

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148405.4 - phase: 0
(609 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
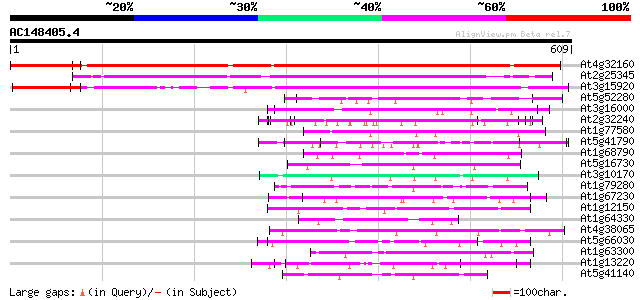
Score E
Sequences producing significant alignments: (bits) Value
At4g32160 unknown protein 448 e-126
At2g25345 putative protein 371 e-103
At3g15920 hypothetical protein 291 7e-79
At5g52280 hyaluronan mediated motility receptor-like protein 60 3e-09
At3g16000 myosin heavy chain like protein 60 4e-09
At2g32240 putative myosin heavy chain 57 4e-08
At1g77580 unknown protein 54 2e-07
At5g41790 myosin heavy chain-like protein 54 3e-07
At1g68790 putative nuclear matrix constituent protein 1 (NMCP1) 52 1e-06
At5g16730 putative protein 51 2e-06
At3g10170 hypothetical protein 51 2e-06
At1g79280 hypothetical protein 51 2e-06
At1g67230 unknown protein 51 2e-06
At1g12150 hypothetical protein 50 5e-06
At1g64330 unknown protein 49 6e-06
At4g38065 hypothetical protein 48 1e-05
At5g66030 Golgi-localized protein GRIP 47 2e-05
At1g63300 hypothetical protein 47 2e-05
At1g13220 putative nuclear matrix constituent protein 47 2e-05
At5g41140 putative protein 47 4e-05
>At4g32160 unknown protein
Length = 723
Score = 448 bits (1153), Expect = e-126
Identities = 272/538 (50%), Positives = 350/538 (64%), Gaps = 23/538 (4%)
Query: 69 SAFQDASQQNSVKDPDSNNRDYSVQSS--LHSGLSLT-AGSSSVASDYGSDTPYDASEVG 125
SA QD Q S DSNN S SS +H LSL AG S++ SDYGSDT Y+ SEVG
Sbjct: 166 SACQDVDQNAS----DSNNDRSSTSSSPMVHPSLSLFHAGGSTLTSDYGSDTAYETSEVG 221
Query: 126 TPRIGQDDNSEVGTDDLTLDEDTT--NPIEKLVKYGISNIDEGLFMGQTILEQLEGLPRH 183
+P +GQDD SE+GT+DLTLDED T NPIEKLV + +SNIDEGL M +TILEQLE P+H
Sbjct: 222 SPSVGQDDISEIGTEDLTLDEDLTLTNPIEKLVNFSMSNIDEGLSMSETILEQLEDFPKH 281
Query: 184 KVNARHGNNVTGKDRSNGNSYDSSLLGNNTMELFSQPGHAK-VLGHIRKLSNESVGSDGS 242
KV +R+ NN+ GKD NGN+ L NN L S+P + + H R S E
Sbjct: 282 KVRSRYVNNILGKDVYNGNASKGVFLANNGSRLLSEPEPSTHSVMHDRNDSAERF----- 336
Query: 243 SIRGSDISNFGIPNSSGDGSVDLPGSASVSRETDIVGHTKLKSSGDAQLVLPLDQRNKLN 302
++ S G+ SS D +DL VS T +V + + + G AQ+VLPL+ RNKLN
Sbjct: 337 ALHTGQTSTSGLLISSRDSHLDLRQGPGVSLGTGLVCNPERQ--GSAQIVLPLELRNKLN 394
