557 (7%)
Query: 5 KKIIFQFQRVTWKLEKLLSSLPYDDLDISEEVKEQVDLVRNQLRRATDKYGF-------- 56
+++ + V+ KLE+ LS +PY++LDIS+EV+EQV+LV +Q RRA +
Sbjct: 95 EQVTSKLMEVSVKLEQSLSQIPYEELDISDEVREQVELVLSQFRRAKGRVDVSDDELYED 154
Query: 57 ---MISKMPSFDSSQPLAQEISQVLGQSVSGLHKQHSCPENLSELGSIPKSNEGKSCNPF 113
+ +K D+ QP+ + +++ L +L E+ + + E + +
Sbjct: 155 LQSLCNKSSDVDAYQPVLERVAKKL---------------HLMEIPDL--AQESVALHEM 197
Query: 114 GAGSRLERTRSIHASSEVSFSIKTAPESQEISG-----------SGNLPEVKKPDAIVIP 162
A S + +I + V IK ++++ +G +G VIP
Sbjct: 198 VASSGGDVGENIEEMAMVLKMIKDFVQTEDDNGEEQKVGVNSRSNGQTSTAASQKIPVIP 257
Query: 163 EDFLCPISLELMRDPVIVATGQTYERSYIQRWIDCGNTTCPKTQQKLQHLTLTPNYVLRS 222
+DF CPISLE+MRDPVIV++GQTYER+ I++WI+ G++TCPKTQQ L TLTPNYVLRS
Sbjct: 258 DDFRCPISLEMMRDPVIVSSGQTYERTCIEKWIEGGHSTCPKTQQALTSTTLTPNYVLRS 317
Query: 223 LVSQWCIEHNIEQPTGLTNGKIKKSDGSFRDVTGDIAAIETLVRKLSCRSVEESRAAVAE 282
L++QWC ++IE P ++ + +K SF + IE L+ +L+ + E+ R+A E
Sbjct: 318 LIAQWCEANDIEPPKPPSSLRPRKVS-SFSS-PAEANKIEDLMWRLAYGNPEDQRSAAGE 375
Query: 283 IRSLSKRSTDNRILIAEAGAIPVLVSLLTSEDVMTQENAVTSILNLSIYENNKGLIMLAG 342
IR L+KR+ DNR+ IAEAGAIP+LV LL++ D QE++VT++LNLSI ENNKG I+ AG
Sbjct: 376 IRLLAKRNADNRVAIAEAGAIPLLVGLLSTPDSRIQEHSVTALLNLSICENNKGAIVSAG 435
Query: 343 AIPSIVQVLRAGTMEARENAAATLFSLSLADENKIIIGASGAISALVDLLQNGSPRGKKD 402
AIP IVQVL+ G+MEARENAAATLFSLS+ DENK+ IGA GAI LV LL G+ RGKKD
Sbjct: 436 AIPGIVQVLKKGSMEARENAAATLFSLSVIDENKVTIGALGAIPPLVVLLNEGTQRGKKD 495
Query: 403 AATALFNLCIYQGNKGRAIRAGIITALLNMLTDSSKSMVDEALTIMSVLASHQEAKVSIV 462
AATALFNLCIYQGNKG+AIRAG+I L +LT+ MVDEAL I+++L+SH E K I
Sbjct: 496 AATALFNLCIYQGNKGKAIRAGVIPTLTRLLTEPGSGMVDEALAILAILSSHPEGKAIIG 555
Query: 463 KASTIPVLIDLLRTGLPRNKENAAAILLALCKRDTDNLSCISRLGAVIPLSELARTGTER 522
+ +P L++ +RTG PRN+ENAAA+L+ LC D +L +LG + PL +LA GT+R
Sbjct: 556 SSDAVPSLVEFIRTGSPRNRENAAAVLVHLCSGDPQHLVEAQKLGLMGPLIDLAGNGTDR 615
Query: 523 AKRKATSLLEHLRKLQQ 539
KRKA LLE + +L +
Sbjct: 616 GKRKAAQLLERISRLAE 632
>At3g54850 putative protein
Length = 632
Score = 440 bits (1132), Expect = e-124
Identities = 251/542 (46%), Positives = 354/542 (65%), Gaps = 28/542 (5%)
Query: 7 IIFQFQRVTWKLEKLLSSLPYDDLDISEEVKEQVDLVRNQLRRATDKYGFMISKMPS--- 63
++ +F+ +T ++E LS +PY+ +++SEEV+EQV L+ Q +RA +++ ++
Sbjct: 101 LVEKFRDMTVEIEAALSQIPYEKIEVSEEVREQVQLLHFQFKRAKERWEESDLQLSHDLA 160
Query: 64 -----FDSSQPLAQEISQVLG-QSVSGLHKQ-HSCPENLSELGSIPKSNEGKSCNPFGAG 116
D + + +SQ L ++ L K+ H+ E P C
Sbjct: 161 MAENVMDPDPIILKRLSQELQLTTIDELKKESHAIHEYFLSYDGDPDD-----C------ 209
Query: 117 SRLERTRSIHASSEVSFSIKTAPESQEISGSGNLPEVKKPDAIVIPEDFLCPISLELMRD 176
ER S+ + V F + + +GS + + P VIPE F CPISLELM+D
Sbjct: 210 --FERMSSL-LKNLVDFVTMESSDPDPSTGSRIVSRHRSP---VIPEYFRCPISLELMKD 263
Query: 177 PVIVATGQTYERSYIQRWIDCGNTTCPKTQQKLQHLTLTPNYVLRSLVSQWCIEHNIEQP 236
PVIV+TGQTYERS IQ+W+D G+ TCPK+Q+ L H LTPNYVL+SL++ WC + IE P
Sbjct: 264 PVIVSTGQTYERSSIQKWLDAGHKTCPKSQETLLHAGLTPNYVLKSLIALWCESNGIELP 323
Query: 237 TGLTNGKIKKSDGSFRDVTGDIAAIETLVRKLSCRSVEESRAAVAEIRSLSKRSTDNRIL 296
+ + K GS D + +L+ KL+ + E+ RAA E+R L+KR+ DNR+
Sbjct: 324 QNQGSCRTTKIGGSSSSDC-DRTFVLSLLEKLANGTTEQQRAAAGELRLLAKRNVDNRVC 382
Query: 297 IAEAGAIPVLVSLLTSEDVMTQENAVTSILNLSIYENNKGLIMLAGAIPSIVQVLRAGTM 356
IAEAGAIP+LV LL+S D TQE++VT++LNLSI E NKG I+ AGAI IV+VL+ G+M
Sbjct: 383 IAEAGAIPLLVELLSSPDPRTQEHSVTALLNLSINEGNKGAIVDAGAITDIVEVLKNGSM 442
Query: 357 EARENAAATLFSLSLADENKIIIGASGAISALVDLLQNGSPRGKKDAATALFNLCIYQGN 416
EARENAAATLFSLS+ DENK+ IGA+GAI AL+ LL+ G+ RGKKDAATA+FNLCIYQGN
Sbjct: 443 EARENAAATLFSLSVIDENKVAIGAAGAIQALISLLEEGTRRGKKDAATAIFNLCIYQGN 502
Query: 417 KGRAIRAGIITALLNMLTDSSKSMVDEALTIMSVLASHQEAKVSIVKASTIPVLIDLLRT 476
K RA++ GI+ L +L D+ MVDEAL I+++L+++QE K +I +A +IPVL++++RT
Sbjct: 503 KSRAVKGGIVDPLTRLLKDAGGGMVDEALAILAILSTNQEGKTAIAEAESIPVLVEIIRT 562
Query: 477 GLPRNKENAAAILLALCKRDTDNLSCISRLGAVIPLSELARTGTERAKRKATSLLEHLRK 536
G PRN+ENAAAIL LC + + L+ +GA + L EL GT+RAKRKA SLLE +++
Sbjct: 563 GSPRNRENAAAILWYLCIGNIERLNVAREVGADVALKELTENGTDRAKRKAASLLELIQQ 622
Query: 537 LQ 538
+
Sbjct: 623 TE 624
>At2g28830 unknown protein
Length = 654
Score = 416 bits (1068), Expect = e-116
Identities = 244/544 (44%), Positives = 348/544 (63%), Gaps = 18/544 (3%)
Query: 6 KIIFQFQRVTWKLEKLLSSLPYDDLDISEEVKEQVDLVRNQLRRATDKYGF------MIS 59
+++ +FQ+VT LE+ LS +PY++L+IS+E+KEQV+LV QLRR+ K G +