Query: 303 RILSTMQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEVELETSKQKNKENLQQAI 362
RIL RLV AKTDMEDLI RLNQEIA KD+L KV DLE ELET+KQ++KENL+QAI
Sbjct: 395 RILLATNERLVNAKTDMEDLIARLNQEIAVKDYLNKKVNDLEGELETTKQRSKENLEQAI 454
Query: 363 LIERERFTQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLTKESIVQQNDVLLQDLDAAR 422
+ ERERF QMQWDMEELR+KS EMEMKLKS G+ + T +S + + VL ++LDA +
Sbjct: 455 MSERERFNQMQWDMEELRQKSYEMEMKLKSREDGSSHAEPTVQSTISEKHVLSKELDARK 514
Query: 423 EQLEILSKQYGELEAKSKADIKVLVKEVKSLRSSQTELKKELSESIKEKYEAEKLLLHER 482
+QLE LS++Y ELEAKSKAD+KVLVKEVKSLR S EL+KEL+ S+ +K AEKLL ER
Sbjct: 515 QQLEDLSRRYEELEAKSKADMKVLVKEVKSLRRSHVELEKELTHSLTDKTNAEKLLQEER 574
Query: 483 EKSEQAEIAWRKQLEKCGLLLKQLQECSVELPYEDEDRTFLQSSSSTDAFNKLKTSDDQI 542
+ E A +K L C +L +L+E ++ L + S++ +D L SDDQI
Sbjct: 575 KLLENTVAARKKLLSDCRILHDRLKEYNLNLSMDGNGNFVDDSTTISDVLRLLSISDDQI 634
Query: 543 DILLAEVENLEK-DAGSAASNVDKTNDI-KDGVICGDEMRKIISDLFLDNVRLRKQTN 598
+ E + L D +AA ++DKT + + I DE+RKI++++F++N +LRKQ N
Sbjct: 635 E----EAQLLSGFDENAAAEDIDKTLSMDTETRIMEDELRKILANIFVENAKLRKQVN 688
Score = 140 bits (353), Expect = 2e-33
Identities = 59/76 (77%), Positives = 65/76 (84%)
Query: 1 MQRRSPPKHRHDGTSPLPLGMDWSPAPRKWNGRDTEWPHNHRTGWSYCVIIPSWVFVPKS 60
MQRRSPPKHRHDGTSPLPLGMDWSP PRKWNGRDT WPH+ RTGWSYCV IPSW+ +PKS
Sbjct: 2 MQRRSPPKHRHDGTSPLPLGMDWSPPPRKWNGRDTVWPHDPRTGWSYCVTIPSWIVLPKS 61
Query: 61 KNSDPIVMSAFQDASQ 76
+NSDP+V Q + Q
Sbjct: 62 RNSDPVVFYRVQVSVQ 77
>At2g25345 putative protein
Length = 600
Score = 371 bits (952), Expect = e-103
Identities = 228/521 (43%), Positives = 316/521 (59%), Gaps = 42/521 (8%)
Query: 69 SAFQDASQQNSVKDPDSNNRDYSVQSSLHSGLSLTAGSSSVASDYGSDTPYDASEVGTPR 128
SA Q Q S + D N+ S+ S +H + GSS ++SDYGSDT Y+ SE+G+
Sbjct: 122 SACQVVDQNASDSNDDGNST--SLSSLVHPN---SGGSSLLSSDYGSDTAYETSELGSAS 176
Query: 129 IGQDDNSEVGTDDLTLDEDTTNPIEKLVKYGISNIDEGLFMGQTILEQLEGLPRHKVNAR 188
+GQDD SE T DLTLDED TNP EKLVK+ +SNIDEGL M QTI+EQLE P+H+V+
Sbjct: 177 LGQDDVSETDTGDLTLDEDLTNPTEKLVKFSMSNIDEGLSMSQTIIEQLEDFPKHRVHLG 236
Query: 189 HGNNVTGKDRSNGNSYDSSLLGNNTMELFSQPGHAKVLGHIRKLSNESVGSDGSSIRGSD 248