Sbjct: 96 QVMVKFQKVTSLLEQALSIIPYENLEISDELKEQVELVLVQLRRSLGKRGGDVYDDELYK 155
Query: 60 KMPSFDSSQPLAQEISQV--LGQSVSGLHKQHSCPENLSELGSIPKSNEGKSCNPFGAGS 117
+ S S + E V + + + + E+L+ L + S F S
Sbjct: 156 DVLSLYSGRGSVMESDMVRRVAEKLQLMTITDLTQESLALLDMVSSSGGDDPGESFEKMS 215
Query: 118 RLERTRSIHASSEVSFSIKTAPESQEISGSGNLPEVKKPDA--IVIPEDFLCPISLELMR 175
+ + + + ++ AP + +LP+ + D ++ PE+F CPISLELM
Sbjct: 216 MVLKKIKDFVQT-YNPNLDDAP----LRLKSSLPKSRDDDRDMLIPPEEFRCPISLELMT 270
Query: 176 DPVIVATGQTYERSYIQRWIDCGNTTCPKTQQKLQHLTLTPNYVLRSLVSQWCIEHNIEQ 235
DPVIV++GQTYER I++W++ G+ TCPKTQ+ L +TPNYVLRSL++QWC + IE
Sbjct: 271 DPVIVSSGQTYERECIKKWLEGGHLTCPKTQETLTSDIMTPNYVLRSLIAQWCESNGIEP 330
Query: 236 PTGLTNGK-IKKSDGSFRDVTGDIAAIETLVRKLSCRSVEESRAAVAEIRSLSKRSTDNR 294
P + K+ S + IE L+ KL+ + E+ R+A EIR L+K++ NR
Sbjct: 331 PKRPNISQPSSKASSSSSAPDDEHNKIEELLLKLTSQQPEDRRSAAGEIRLLAKQNNHNR 390
Query: 295 ILIAEAGAIPVLVSLLT-SEDVMTQENAVTSILNLSIYENNKGLIMLA-GAIPSIVQVLR 352
+ IA +GAIP+LV+LLT S D TQE+AVTSILNLSI + NKG I+ + GA+P IV VL+
Sbjct: 391 VAIAASGAIPLLVNLLTISNDSRTQEHAVTSILNLSICQENKGKIVYSSGAVPGIVHVLQ 450
Query: 353 AGTMEARENAAATLFSLSLADENKIIIGASGAISALVDLLQNGSPRGKKDAATALFNLCI 412
G+MEARENAAATLFSLS+ DENK+ IGA+GAI LV LL GS RGKKDAATALFNLCI
Sbjct: 451 KGSMEARENAAATLFSLSVIDENKVTIGAAGAIPPLVTLLSEGSQRGKKDAATALFNLCI 510
Query: 413 YQGNKGRAIRAGIITALLNMLTDSSKSMVDEALTIMSVLASHQEAKVSIVKASTIPVLID 472
+QGNKG+A+RAG++ L+ +LT+ MVDE+L+I+++L+SH + K + A +PVL+D
Sbjct: 511 FQGNKGKAVRAGLVPVLMRLLTEPESGMVDESLSILAILSSHPDGKSEVGAADAVPVLVD 570
Query: 473 LLRTGLPRNKENAAAILLALCKRDTDNLSCISRLGAVIPLSELARTGTERAKRKATSLLE 532
+R+G PRNKEN+AA+L+ LC + +L +LG + L E+A GT+R KRKA LL
Sbjct: 571 FIRSGSPRNKENSAAVLVHLCSWNQQHLIEAQKLGIMDLLIEMAENGTDRGKRKAAQLLN 630
Query: 533 HLRK 536
+
Sbjct: 631 RFSR 634
>At5g42340 arm repeat containing protein
Length = 656
Score = 365 bits (936), Expect = e-101
Identities = 216/547 (39%), Positives = 323/547 (58%), Gaps = 44/547 (8%)
Query: 10 QFQRVTWKLEKLLSSLPYDDLDISEEVKEQVDLVRNQLRRA-----TDKYGFMISKMPSF 64
+F + KL ++L P+D+L IS + K+++D + QL++A T + M F
Sbjct: 138 RFHSIYEKLNRVLVKAPFDELMISGDAKDEIDSLCKQLKKAKRRTDTQDIELAVDMMVVF 197
Query: 65 DSSQP------LAQEISQVLG-QSVSGLHKQHSCPENLSE----LGSIPKSNEGKSCNPF 113
+ P + + +++ L Q++ L + ++L + L K + + N F
Sbjct: 198 SKTDPRNADSAIIERLAKKLELQTIDDLKTETIAIQSLIQDKGGLNIETKQHIIELLNKF 257
Query: 114 GAGSRLERTRSIHASSEVSFSIKTAPESQEISGSGNLPEVKKPDAIVIPEDFLCPISLEL 173
LE T ++ Q + + K ++++P +FLCPI+LE+
Sbjct: 258 KKLQGLEATDILY---------------QPVINKA----ITKSTSLILPHEFLCPITLEI 298
Query: 174 MRDPVIVATGQTYERSYIQRWIDCGNTTCPKTQQKLQHLTLTPNYVLRSLVSQWCIEHNI 233
M DPVI+ATGQTYE+ IQ+W D G+ TCPKT+Q+L HL+L PN+ L++L+ QWC ++N
Sbjct: 299 MLDPVIIATGQTYEKESIQKWFDAGHKTCPKTRQELDHLSLAPNFALKNLIMQWCEKNNF 358
Query: 234 EQPTGLTNGKIKKSDGSFRDVTGDIAAIETLVRKLSCRSVEESRAAVAEIRSLSKRSTDN 293
KI + + S + LV LS +EE R +V ++R L++ + +N
Sbjct: 359 ---------KIPEKEVSPDSQNEQKDEVSLLVEALSSSQLEEQRRSVKQMRLLARENPEN 409
Query: 294 RILIAEAGAIPVLVSLLTSEDVMTQENAVTSILNLSIYENNKGLIMLAGAIPSIVQVLRA 353
R+LIA AGAIP+LV LL+ D QENAVT++LNLSI E NK LI GAIP+I+++L
Sbjct: 410 RVLIANAGAIPLLVQLLSYPDSGIQENAVTTLLNLSIDEVNKKLISNEGAIPNIIEILEN 469
Query: 354 GTMEARENAAATLFSLSLADENKIIIGASGAISALVDLLQNGSPRGKKDAATALFNLCIY 413
G EAREN+AA LFSLS+ DENK+ IG S I LVDLLQ+G+ RGKKDA TALFNL +
Sbjct: 470 GNREARENSAAALFSLSMLDENKVTIGLSNGIPPLVDLLQHGTLRGKKDALTALFNLSLN 529
Query: 414 QGNKGRAIRAGIITALLNMLTDSSKSMVDEALTIMSVLASHQEAKVSIVKASTIPVLIDL 473
NKGRAI AGI+ LLN+L D + M+DEAL+I+ +LASH E + +I + S I L++
Sbjct: 530 SANKGRAIDAGIVQPLLNLLKDKNLGMIDEALSILLLLASHPEGRQAIGQLSFIETLVEF 589
Query: 474 LRTGLPRNKENAAAILLALCKRDTDNLSCISRLGAVIPLSELARTGTERAKRKATSLLEH 533
+R G P+NKE A ++LL L ++ + + G L E+ +GT RA+RKA +L++
Sbjct: 590 IRQGTPKNKECATSVLLELGSNNSSFILAALQFGVYEYLVEITTSGTNRAQRKANALIQL 649
Query: 534 LRKLQQL 540
+ K +Q+
Sbjct: 650 ISKSEQI 656
>At1g29340 arm repeat-containing protein, putative
Length = 729
Score = 279 bits (714), Expect = 2e-75
Identities = 201/545 (36%), Positives = 299/545 (53%), Gaps = 31/545 (5%)
Query: 11 FQRVTWKLEKLLSSLPYDDLDISEEVKEQVDLVRNQLRRAT---DKYGFMI-----SKMP 62
F + ++ LL LP +DL +S++++EQ++L++ Q R+A DK + S +
Sbjct: 143 FHDLNQEISTLLDVLPVNDLGLSDDIREQIELLQRQSRKARLYIDKNDESLRESFYSFLD 202
Query: 63 SFDSSQ-PLAQEISQVLGQSVSGLHKQHSCPENLSELGSIPKSNEG------KSCNPFGA 115
F++ + P + ++ + + G+ SC + L +++G N F A
Sbjct: 203 GFENGKIPSSVDLRMFFVEKL-GIRDSKSCRSEIEFLEEQIVNHDGDLEPTGSVINGFVA 261
Query: 116 GSRLERTRSIHASSE-VSFSIKTAPESQEISGSGNLPEVKKPDAIVIPEDFLCPISLELM 174
+R R + + + I+ P+ G + + I +P+DF+CPISL+LM
Sbjct: 262 ITRYCRFLLFGFEEDGMEWWIENNPKKPR---KGFVAQEIGDTFITVPKDFVCPISLDLM 318
Query: 175 RDPVIVATGQTYERSYIQRWIDCGNTTCPKTQQKLQHLTLTPNYVLRSLVSQWCIEHNIE 234
DPVI++TGQTY+R+ I RWI+ G+ TCPKT Q L + PN L++L+ QWC I
Sbjct: 319 TDPVIISTGQTYDRNSIARWIEEGHCTCPKTGQMLMDSRIVPNRALKNLIVQWCTASGIS 378
Query: 235 QPTGLT---NGKIKKSDGSFRDVTGDIAAIETLVRKLSCRSVEESRAAVAEIRSLSKRST 291
+ T N + + V + A + L++ L+ S A EIR L+K
Sbjct: 379 YESEFTDSPNESFASALPTKAAVEANKATVSILIKYLADGSQAAQTVAAREIRLLAKTGK 438
Query: 292 DNRILIAEAGAIPVLVSLLTSEDVMTQENAVTSILNLSIYENNKGLIMLAG-AIPSIVQV 350
+NR IAEAGAIP L LLTSE+ + QEN+VT++LNLSIYE NK IM G + SIV V
Sbjct: 439 ENRAYIAEAGAIPHLCRLLTSENAIAQENSVTAMLNLSIYEKNKSRIMEEGDCLESIVSV 498
Query: 351 LRAG-TMEARENAAATLFSLSLADE-NKIIIGASGAISALVDLLQNGSPRGKKDAATALF 408
L +G T+EA+ENAAATLFSLS E K I + AL LLQNG+PRGKKDA TAL+
Sbjct: 499 LVSGLTVEAQENAAATLFSLSAVHEYKKRIAIVDQCVEALALLLQNGTPRGKKDAVTALY 558
Query: 409 NLCIYQGNKGRAIRAGIITALLNMLTDSSKSMVDEALTIMSVLASHQEAKVSIVKA-STI 467
NL + N R I G +++L+ L ++ + +EA +++L +I K S +
Sbjct: 559 NLSTHPDNCSRMIEGGGVSSLVGAL--KNEGVAEEAAGALALLVRQSLGAEAIGKEDSAV 616
Query: 468 PVLIDLLRTGLPRNKENAAAILLALCKRDTDNLS-CISRLGAVIPLSE-LARTGTERAKR 525
L+ ++R G PR KENA A LL LC+ ++ + R A+ L + L TGT+RA+R
Sbjct: 617 AGLMGMMRCGTPRGKENAVAALLELCRSGGAAVAEKVLRAPAIAGLLQTLLFTGTKRARR 676
Query: 526 KATSL 530
KA SL
Sbjct: 677 KAASL 681
>At3g07360 unknown protein
Length = 460
Score = 255 bits (652), Expect = 4e-68
Identities = 147/392 (37%), Positives = 232/392 (58%), Gaps = 13/392 (3%)
Query: 157 DAIVIPEDFLCPISLELMRDPVIVATGQTYERSYIQRWIDCGNTTCPKTQQKLQHLTLTP 216
+ + PE+F CP+S ELMRDPV++A+GQTY++ +IQ+W+ GN TCPKTQQ L H LTP
Sbjct: 70 ETVSCPEEFRCPLSNELMRDPVVLASGQTYDKLFIQKWLSSGNRTCPKTQQVLPHTALTP 129
Query: 217 NYVLRSLVSQWCIEHNIEQPTGLTNGKIKKSDGSFRDVTGDIAAIETLVRKLSCRSVEES 276
N ++R ++S+WC ++ +E + + + + R D +L+ K+S ++++
Sbjct: 130 NLLIREMISKWCKKNGLETKSQYHPNLVNEDETVTR---SDREIFNSLLCKVSSSNLQDQ 186
Query: 277 RAAVAEIRSLSKRSTDNRILIAEA-GAIPVLVSLL---TSEDVMTQENAVTSILNLSIYE 332
++A E+R L+++ T+ R L E+ I LV+ L ++ D QE+ VT++LN+SI++
Sbjct: 187 KSAAKELRLLTRKGTEFRALFGESPDEITRLVNPLLHGSNPDEKLQEDVVTTLLNISIHD 246
Query: 333 --NNKGLIMLAGAIPSIVQVLRAGTMEARENAAATLFSLSLADENKIIIGASGAISALVD 390
N K + IP ++ LR GT+ R NAAA +F+LS D NK++IG SG + L+D
Sbjct: 247 DSNKKLVCENPNVIPLLIDALRRGTVATRSNAAAAIFTLSALDSNKVLIGKSGILKPLID 306
Query: 391 LLQNGSPRGKKDAATALFNLCIYQGNKGRAIRAGIITALLNMLTDSSKSMVDEALTIMSV 450
LL+ G+P KD A A+F LCI N+ RA+R G + L + S+ VDE L I+++
Sbjct: 307 LLEEGNPLAIKDVAAAIFTLCIAHENRSRAVRDGAVRVLGKKI--SNGLYVDELLAILAM 364
Query: 451 LASHQEAKVSIVKASTIPVLIDLLR-TGLPRNKENAAAILLALCKRDTDNLSCI-SRLGA 508
L +H +A + + + L+ + R + RNKENA IL +C D I A
Sbjct: 365 LVTHWKAVEELGELGGVSWLLKITRESECKRNKENAIVILHTICFSDRTKWKEIKEEENA 424
Query: 509 VIPLSELARTGTERAKRKATSLLEHLRKLQQL 540
+++L+R GT RA+RKA +L+ LRK L
Sbjct: 425 HGTITKLSREGTSRAQRKANGILDRLRKAMNL 456
>At5g67340 unknown proteins
Length = 707
Score = 251 bits (641), Expect = 7e-67
Identities = 166/465 (35%), Positives = 243/465 (51%), Gaps = 89/465 (19%)
Query: 161 IPEDFLCPISLELMRDPVIVATGQTYERSYIQRWIDCGNTTCPKTQQKLQHLTLTPNYVL 220
+P DF C +SLELM DPVIVA+GQT+ER +IQ+WID G CPKT+Q L H TLTPN+++
Sbjct: 240 VPSDFRCSLSLELMTDPVIVASGQTFERVFIQKWIDMGLMVCPKTRQALSHTTLTPNFIV 299
Query: 221 RSLVSQWCIEHNIEQPTGL---------------------TNGKIKKSDG-SFRDVTGDI 258
R+ ++ WC +N+ P L NG + D R V
Sbjct: 300 RAFLASWCETNNVYPPDPLELIHSSEPFPLLVESVRASSSENGHSESLDAEELRQVFSRS 359
Query: 259 AAIETLVRKLSCRSVEESRAAVAEIRSLSKRSTDNRI--------------LIAEAGA-- 302
A+ +V ++ C++ + AA RSL++ +T + + E G+
Sbjct: 360 ASAPGIVSEVVCKTKRNNNAAAD--RSLTRSNTPWKFPEERHWRHPGIIPATVRETGSSS 417
Query: 303 -----IPVLVSLLTSEDVMTQENA------------------------------------ 321
+ L+ L S + TQ A
Sbjct: 418 SIETEVKKLIDDLKSSSLDTQREATARIRILARNSTDNRIVIARCEAIPSLVSLLYSTDE 477
Query: 322 ------VTSILNLSIYENNKGLIMLAGAIPSIVQVLRAGTM-EARENAAATLFSLSLADE 374
VT +LNLSI +NNK LI +GAI ++ VL+ G + EA+ N+AATLFSLS+ +E
Sbjct: 478 RIQADAVTCLLNLSINDNNKSLIAESGAIVPLIHVLKTGYLEEAKANSAATLFSLSVIEE 537
Query: 375 NKIIIGASGAISALVDLLQNGSPRGKKDAATALFNLCIYQGNKGRAIRAGIITALLNMLT 434