+ N++T + NG + NN + S+ + + H RKLS ES +DG S+ +
Sbjct: 237 YANDITETNSYNGKASKGVFRANNDLRCRSESETSHSVMHDRKLSLES--ADGVSLLAGE 294
Query: 249 ISNFGIPNSSGDGSVDLPGSASVSRETDIVGHTKLKSSGDAQLVLPLDQRNKLNRILSTM 308
S I +S +D+ SV L+ G+ ++VLPL +KLNRIL TM
Sbjct: 295 TSTSSILSSIVHSQLDVNHDISVGN---------LEIPGNGRIVLPLKMHSKLNRILLTM 345
Query: 309 QRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEVELETSKQKNKENLQQAILIERER 368
RL+ +KTDMEDLI RLNQE A K++L KV DLEVELET+KQ+NKENL+QA++ ER+
Sbjct: 346 NERLLNSKTDMEDLIARLNQETAVKEYLNRKVDDLEVELETTKQRNKENLEQALMTERQS 405
Query: 369 FTQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLTKESIVQQNDVLLQDLDAAREQLEIL 428
T+MQWDMEELR+K+ EME+KLKS+ G+ + S + ++ LLQ++DA ++QLE L
Sbjct: 406 VTKMQWDMEELRQKTFEMELKLKSKEDGSSDSKTSGNSTISESHELLQEMDATKQQLEDL 465
Query: 429 SKQYGELEAKSKADIKVLVKEVKSLRSSQTELKKELSESIKEKYEAEKLLLHEREKSEQA 488
S++Y ELEAKSKADIKVLV+EVKSLR S E++KEL+ S+ EK + EKLL ER E
Sbjct: 466 SRRYVELEAKSKADIKVLVREVKSLRRSHMEMEKELTRSLTEKSDTEKLLQQERIIVENT 525
Query: 489 EIAWRKQLEKCGLLLKQLQECSVELPYEDEDRTFLQSSSSTDAFNKLKTSDDQIDILLAE 548
A R+ C +L +L+ + L ++ +S+++ D L+E
Sbjct: 526 LEARRRLYSDCEILHDRLKVNNTNLSMDE-------------------SSNNRED--LSE 564
Query: 549 VENLEKDAGSAASNVDKTNDIKDGVICGDEMRKIISDLFLD 589
V N +D A + D DE+RKI++D++ D
Sbjct: 565 VSNALQDQIEAQLLLG-----FDETASEDELRKIMADMYED 600
>At3g15920 hypothetical protein
Length = 755
Score = 291 bits (745), Expect = 7e-79
Identities = 190/546 (34%), Positives = 314/546 (56%), Gaps = 52/546 (9%)
Query: 67 VMSAFQDASQQNSVKDPDSNNRDYSVQSSLHSGLSLTAGSSSVASDYGSDTPYDASEVGT 126
V S F D Q+ D + S L + +S GSSSV D+ +D+ + S T
Sbjct: 223 VRSYFNDEYQETEDTSGD-------IPSLLPTTISDVPGSSSVTIDHDNDSADETSNAST 275
Query: 127 PRIGQDDNSEVGTDDLTLDEDTTNPIEKLVKYGISNIDEGLFMGQTILEQLEGLPRHKVN 186
+ + + + + + T ++ T+ E + +YG+ +D+ F E++E +++
Sbjct: 276 MKHDEANLKNLFSRNSTAVDNVTDWHELITEYGL--LDQSSFQ-----EKVE-----RLS 323
Query: 187 ARHGNNVTGKDRSNGNSYDSSLLGNNTMELFSQPGHAKVLGHIRKLSNESVGSDGSSIRG 246
+ +G+ TG G S S G ++ G RK ++ SI+
Sbjct: 324 STNGDAATGTVTRGGIS--------------SGVGIQRLDGSDRKFQELTI----ESIKK 365
Query: 247 SDISNFGI-----PNSSGDGSVDLPGSASVSRETDIVGHTKLKSSGDAQLVLPLDQRNKL 301