K IG +GAI LVDLL +GS GKKDAATALFNL I+ NK + I AG + L+ ++
Sbjct: 538 YKTEIGEAGAIEPLVDLLGSGSLSGKKDAATALFNLSIHHENKTKVIEAGAVRYLVELM- 596
Query: 435 DSSKSMVDEALTIMSVLASHQEAKVSIVKASTIPVLIDLLRTGLPRNKENAAAILLALCK 494
D + MV++A+ +++ LA+ +E K++I + IPVL++++ G R KENA A LL LC
Sbjct: 597 DPAFGMVEKAVVVLANLATVREGKIAIGEEGGIPVLVEVVELGSARGKENATAALLQLCT 656
Query: 495 RDTDNLSCISRLGAVIPLSELARTGTERAKRKATSLLEHLRKLQQ 539
+ + R G + PL L ++GT R K KA +LL++ + +Q
Sbjct: 657 HSPKFCNNVIREGVIPPLVALTKSGTARGKEKAQNLLKYFKAHRQ 701
>At2g23140 hypothetical protein
Length = 924
Score = 236 bits (602), Expect = 2e-62
Identities = 134/280 (47%), Positives = 191/280 (67%), Gaps = 1/280 (0%)
Query: 247 SDGSFRDVTGDIAAIETLVRKLSCRSVEESRAAVAEIRSLSKRSTDNRILIAEAGAIPVL 306
S+ + RD++ ++ LV +L S++ R A AE+R L+K + DNRI+I +GAI +L
Sbjct: 608 SNETRRDLSEVETQVKKLVEELKSSSLDTQRQATAELRLLAKHNMDNRIVIGNSGAIVLL 667
Query: 307 VSLLTSEDVMTQENAVTSILNLSIYENNKGLIMLAGAIPSIVQVLRAGTMEARENAAATL 366
V LL S D TQENAVT++LNLSI +NNK I AGAI ++ VL G+ EA+EN+AATL
Sbjct: 668 VELLYSTDSATQENAVTALLNLSINDNNKKAIADAGAIEPLIHVLENGSSEAKENSAATL 727
Query: 367 FSLSLADENKIIIGASGAISALVDLLQNGSPRGKKDAATALFNLCIYQGNKGRAIRAGII 426
FSLS+ +ENKI IG SGAI LVDLL NG+PRGKKDAATALFNL I+Q NK +++G +
Sbjct: 728 FSLSVIEENKIKIGQSGAIGPLVDLLGNGTPRGKKDAATALFNLSIHQENKAMIVQSGAV 787
Query: 427 TALLNMLTDSSKSMVDEALTIMSVLASHQEAKVSIVKASTIPVLIDLLRTGLPRNKENAA 486
L++++ D + MVD+A+ +++ LA+ E + +I + IP+L++++ G R KENAA
Sbjct: 788 RYLIDLM-DPAAGMVDKAVAVLANLATIPEGRNAIGQEGGIPLLVEVVELGSARGKENAA 846
Query: 487 AILLALCKRDTDNLSCISRLGAVIPLSELARTGTERAKRK 526
A LL L + + + GAV PL L+++GT RA+ K
Sbjct: 847 AALLQLSTNSGRFCNMVLQEGAVPPLVALSQSGTPRAREK 886
Score = 118 bits (295), Expect = 9e-27
Identities = 118/442 (26%), Positives = 195/442 (43%), Gaps = 72/442 (16%)
Query: 158 AIVIPEDFLCPISLELMRDPVIVATGQTYERSYIQRWIDCGNTTCPKTQQKLQHLTLTPN 217
++ I DF CP+SLE+M DPVIV++GQTYE+++I+RWID G CPKT+Q L H TL PN
Sbjct: 306 SVAILADFFCPLSLEVMTDPVIVSSGQTYEKAFIKRWIDLGLKVCPKTRQTLTHTTLIPN 365
Query: 218 YVLRSLVSQWCIEHNIEQPTGLTNGKIKK---------------SDGSFRDVTG-----D 257
Y +++L++ WC ++++ P + + + +D S R V+ D
Sbjct: 366 YTVKALIANWCETNDVKLPDPNKSTSLNELSPLLSCTDSIPSTGADVSARKVSNKSHDWD 425
Query: 258 IAAIETLVRKLSCRSVEE-----SRAAVA--------------------EIRSLSKRSTD 292
++ ET S R+ E SR A A + RS D
Sbjct: 426 ASSSETGKPSFSSRATEREGASPSRPASALGASSPGISGNGYGLDARRGSLNDFEDRSND 485
Query: 293 NRILIAEAGAIPVLVSLLTSEDVMTQENAVTSILNLSIYENNKGLIMLAGAIPSIVQVLR 352
+R L +A S ++S + EN TS EN+ A + S + R
Sbjct: 486 SRELRTDAPG----RSSVSSTTRGSVENGQTS-------ENHHHRSPSATSTVSNEEFPR 534
Query: 353 AGTME-ARENAAATLFSLSLADENKIIIGASGAISALVDLLQNGSPRGKKDAATALF--- 408
A E + E+A AT +S + E + A+ +A L + SP+ F
Sbjct: 535 ADANENSEESAHATPYSSDASGEIRSGPLAATTSAATRRDLSDFSPKFMDRRTRGQFWRR 594
Query: 409 -------NLCIYQGNKGRAIRAGIITALLNMLTDSSKSMVD---EALTIMSVLASH-QEA 457
+ N+ R + + T + ++ + S +D +A + +LA H +
Sbjct: 595 PSERLGSRIVSAPSNETRRDLSEVETQVKKLVEELKSSSLDTQRQATAELRLLAKHNMDN 654
Query: 458 KVSIVKASTIPVLIDLLRTGLPRNKENAAAILLALCKRDTDNLSCISRLGAVIPLSELAR 517
++ I + I +L++LL + +ENA LL L D +N I+ GA+ PL +
Sbjct: 655 RIVIGNSGAIVLLVELLYSTDSATQENAVTALLNLSIND-NNKKAIADAGAIEPLIHVLE 713
Query: 518 TGTERAKRKATSLLEHLRKLQQ 539
G+ AK + + L L +++
Sbjct: 714 NGSSEAKENSAATLFSLSVIEE 735
Score = 69.3 bits (168), Expect = 5e-12
Identities = 61/211 (28%), Positives = 94/211 (43%), Gaps = 45/211 (21%)
Query: 237 TGLTNGKIKKSDGSFRDVTGDIAAIETLVRKLSCRSVEESRAAVAEIRSLSKRSTDNRIL 296
T L N I ++ + D AIE L+ L S E + A + SLS +N+I
Sbjct: 684 TALLNLSINDNN---KKAIADAGAIEPLIHVLENGSSEAKENSAATLFSLSVIE-ENKIK 739
Query: 297 IAEAGAIPVLVSLLTSEDVMTQENAVTSILNLSIYENNKGLIMLAGA------------- 343
I ++GAI LV LL + +++A T++ NLSI++ NK +I+ +GA
Sbjct: 740 IGQSGAIGPLVDLLGNGTPRGKKDAATALFNLSIHQENKAMIVQSGAVRYLIDLMDPAAG 799
Query: 344 ---------------------------IPSIVQVLRAGTMEARENAAATLFSLSLADENK 376
IP +V+V+ G+ +ENAAA L LS
Sbjct: 800 MVDKAVAVLANLATIPEGRNAIGQEGGIPLLVEVVELGSARGKENAAAALLQLSTNSGRF 859
Query: 377 I-IIGASGAISALVDLLQNGSPRGKKDAATA 406
++ GA+ LV L Q+G+PR ++ TA
Sbjct: 860 CNMVLQEGAVPPLVALSQSGTPRAREKKPTA 890
Score = 60.8 bits (146), Expect = 2e-09
Identities = 59/217 (27%), Positives = 98/217 (44%), Gaps = 43/217 (19%)
Query: 319 ENAVTSILNLSIYENNKGLIMLAGAIPSIVQVLRAGTMEARENAAATLFSLSLAD-ENKI 377
E + I++ E + L + + +V+ L++ +++ + A A L L+ + +N+I