+ +S+F P+ G++D+ G A + + G T+ + D +V ++R+KL
Sbjct: 366 THVSDFEASTAVEPDLVNQGAMDIHGEAHGNMYGAVGGDTETQK--DLAIVFQSEERHKL 423
Query: 302 NRILSTMQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEVELETSKQKNKENLQQA 361
R++ T+++RL TAK D EDLI RLNQE+A + FL+TKV+DLEVELET+++ K+ +++
Sbjct: 424 KRVIDTLKQRLETAKADTEDLISRLNQELAVRQFLSTKVRDLEVELETTRESCKQGMEKT 483
Query: 362 ILIERERFTQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLTKESIVQQNDVLLQDLDAA 421
+L E+ERFTQ+QWDMEELR++ +EME L S + ES+VQ+N +LLQ ++
Sbjct: 484 VLDEKERFTQIQWDMEELRKQCMEMESFLNSIKDEKTHIETANESLVQENQMLLQQINDI 543
Query: 422 REQLEILSKQYGELEAKSKADIKVLVKEVKSLRSSQTELKKELSESIKEKYEAEKLLLHE 481
RE E K++ ELE K+KA++KVLVKEVKSLR++Q++L++ELS +KEK E E+++ E
Sbjct: 544 RENFENFHKEHEELEVKAKAELKVLVKEVKSLRTTQSDLRQELSGIMKEKLEMERIVQRE 603
Query: 482 REKSEQAEIAWRKQLEKCGLLLKQLQECSVELPYEDEDRTFLQSSSSTDAFNKLKTSDDQ 541
+++ E A+ A +K L +C +L +LQEC+V+ E+E + + SSS ++A L TSD++
Sbjct: 604 KDREETAKNADKKLLHECDVLQNRLQECNVKFDIEEEGKLIMDSSSLSEAIELLATSDNR 663
Query: 542 IDILLAEVENLEKDAGSAASNVDKTNDIKDGVICGDEM-RKIISDLFLDNVRLRKQTNRV 600
I +L+AE + L ++ V+K G D++ RK+++++ +DN RLRKQ N V
Sbjct: 664 IGLLIAETQLLSEE-------VEKLKLTSGGHRGTDDLVRKMLTEVLIDNARLRKQVNSV 716
Query: 601 TRHMLS 606
R LS
Sbjct: 717 LRCSLS 722
Score = 111 bits (278), Expect = 1e-24
Identities = 49/73 (67%), Positives = 53/73 (72%)
Query: 4 RSPPKHRHDGTSPLPLGMDWSPAPRKWNGRDTEWPHNHRTGWSYCVIIPSWVFVPKSKNS 63
R+ P HRHDG SPLPLGMDWS PR GRDT WPH+HRTGWSYCV +PSWV +PKS S
Sbjct: 64 RASPPHRHDGRSPLPLGMDWSSPPRHLEGRDTVWPHDHRTGWSYCVTVPSWVDLPKSSVS 123
Query: 64 DPIVMSAFQDASQ 76
DP V Q A Q
Sbjct: 124 DPAVFYRVQVAIQ 136
>At5g52280 hyaluronan mediated motility receptor-like protein
Length = 853
Score = 60.5 bits (145), Expect = 3e-09
Identities = 64/273 (23%), Positives = 115/273 (41%), Gaps = 41/273 (15%)
Query: 312 LVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEVELETSKQKNKENLQQAILIERERFTQ 371
L+ A D+ +++ + N EI++ + L + K LE + K ++ I +++
Sbjct: 382 LILAVRDLNEMLEQKNNEISSLNSLLEEAKKLE------EHKGMDSGNNEIDTLKQQIED 435
Query: 372 MQWDMEELRRKSLEMEM----------KLKSESGGNVGQDLTKESIVQQNDVLLQD---L 418
+ W+++ ++K+ E E+ LK E+ NV L ++ D L +
Sbjct: 436 LDWELDSYKKKNEEQEILLDELTQEYESLKEENYKNVSSKLEQQECSNAEDEYLDSKDII 495
Query: 419 DAAREQLEILSKQYGELEAKSKADIKVLVKEVKSLRSSQTELKKELSESIKEKYEAEKLL 478
D + Q+EIL G+L+ +S + L+ V L S ELKKEL + + E +
Sbjct: 496 DELKSQIEILE---GKLKQQSLEYSECLIT-VNELESQVKELKKELEDQAQAYDEDIDTM 551
Query: 479 LHEREKSEQAEIAWRKQLEKC----GLLLKQLQECSVELPYEDEDRTFLQSSSSTDAFNK 534
+ E+ + EQ I + L K + ++LQE L E E +K
Sbjct: 552 MREKTEQEQRAIKAEENLRKTRWNNAITAERLQEKCKRLSLEME--------------SK 597
Query: 535 LKTSDDQIDILLAEVENLEKDAGSAASNVDKTN 567
L ++ LAE NL + +KT+
Sbjct: 598 LSEHENLTKKTLAEANNLRLQNKTLEEMQEKTH 630
Score = 55.8 bits (133), Expect = 7e-08
Identities = 71/323 (21%), Positives = 136/323 (41%), Gaps = 63/323 (19%)
Query: 299 NKLNRILSTMQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEVELETSKQKNKENL 358
++L + ++ +L + + ++ +N+ L ++VK+L+ ELE Q E++
Sbjct: 496 DELKSQIEILEGKLKQQSLEYSECLITVNE-------LESQVKELKKELEDQAQAYDEDI 548
Query: 359 -----------QQAILIERERFTQMQWD--------MEELRRKSLEMEMKLKSESGGNVG 399
Q+AI E E + +W+ E+ +R SLEME KL
Sbjct: 549 DTMMREKTEQEQRAIKAE-ENLRKTRWNNAITAERLQEKCKRLSLEMESKLSEH------ 601
Query: 400 QDLTKESIVQQNDVLLQD--LDAAREQLEILSKQYGELEAKSKADIKVLVKEVKSLRSSQ 457
++LTK+++ + N++ LQ+ L+ +E+ Q E + K L +V+ L S
Sbjct: 602 ENLTKKTLAEANNLRLQNKTLEEMQEKTHTEITQEKEQRKHVEEKNKALSMKVQMLESEV 661
Query: 458 TELKKELSESIKEKYEAEKLLLHEREKSEQAEIAWRKQLEKCGLLLKQLQECSVELPYED 517
+L K ES E EK++ R++ ++ E K L + + EL
Sbjct: 662 LKLTKLRDESSAAATETEKIIQEWRKERDEFE-------RKLSLAKEVAKTAQKELT--- 711
Query: 518 EDRTFLQSSSSTDAFNKLKTSDDQIDILLAEVENLEKDAGSAASNVDKTNDIKDGVICGD 577
L SS+ D +L+ +++ L + L+ N + D
Sbjct: 712 -----LTKSSNDDKETRLRNLKTEVEGLSLQYSELQ-------------NSFVQEKMEND 753
Query: 578 EMRKIISDLFLDNVRLRKQTNRV 600
E+RK +S+L +D R ++ ++
Sbjct: 754 ELRKQVSNLKVDIRRKEEEMTKI 776
>At3g16000 myosin heavy chain like protein
Length = 727
Score = 60.1 bits (144), Expect = 4e-09
Identities = 71/305 (23%), Positives = 130/305 (42%), Gaps = 16/305 (5%)
Query: 288 DAQLVLPLDQRNKLNRILSTMQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEVEL 347