Sbjct: 597 ERLGSRIVSAPSNETRRDLSEVETQVKKLVEELKSSSLDTQRQATAELRLLAKHNMDNRI 656
Query: 378 IIGASGAISALVDLLQNGSPRGKKDAATALFNLCIYQGNKGRAIRAGIITALLNMLTDSS 437
+IG SGAI LV+LL + D+AT + +TALLN
Sbjct: 657 VIGNSGAIVLLVELLYS------TDSAT----------------QENAVTALLN------ 688
Query: 438 KSMVDEALTIMSVLASHQEAKVSIVKASTIPVLIDLLRTGLPRNKENAAAILLALCKRDT 497
L+ + K +I A I LI +L G KEN+AA L +L +
Sbjct: 689 -------------LSINDNNKKAIADAGAIEPLIHVLENGSSEAKENSAATLFSLSVIEE 735
Query: 498 DNLSCISRLGAVIPLSELARTGTERAKRKATSLLEHL 534
+ + I + GA+ PL +L GT R K+ A + L +L
Sbjct: 736 NKIK-IGQSGAIGPLVDLLGNGTPRGKKDAATALFNL 771
>At5g01830 unknown protein
Length = 674
Score = 234 bits (597), Expect = 9e-62
Identities = 182/555 (32%), Positives = 272/555 (48%), Gaps = 59/555 (10%)
Query: 7 IIFQFQRVTWKLEKLLSSLPYDDLDISEEVKEQVDLVRNQLRRATDKYGFMISKMPSFDS 66
+ F F + L +L LP D D+S++ ++ + L+ Q S F
Sbjct: 125 VAFNFHELVTDLSTVLDILPLHDFDLSDDAQDLISLLTKQC-----------SDSVQFVD 173
Query: 67 SQPLAQEISQVLGQSVSGLHKQHSCPENLSELGSIPKSNEGKSCNPFGAGSRLERTRSIH 126
++ +A + + + +++G+ Q S P++ + + K N G T I
Sbjct: 174 ARDVA--LRRKVTDTIAGIKHQIS-PDHSTLI---------KIFNDLGLSDSASLTDEIQ 221
Query: 127 A-SSEVSFSIKTAPESQEISGSGNL---------PEVKKPDA--------IVIPEDFLCP 168
E+ I +S S G + P PD IP DF CP
Sbjct: 222 RLEDEIQDQIDDRSKSAAASLIGLVRYSKCVLYGPSTPAPDFRRHQSLSDANIPADFRCP 281
Query: 169 ISLELMRDPVIVATGQTYERSYIQRWIDCGNTTCPKTQQKLQHLTLTPNYVLRSLVSQWC 228
I+LELMRDPV+VATGQTY+R I WI G+ TCPKT Q L+H +L PN L++L+ WC
Sbjct: 282 ITLELMRDPVVVATGQTYDRESIDLWIQSGHNTCPKTGQVLKHTSLVPNRALKNLIVLWC 341
Query: 229 IEHNIEQPTGLTNGKIKKSDGSFRDVTGDIAAIETLVRKLSCRSVEESRAAVAEIRSLSK 288
+ I P L + V + L+ KL SV +S V E+R+L+K
Sbjct: 342 RDQKI--PFELYGDGGGEPAPCKEAVEFTKMMVSFLIEKL---SVADSNGVVFELRALAK 396
Query: 289 RSTDNRILIAEAGAIPVLVSLLTSEDVMTQENAVTSILNLSIYENNKGLIM-LAGAIPSI 347
T R IAEAGAIP LV L +E Q NAVT+ILNLSI E NK IM GA+ +
Sbjct: 397 SDTVARACIAEAGAIPKLVRYLATECPSLQINAVTTILNLSILEQNKTRIMETDGALNGV 456
Query: 348 VQVLRAG-TMEARENAAATLFSLSLADENKIIIGASG-AISALVDLLQNGSPRGKKDAAT 405
++VLR+G T EA+ NAAATLFSL+ + +G +S LVDL + G K+DA
Sbjct: 457 IEVLRSGATWEAKANAAATLFSLAGVSAYRRRLGRKARVVSGLVDLAKQGPTSSKRDALV 516
Query: 406 ALFNLCIYQGNKGRAIRAGIITALLNMLTDSSKSMVDEALTIMSVLASHQEAKVSIVKAS 465
A+ NL + N GR + AG++ A D+ + + +EA+ ++ + S
Sbjct: 517 AILNLVAERENVGRFVEAGVMGA----AGDAFQELPEEAVAVVEAVVRRGGLMAVSAAFS 572
Query: 466 TIPVLIDLLRTGLPRNKENAAAILLALCKRDTDNLSCISRLGAVIPLS----ELARTGTE 521
I +L +++R G +E+AAA L+ +C++ L ++ + A+ + E+ GT
Sbjct: 573 LIRLLGEVMREGADTTRESAAATLVTMCRKGGSEL--VAEMAAIPGIERVIWEMIGAGTA 630
Query: 522 RAKRKATSLLEHLRK 536
R RKA SL+ +LR+
Sbjct: 631 RGGRKAASLMRYLRR 645
>At3g01400 unknown protein
Length = 355
Score = 228 bits (582), Expect = 5e-60
Identities = 124/268 (46%), Positives = 181/268 (67%)
Query: 269 SCRSVEESRAAVAEIRSLSKRSTDNRILIAEAGAIPVLVSLLTSEDVMTQENAVTSILNL 328
S S++E + A EIR LSK +NRI IA+AGAI L+SL++S D+ QE VT+ILNL
Sbjct: 73 SSYSIDEQKQAAMEIRLLSKNKPENRIKIAKAGAIKPLISLISSSDLQLQEYGVTAILNL 132
Query: 329 SIYENNKGLIMLAGAIPSIVQVLRAGTMEARENAAATLFSLSLADENKIIIGASGAISAL 388
S+ + NK I +GAI +V+ L+ GT A+ENAA L LS +ENK+ IG SGAI L
Sbjct: 133 SLCDENKESIASSGAIKPLVRALKMGTPTAKENAACALLRLSQIEENKVAIGRSGAIPLL 192
Query: 389 VDLLQNGSPRGKKDAATALFNLCIYQGNKGRAIRAGIITALLNMLTDSSKSMVDEALTIM 448
V+LL+ G R KKDA+TAL++LC + NK RA+++GI+ L+ ++ D +MVD++ +M
Sbjct: 193 VNLLETGGFRAKKDASTALYSLCSAKENKIRAVQSGIMKPLVELMADFGSNMVDKSAFVM 252
Query: 449 SVLASHQEAKVSIVKASTIPVLIDLLRTGLPRNKENAAAILLALCKRDTDNLSCISRLGA 508
S+L S E+K +IV+ +PVL++++ G R KE A +ILL LC+ + ++R GA
Sbjct: 253 SLLMSVPESKPAIVEEGGVPVLVEIVEVGTQRQKEMAVSILLQLCEESVVYRTMVAREGA 312
Query: 509 VIPLSELARTGTERAKRKATSLLEHLRK 536
+ PL L++ GT RAK+KA +L+E LR+
Sbjct: 313 IPPLVALSQAGTSRAKQKAEALIELLRQ 340
>At5g58680 unknown protein
Length = 357
Score = 221 bits (562), Expect = 1e-57
Identities = 125/281 (44%), Positives = 182/281 (64%), Gaps = 2/281 (0%)
Query: 261 IETLVRKL-SCRSVEESRAAVAEIRSLSKRSTDNRILIAEAGAIPVLVSLLTSEDVMTQE 319
I L+ L S S+EE + A EIR LSK +NRI +A+AGAI LVSL++S D+ QE
Sbjct: 62 IRNLITHLESSSSIEEQKQAAMEIRLLSKNKPENRIKLAKAGAIKPLVSLISSSDLQLQE 121
Query: 320 NAVTSILNLSIYENNKGLIMLAGAIPSIVQVLRAGTMEARENAAATLFSLSLADENKIII 379
VT++LNLS+ + NK +I+ +GA+ +V LR GT +ENAA L LS +ENKI I
Sbjct: 122 YGVTAVLNLSLCDENKEMIVSSGAVKPLVNALRLGTPTTKENAACALLRLSQVEENKITI 181
Query: 380 GASGAISALVDLLQNGSPRGKKDAATALFNLCIYQGNKGRAIRAGIITALLNMLTDSSKS 439