+A+L ++R K Q L+ +DL+ L +E++++ L K+KD L
Sbjct: 181 EAKLQHEQEERKKEVEKAKEEQLSLINQLNSAKDLVTELGRELSSEKKLCEKLKDQIESL 240
Query: 348 ETSKQKNKENLQQAILIERERFTQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLTKESI 407
E S K E+ + RE+ ++ + + SLE++ + N + +
Sbjct: 241 ENSLSKAGEDKEALETKLREKLDLVEGLQDRINLLSLELKDSEEKAQRFNASLAKKEAEL 300
Query: 408 VQQNDVLLQ-DLDAAREQLEILSKQYGELEAKSKADIKVLVKEVKSLRSSQTELKKE--- 463
+ N + Q D A +LEI ++ + +S+ D K E + R + +KE
Sbjct: 301 KELNSIYTQTSRDLAEAKLEIKQQKEELIRTQSELDSKNSAIEELNTRITTLVAEKESYI 360
Query: 464 -LSESIKEKYEAEKLLLHEREKSEQAEIAWRKQLEKCGLLLKQLQECSVELPYEDEDRTF 522
+SI + Y A K L E + + AE+ RK+ E ++QL E +++ +D +++
Sbjct: 361 QKLDSISKDYSALK-LTSETQAAADAELISRKEQE-----IQQLNE-NLDRALDDVNKS- 412
Query: 523 LQSSSSTDAFNKLKTSDDQIDILLAEVENLEKDAGSAASNVDKTND-IKDGVICGDEMRK 581
D K + S +DI L V+NL + + + D + D DE R
Sbjct: 413 --KDKVADLTEKYEDSKRMLDIELTTVKNLRHELEGTKKTLQASRDRVSDLETMLDESRA 470
Query: 582 IISDL 586
+ S L
Sbjct: 471 LCSKL 475
Score = 56.2 bits (134), Expect = 5e-08
Identities = 71/329 (21%), Positives = 141/329 (42%), Gaps = 45/329 (13%)
Query: 281 TKLKSSGDAQLVLPLDQR-NKLNRILSTMQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTK 339
++ +++ DA+L+ +Q +LN L + +K + DL + D T
Sbjct: 377 SETQAAADAELISRKEQEIQQLNENLDRALDDVNKSKDKVADLTEKYEDSKRMLDIELTT 436
Query: 340 VKDLEVELETSKQKNKENLQQAILIERERFTQMQWDMEELRRKSLEMEMKL--------- 390
VK+L ELE +K+ + R+R + ++ ++E R ++E +L
Sbjct: 437 VKNLRHELEGTKK--------TLQASRDRVSDLETMLDESRALCSKLESELAIVHEEWKE 488
Query: 391 -KSESGGNVGQDLTKESIVQQNDVLLQDLDA-AREQLEILSKQYGELEAKSKADIKVLVK 448
K N+ + K I L +DL +++LE ++ + E K+++ K LV+
Sbjct: 489 AKERYERNLDAEKQKNEISASELALEKDLRRRVKDELEGVTHELKESSVKNQSLQKELVE 548
Query: 449 EVKSLRSSQTELKKELSESI---KEKYEAEKLLLHEREKSEQAEIAWRKQLEKC------ 499
K + +S EL++E + KE EK +L ERE + E + ++
Sbjct: 549 IYKKVETSNKELEEEKKTVLSLNKEVKGMEKQILMEREARKSLETDLEEAVKSLDEMNKN 608
Query: 500 -GLLLKQLQECSVELPYEDEDRTFLQSSSSTDAFNKLKTSDDQID--------------I 544
+L ++L++ + ++++ LQ S +A N K + + ++ +
Sbjct: 609 TSILSRELEKVNTHASNLEDEKEVLQRSLG-EAKNASKEAKENVEDAHILVMSLGKEREV 667
Query: 545 LLAEVENLEKDAGSAASNVDKTNDIKDGV 573