G SGAI LV+LL+NG R KKDA+TAL++LC NK RA+ +GI+ L+ ++ D
Sbjct: 182 GRSGAIPLLVNLLENGGFRAKKDASTALYSLCSTNENKTRAVESGIMKPLVELMIDFESD 241
Query: 440 MVDEALTIMSVLASHQEAKVSIVKASTIPVLIDLLRTGLPRNKENAAAILLALCKRDTDN 499
MVD++ +M++L S E+K ++V+ +PVL++++ G R KE + +ILL LC+
Sbjct: 242 MVDKSAFVMNLLMSAPESKPAVVEEGGVPVLVEIVEAGTQRQKEISVSILLQLCEESVVY 301
Query: 500 LSCISRLGAVIPLSELARTGTER-AKRKATSLLEHLRKLQQ 539
+ ++R GAV PL L++ R AK KA +L+E LR+ +Q
Sbjct: 302 RTMVAREGAVPPLVALSQGSASRGAKVKAEALIELLRQPRQ 342
>At3g54790 unknown protein
Length = 760
Score = 210 bits (534), Expect = 2e-54
Identities = 126/273 (46%), Positives = 176/273 (64%), Gaps = 2/273 (0%)
Query: 264 LVRKLSCRSVEESRAAVAEIRSLSKRSTDNRILIAEAGAIPVLVSLLTSEDVMTQENAVT 323
LV L S + AA AEIR L+ S +NR+ I GAI L+SLL SE+ +TQE+AVT
Sbjct: 477 LVEDLKSGSNKVKTAAAAEIRHLTINSIENRVHIGRCGAITPLLSLLYSEEKLTQEHAVT 536
Query: 324 SILNLSIYENNKGLIMLAGAIPSIVQVLRAGTMEARENAAATLFSLSLADENKIIIGAS- 382
++LNLSI E NK +I+ GAI +V VL G A+EN+AA+LFSLS+ N+ IG S
Sbjct: 537 ALLNLSISELNKAMIVEVGAIEPLVHVLNTGNDRAKENSAASLFSLSVLQVNRERIGQSN 596
Query: 383 GAISALVDLLQNGSPRGKKDAATALFNLCIYQGNKGRAIRAGIITALLNMLTDSSKSMVD 442
AI ALV+LL G+ RGKKDAA+ALFNL I NK R ++A + L+ +L D MVD
Sbjct: 597 AAIQALVNLLGKGTFRGKKDAASALFNLSITHDNKARIVQAKAVKYLVELL-DPDLEMVD 655
Query: 443 EALTIMSVLASHQEAKVSIVKASTIPVLIDLLRTGLPRNKENAAAILLALCKRDTDNLSC 502
+A+ +++ L++ E + +IV+ IP+L++ + G R KENAA++LL LC +
Sbjct: 656 KAVALLANLSAVGEGRQAIVREGGIPLLVETVDLGSQRGKENAASVLLQLCLNSPKFCTL 715
Query: 503 ISRLGAVIPLSELARTGTERAKRKATSLLEHLR 535
+ + GA+ PL L+++GT+RAK KA LL H R
Sbjct: 716 VLQEGAIPPLVALSQSGTQRAKEKAQQLLSHFR 748
Score = 124 bits (312), Expect = 1e-28
Identities = 119/433 (27%), Positives = 200/433 (45%), Gaps = 61/433 (14%)
Query: 159 IVIPEDFLCPISLELMRDPVIVATGQTYERSYIQRWIDCGNTTCPKTQQKLQHLTLTPNY 218
I IP F CP+S ELM DPVIVA+GQT++R+ I++W+D G CP+T+Q L H L PNY
Sbjct: 236 ISIPPYFRCPLSTELMLDPVIVASGQTFDRTSIKKWLDNGLAVCPRTRQVLTHQELIPNY 295
Query: 219 VLRSLVSQW------------CIEHNIEQPTGLTNG--------------KIKKSDGSFR 252
++++++ W C +++ + + N ++ S + R
Sbjct: 296 TVKAMIASWLEANRINLATNSCHQYDGGDASSMANNMGSQDFNRTESFRFSLRSSSLTSR 355
Query: 253 DVTGDIAAIETLVRKLSCRSVEESRAAVAEIRSL--------SKRSTDNRILIAEAGAIP 304
E L +S ES++ EI L RS +++ +P
Sbjct: 356 SSLETGNGFEKLKINVSASLCGESQSKDLEIFELLSPGQSYTHSRSESVCSVVSSVDYVP 415
Query: 305 VLV----SLL---------------TSEDVMTQENAVTSILNLSIYENNKGLIMLAGAIP 345
+ S+L S + + E++ S+++ + M
Sbjct: 416 SVTHETESILGNHQSSSEMSPKKNLESSNNVNHEHSAAKTYECSVHDLDDSGTMTTSHTI 475
Query: 346 SIVQVLRAGTMEARENAAATLFSLSL-ADENKIIIGASGAISALVDLLQNGSPRGKKDAA 404
+V+ L++G+ + + AAA + L++ + EN++ IG GAI+ L+ LL + ++ A
Sbjct: 476 KLVEDLKSGSNKVKTAAAAEIRHLTINSIENRVHIGRCGAITPLLSLLYSEEKLTQEHAV 535
Query: 405 TALFNLCIYQGNKGRAIRAGIITALLNML---TDSSKSMVDEALTIMSVLASHQEAKVSI 461
TAL NL I + NK + G I L+++L D +K +L +SVL ++E ++
Sbjct: 536 TALLNLSISELNKAMIVEVGAIEPLVHVLNTGNDRAKENSAASLFSLSVLQVNRE-RIGQ 594
Query: 462 VKASTIPVLIDLLRTGLPRNKENAAAILLALCKRDTDNLSCISRLGAVIPLSELARTGTE 521
A+ I L++LL G R K++AA+ L L DN + I + AV L EL E
Sbjct: 595 SNAA-IQALVNLLGKGTFRGKKDAASALFNL-SITHDNKARIVQAKAVKYLVELLDPDLE 652
Query: 522 RAKRKATSLLEHL 534
KA +LL +L
Sbjct: 653 MVD-KAVALLANL 664
>At5g18320 putative protein
Length = 474
Score = 203 bits (516), Expect = 2e-52
Identities = 147/424 (34%), Positives = 227/424 (52%), Gaps = 39/424 (9%)
Query: 135 IKTAPESQEISGSGNLPEVKKPDA----IVIPEDFLCPISLELMRDPVIVATGQTYERSY 190
+K E+ I E K P++ + +P++F+C +S +M +PVI+A+GQTYE+ Y
Sbjct: 42 VKAIDEAVRILTCLRKVESKIPESDISPVEVPKEFICTLSNTIMIEPVIIASGQTYEKRY 101
Query: 191 IQRWIDCGNTTCPKTQQKLQHLTLTPNYVLRSLVSQWCIEHNIEQPTGLTNGKIKKSDGS 250
I W+ TCPKT+Q L H PN+++ L++QWC+ + + K SD
Sbjct: 102 ITEWLK-HERTCPKTKQVLSHRLWIPNHLISDLITQWCLVNKYDHQ--------KPSDEL 152
Query: 251 FRDV-TGDIAAIETLVRKLSCRSVEESRAAVAEIRSLSKRSTDNRILIAEA--GAIPVLV 307
++ T DI A+ L R S SV + A E+R +K+ + R+ +I L+
Sbjct: 153 VAELFTSDIEAL--LQRVSSSSSVADQIEAAKELRHQTKKFPNVRVFFVAGIHDSITRLL 210
Query: 308 SLLTSED------VMTQENAVTSILNLSIYENNKGLIML-AGAIPSIVQVLRAGTMEARE 360
S L++ D + QEN VT++ NLSI E+NK +I IP + + L+ GT E R
Sbjct: 211 SPLSTLDEAVDSSLELQENIVTALFNLSILESNKTVIAENCLVIPLLTKSLKQGTDETRR 270
Query: 361 NAAATLFSLSLADENKIIIGASGAISALVDLLQNGSPRGKKDAATALFNLCIYQGNKGRA 420
NAAATL SLS D NKIIIG S A+ AL+DL++ G K+A + +FNLCI NKG+
Sbjct: 271 NAAATLSSLSAIDSNKIIIGNSEAVKALIDLIEEGDLLATKEATSTVFNLCIVLENKGKV 330
Query: 421 IRAGIITALLNMLTDSSKSMVDEALTIMSVLASHQEAKVSIVKASTIPVLIDLLR-TGLP 479
+ AG+I A + + S VDE L++++++++H A + K I L +LR
Sbjct: 331 VSAGLIHAATKKI--KAGSNVDELLSLLALISTHNRAVEEMDKLGFIYDLFSILRKPSSL 388
Query: 480 RNKENAAAILLALCKRDTDNLSCISRLGAV-------IPLSELARTGTERAKRKATSLLE 532
ENA I+ + R+ D SRL V ++LA+ G+ RA RKA +L+
Sbjct: 389 LTGENAVVIVFNMYDRNRDR----SRLKVVGEEENQHGTFTKLAKQGSVRAARKAQGILQ 444
Query: 533 HLRK 536
+++
Sbjct: 445 WIKR 448
>At4g21350 putative protein
Length = 374
Score = 175 bits (443), Expect = 7e-44
Identities = 124/380 (32%), Positives = 202/380 (52%), Gaps = 17/380 (4%)
Query: 158 AIVIPEDFLCPISLELMRDPVIVATGQTYERSYIQRWIDCGNTTCPKTQQKLQHLT-LTP 216
A +P DF CPISLE+M DPVI+ +G T++R IQ+WID GN TCP T+ L L P
Sbjct: 2 AFDLPNDFRCPISLEIMSDPVILQSGHTFDRVSIQQWIDSGNRTCPITKLPLSETPYLIP 61
Query: 217 NYVLRSLVSQWCIEHNIEQPTGLTNGKIKKSDGSFRDVTGDIAAIETLVRKLSCRSVEES 276
N+ LRSL+ + E T + S A I TLV + S S
Sbjct: 62 NHALRSLILNFAHVSLKESSRPRTQQEHSHSQSQ--------ALISTLVSQSS--SNASK 111
Query: 277 RAAVAEIRSLSKRSTDNRILIAEAGAIPVLVSLLTSEDVMTQENAVTSILNLSIYENNKG 336
++ + L+KR + R + E+GA+ + + S + + QE +++ +LNLS+ ++NK
Sbjct: 112 LESLTRLVRLTKRDSSIRRKVTESGAVRAALDCVDSCNQVLQEKSLSLLLNLSLEDDNKV 171
Query: 337 LIMLAGAIPSIVQVLRAGTMEARENAAATLFSLSLADENKIIIGA-SGAISALVDLLQNG 395
++ G I IV VLR G+ + + AA L SL++ + NK IG+ AISALV LL+ G
Sbjct: 172 GLVADGVIRRIVTVLRVGSPDCKAIAATLLTSLAVVEVNKATIGSYPDAISALVSLLRVG 231
Query: 396 SPRGKKDAATALFNLCIYQGNKGRAIRAGIITALLNMLTDSSKSMVDEALTIMSVLASHQ 455
+ R +K++ATAL+ LC + N+ R + G + +L +++ S ++ A+ ++ +L +
Sbjct: 232 NDRERKESATALYALCSFPDNRKRVVDCGSVP----ILVEAADSGLERAVEVLGLLVKCR 287
Query: 456 EAKVSIVKAS-TIPVLIDLLRTGLPRNKENAAAILLALCKRDTDNLSCISRLGAVIPLSE 514
+ + K S + VL+++LR G + + + IL LC + + + R G V
Sbjct: 288 GGREEMSKVSGFVEVLVNVLRNGNLKGIQYSLFILNCLCCCSGEIVDEVKREGVVEICFG 347
Query: 515 LARTGTERAKRKATSLLEHL 534
+E+ +R AT L+ L
Sbjct: 348 FEDNESEKIRRNATILVHTL 367
>At3g19380 unknown protein
Length = 421
Score = 164 bits (415), Expect = 1e-40
Identities = 127/391 (32%), Positives = 194/391 (49%), Gaps = 19/391 (4%)
Query: 159 IVIPEDFLCPISLELMRDPVIVATGQTYERSYIQRWIDCG-NTTCPKTQQKLQHLTLTPN 217
I IP F CPISLELM+DPV V TGQTY+R+ I+ W+ G NTTCP T+ L TL PN
Sbjct: 12 IQIPYHFRCPISLELMQDPVTVCTGQTYDRASIESWVSIGNNTTCPVTRAPLSDFTLIPN 71
Query: 218 YVLRSLVSQWCIEHNIEQPTGLTNGKIKKSDGSFRDVTGDIAAIETLVRKLSCRSVEESR 277
+ LR L+ +WC+ + + K S R + +AI + SV
Sbjct: 72 HTLRRLIQEWCVANRSNGVERIPTPKQPADPTSVRALLSQASAITG-----THVSVRSRA 126
Query: 278 AAVAEIRSLSKRSTDNRILIAEAGAIPVLVSLLTSEDVMTQ--ENAVTSILNLSIYENNK 335
AA+ +R ++ S NR+LIA A +L+ +L SE ++ ++ ++ L I E N+
Sbjct: 127 AALRRLRGFARDSDKNRVLIAAHNATEILIKILFSETTSSELVSESLALLVMLPITEPNQ 186
Query: 336 GLIMLA--GAIPSIVQVLRAGTMEARENAAATLFSLSL----ADENKIIIGASGAISALV 389
+ + + G + + ++L ++E R NAAA + +S AD I + ++
Sbjct: 187 FVSISSDPGRVEFLTRLLFDSSIETRVNAAALIEIVSTGTKSADLKGSISNSESVFEGVL 246
Query: 390 DLLQN--GSPRGKKDAATALFNLCIYQGNKGRAIRAGIITALLNML-TDSSKSMVDEALT 446
DLL+N S R K LF LC + + AI AG L++ L D + + AL
Sbjct: 247 DLLRNPISSRRALKIGIKTLFALCSVKSTRHIAITAGAPEILIDRLAADFDRCDTERALA 306
Query: 447 IMSVLASHQEAKVSIVK-ASTIPVLIDLLRTGLPRNKENAAAILLALCKRDTDNLSCISR 505
+ +L E + + A T+P+L+ + R E AA LLALC + +
Sbjct: 307 TVELLCRTPEGCAAFGEHALTVPLLVKTILRVSDRATEYAAGALLALCTAEERWREEAAG 366
Query: 506 LGAVIPLSELARTG-TERAKRKATSLLEHLR 535
G V+ L + ++ TERAK+KA LL+ LR
Sbjct: 367 AGVVVQLLLMVQSECTERAKKKAQKLLKLLR 397
>At1g49780 unknown protein
Length = 421
Score = 159 bits (402), Expect = 4e-39
Identities = 126/394 (31%), Positives = 192/394 (47%), Gaps = 23/394 (5%)
Query: 159 IVIPEDFLCPISLELMRDPVIVATGQTYERSYIQRWIDCGNTTCPKTQQKLQHLTLTPNY 218
I IP F CPISL+LM DPV ++TGQTY+R+ I WI GNTTCP T+ L TL PN+
Sbjct: 12 IQIPYHFRCPISLDLMSDPVTISTGQTYDRTSIDSWIAMGNTTCPVTRVALSDFTLIPNH 71
Query: 219 VLRSLVSQWCIEHNIEQPTGLTNGKIKKSDGSFRDVTGDIAAIETLVRKLSCRSVEESRA 278
LR L+ +WC+ + + K S R + +AI + SV A
Sbjct: 72 TLRRLIQEWCVANRSNGVERIPTPKQPADPISVRSLLSQASAITG-----THVSVRSRAA 126
Query: 279 AVAEIRSLSKRSTDNRILIAEAGAIPVLVSLL--------TSEDVMTQENAVTSILNLSI 330
A+ +R L++ S NR+LIA A +LV +L S +++++ A+ +L+++
Sbjct: 127 AIRRLRGLARDSEKNRVLIAGHNAREILVRILFADIETTSLSSELVSESLALLVLLHMTE 186
Query: 331 YENNKGLIMLAGAIPSIVQVLRAGTMEARENAAA----TLFSLSLADENKIIIGASGAIS 386