L +V+ LE+D GSA + + D V
Sbjct: 668 LEKKVKKLEEDLGSAKGEILRMRSQPDSV 696
>At2g32240 putative myosin heavy chain
Length = 1333
Score = 56.6 bits (135), Expect = 4e-08
Identities = 70/297 (23%), Positives = 131/297 (43%), Gaps = 21/297 (7%)
Query: 282 KLKSSGDAQLVLPLDQRNKLNRI---LSTMQRRLVTAKTDMEDLIVRLNQEI----AAKD 334
KL+++ D V + N L I L+ Q +L + + D++ ++ ++ + +A++
Sbjct: 715 KLEATVDEYSVKISESENLLESIRNELNVTQGKLESIENDLKAAGLQESEVMEKLKSAEE 774
Query: 335 FLTTKVKDLEVELETSKQKNKENLQQAILIERER--------FTQMQWDMEELRRKSLEM 386
L K + E++ T+K+ E L Q++ I+ E FT + L K ++
Sbjct: 775 SLEQKGR--EIDEATTKRMELEALHQSLSIDSEHRLQKAMEEFTSRDSEASSLTEKLRDL 832
Query: 387 EMKLKSESGGNVGQDLTKESIVQQNDVLLQDLDAAREQLEILSKQYGELEAKS---KADI 443
E K+KS S+ ++ + L L AA E L +++ + + KS ++
Sbjct: 833 EGKIKSYEEQLAEASGKSSSLKEKLEQTLGRLAAAESVNEKLKQEFDQAQEKSLQSSSES 892
Query: 444 KVLVKEVKSLRSSQTELKKELSESIKEKYEAEKLLLHEREKSEQAEIAWRKQLEKCGLLL 503
++L + L+ EL+ + EK A K L E+ Q E +EK
Sbjct: 893 ELLAETNNQLKIKIQELEGLIGSGSVEKETALKRLEEAIERFNQKETESSDLVEKLKTHE 952
Query: 504 KQLQECSVELPYEDEDRTFLQSSSSTDAFNKLKTSDDQIDILLAEVENLEKDAGSAA 560
Q++E +L +E + DA +KLK + I+ L A+ + LEK++G A
Sbjct: 953 NQIEEYK-KLAHEASGVADTRKVELEDALSKLKNLESTIEELGAKCQGLEKESGDLA 1008
Score = 53.5 bits (127), Expect = 3e-07
Identities = 70/309 (22%), Positives = 133/309 (42%), Gaps = 35/309 (11%)
Query: 281 TKLKSSGDAQLVLPLDQRNKLNRILSTMQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKV 340
T++K + DA + R KL + ++R A+ E+L Q + D + K
Sbjct: 180 TEVKEAFDALGIELESSRKKLIELEEGLKRSAEEAQK-FEELH---KQSASHADSESQKA 235
Query: 341 KDLEVELETSKQKNKENLQQAILIERERFTQMQWDMEELRRKSLE---MEMKLKSESGG- 396
+ L+++K+ KE E+ +Q +++EL K E +E LKS +G
Sbjct: 236 LEFSELLKSTKESAKEM--------EEKMASLQQEIKELNEKMSENEKVEAALKSSAGEL 287
Query: 397 ---------NVGQDLTKESIVQQNDVLLQDLDAAREQLEILSKQYGELEAKSKADIKVLV 447
+ + L E V + L+ +L EQ + ++ E E D+
Sbjct: 288 AAVQEELALSKSRLLETEQKVSSTEALIDELTQELEQKKASESRFKE-ELSVLQDLDAQT 346
Query: 448 KEVKSLRSSQTELKKELSESIKEKYEAEKLLLHEREKSEQAEIAWRKQLEKCGLLLKQLQ 507
K +++ S Q + +L+E +KEK E L + EK A + L++ L +
Sbjct: 347 KGLQAKLSEQEGINSKLAEELKEKELLESLSKDQEEKLRTANEKLAEVLKEKEALEANVA 406