E + + + + ++L ++E R NAAA L D II G+
Sbjct: 187 TE-CEAVASDPSRVGFMTRLLFDSSIEIRVNAAALIEMVLTGAKSMDLKLIISGSDSIFE 245
Query: 387 ALVDLLQN--GSPRGKKDAATALFNLCIYQGNKGRAIRAGIITALLNML-TDSSKSMVDE 443
++DLL+N S R K A+F LC+ + + AI AG L++ L D + +
Sbjct: 246 GVLDLLKNPISSRRALKIGIKAIFALCLVKQTRHLAISAGAPGILIDRLAADFDRCDTER 305
Query: 444 ALTIMSVLASHQEAKVSIVK-ASTIPVLIDLLRTGLPRNKENAAAILLALCKRDTDNLSC 502
L + +L E + + A T+P+++ + R E AA LLALC +
Sbjct: 306 GLATVELLCRLPEGCAAFGEHALTVPLMVKTILRVSDRATEYAAGALLALCTAEERCRDE 365
Query: 503 ISRLGAVIPLSELARTG-TERAKRKATSLLEHLR 535
+ G V L L ++ TERAKRKA LL+ LR
Sbjct: 366 AAAAGLVTQLLLLVQSDCTERAKRKAQMLLKLLR 399
>At5g18330 putative protein
Length = 445
Score = 157 bits (397), Expect = 1e-38
Identities = 113/358 (31%), Positives = 191/358 (52%), Gaps = 31/358 (8%)
Query: 152 EVKKPDA----IVIPEDFLCPISLELMRDPVIVATGQTYERSYIQRWIDCGNTTCPKTQQ 207
E K P++ + +P++F+C +S ++M +P+++A+GQT+E+SYI W+ TCP+T+Q
Sbjct: 52 ESKNPESDISPVEVPKEFICTLSNKIMIEPMLIASGQTFEKSYILEWLK-HERTCPRTKQ 110
Query: 208 KLQHLTLTPNYVLRSLVSQWCIEHNIEQPTGLTNGKIKKSDGSFRDVTGDIAAIETLVRK 267
L H + PN+++ ++ +WC+ HN ++P K SD TGD+ E+L+++
Sbjct: 111 VLYHRFMIPNHLINEVIKEWCLIHNFDRP--------KTSDEVIDLFTGDL---ESLLQR 159
Query: 268 LSC-RSVEESRAAVAEIRSLSKRSTDNRILIAEAGAIP-----VLVSLLTSEDVMTQ--E 319
+S SVE+ A E+ +KR + + + IP +L L SED + E
Sbjct: 160 ISSPSSVEDQTEAAKELALKAKRFSS--VCVYFVAKIPDSITRLLTPLSISEDSNPEFLE 217
Query: 320 NAVTSILNLSIYENNKGLIMLAGAI-PSIVQVLRAGTMEARENAAATLFSLSLADENKII 378
N VT++ S E NK L+ + P + + ++ GT+ R ++AAT+ SLS D NKII
Sbjct: 218 NIVTALHIFSTSEKNKTLVAENPLVLPLLAKYMKQGTVLTRIHSAATVNSLSYTDSNKII 277
Query: 379 IGASGAISALVDLLQNGSPRGKKDAATALFNLCIYQGNKGRAIRAGIITALLNML-TDSS 437
IG S + AL+ +++ G +A +AL NLC + +A+ G+I A + + S+
Sbjct: 278 IGNSEVLKALIHVIEEGDSLATSEAFSALSNLCPVKEISEKAVSEGLIRAAIKKIKAGSN 337
Query: 438 KSMVDEALTIMSVLASHQEAKVSIVKASTIPVLIDLLRTGLPR-NKENAAAILLALCK 494
SM+ L +S +HQ + + I L +LR N ENA I+ +CK
Sbjct: 338 VSMLLSLLAFVST-QNHQTTE-EMDNLGLIYDLFSILRNSNSLVNDENAVVIVYNICK 393
>At1g60190
Length = 686
Score = 155 bits (391), Expect = 7e-38
Identities = 116/390 (29%), Positives = 209/390 (52%), Gaps = 15/390 (3%)
Query: 163 EDFLCPISLELMRDPVIVATGQTYERSYIQRWIDCGNTTCPKTQQKLQHLTLTPNYVLRS 222
+D CPISLE+M DPV++ +G TY+RS I +W GN TCPKT + L L N+ ++
Sbjct: 280 DDLRCPISLEIMSDPVVLESGHTYDRSSITKWFASGNITCPKTGKTLVSTVLVDNFSVKQ 339
Query: 223 LVSQWCIEHNIEQPTGLTNGKIKKSDGSFRDVTGDIAAIETLVRKLSCRSVEESRAAVAE 282
++ + ++ + K+ ++ + G + A E L +L EE A+ E
Sbjct: 340 VIQSYSKQNGVVMGQ-KGKKKVDVAESLAAEEAGKLTA-EFLAGELIKGDEEEMVKALVE 397
Query: 283 IRSLSKRSTDNRILIAEAGAIPVLVSLLTSEDVMTQENAVTSILNLSIYENNKGLIM--L 340
IR L+K ST R + EAG + L+ +L S+D QENA+ I+NLS K I+
Sbjct: 398 IRILTKTSTFYRSCLVEAGVVESLMKILRSDDPRIQENAMAGIMNLSKDIAGKTRIVGED 457
Query: 341 AGAIPSIVQVLRAGT-MEARENAAATLFSL-SLADENKIIIGASGAISALVDLLQ--NGS 396
G + IV+VL G E+R+ AAA LF L SL D +++I S AI LV +++ +
Sbjct: 458 GGGLRLIVEVLNDGARRESRQYAAAALFYLSSLGDYSRLIGEISDAIPGLVRIVKSCDYG 517
Query: 397 PRGKKDAATALFNLCIYQ-GNKGRAIRAGIITALLNML--TDSSKSMVDEALTIMSVLAS 453
K++A A+ +L + Q N R + AGI+ LL+++ + S + +++ I++ +A
Sbjct: 518 DSAKRNALIAIRSLLMNQPDNHWRILAAGIVPVLLDLVKSEEISDGVTADSMAILAKMAE 577
Query: 454 HQEAKVSIVKASTIPVLIDLLRTG--LPRNKENAAAILLALCKR-DTDNLSCISRLGAVI 510
+ + +S+++ + + + +L + P K++ A+LL LC +D + +++ +++
Sbjct: 578 YPDGMISVLRRGGLKLAVKILGSSEVSPATKQHCVALLLNLCHNGGSDVVGSLAKNPSIM 637
Query: 511 -PLSELARTGTERAKRKATSLLEHLRKLQQ 539
L + G +KA++L++ + + Q+
Sbjct: 638 GSLYTASSNGELGGGKKASALIKMIHEFQE 667
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.315 0.131 0.359
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,947,139
Number of Sequences: 26719
Number of extensions: 446633
Number of successful extensions: 2037
Number of sequences better than 10.0: 136
Number of HSP's better than 10.0 without gapping: 99
Number of HSP's successfully gapped in prelim test: 37
Number of HSP's that attempted gapping in prelim test: 1405
Number of HSP's gapped (non-prelim): 264
length of query: 540
length of database: 11,318,596
effective HSP length: 104
effective length of query: 436
effective length of database: 8,539,820
effective search space: 3723361520
effective search space used: 3723361520
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 62 (28.5 bits)
Medicago: description of AC148469.7