Query: 508 ECSVELP--------YEDEDRTFLQSSSSTDAF-NKLKTSDDQIDILLAEVENLEKDAGS 558
E + + E++ +T ++ S TDA ++ +++ +++ L +E L +AGS
Sbjct: 407 EVTSNVATVTEVCNELEEKLKTSDENFSKTDALLSQALSNNSELEQKLKSLEELHSEAGS 466
Query: 559 AASNVDKTN 567
AA+ + N
Sbjct: 467 AAAAATQKN 475
Score = 50.8 bits (120), Expect = 2e-06
Identities = 75/312 (24%), Positives = 137/312 (43%), Gaps = 32/312 (10%)
Query: 284 KSSGDAQLVLPLDQRNKLNRILSTMQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKD- 342
K GD ++V Q+ + +I+ +R K+ +ED + Q AKD T+VK+
Sbjct: 135 KKHGDLEVV----QKKQQEKIVEGEERHSSQLKS-LEDAL----QSHDAKDKELTEVKEA 185
Query: 343 ---LEVELETSKQKN-------KENLQQAILIERERFTQMQWDMEELRRKSLEMEMKLKS 392
L +ELE+S++K K + ++A E E Q + +K+LE LKS
Sbjct: 186 FDALGIELESSRKKLIELEEGLKRSAEEAQKFE-ELHKQSASHADSESQKALEFSELLKS 244
Query: 393 ESGGNVGQDLTKESIVQQNDVLLQDLDAAREQLEILSKQYGELEAKSK--ADIKVLVKEV 450
+ S+ Q+ L + + + L GEL A + A K + E
Sbjct: 245 TKESAKEMEEKMASLQQEIKELNEKMSENEKVEAALKSSAGELAAVQEELALSKSRLLET 304
Query: 451 KSLRSSQTELKKELSESIKEKYEAEKLLLHEREKSEQAEIAWRKQLEKCGLLLKQLQECS 510
+ SS L EL++ +++K +E E + + + K L + + +
Sbjct: 305 EQKVSSTEALIDELTQELEQKKASESRFKEELSVLQDLDAQTKGLQAK----LSEQEGIN 360
Query: 511 VELPYEDEDRTFLQSSSSTDAFNKLKTSDDQIDILLAEVENLEKDAGSAASNV----DKT 566
+L E +++ L+S S D KL+T+++++ +L E E LE + SNV +
Sbjct: 361 SKLAEELKEKELLESLSK-DQEEKLRTANEKLAEVLKEKEALEANVAEVTSNVATVTEVC 419
Query: 567 NDIKDGVICGDE 578
N++++ + DE
Sbjct: 420 NELEEKLKTSDE 431
Score = 47.4 bits (111), Expect = 2e-05
Identities = 60/247 (24%), Positives = 105/247 (42%), Gaps = 14/247 (5%)
Query: 310 RRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEVELETSKQKNKENLQQAILIERERF 369
+ L T T E L Q++ + L K D E EL+ +K+ E LQ AI + E
Sbjct: 497 KELETKFTAAEQKNAELEQQL---NLLQLKSSDAERELKELSEKSSE-LQTAIEVAEEEK 552
Query: 370 TQMQWDMEELRRKSLEMEMKLKSESGGN--VGQDLTKESIVQQNDVLLQDLDAAREQLEI 427
Q M+E ++K+ E+E+ L S N + +DL I Q +D Q I
Sbjct: 553 KQATTQMQEYKQKASELELSLTQSSARNSELEEDL---RIALQKGAEHEDRANTTHQRSI 609
Query: 428 LSKQYGELEAKSKADIKVLVKEVKSLRSSQTELKKELSESIK--EKYEAEKLLLHEREKS 485
+ + D + +K+++ L ++ +EL E + EK E +
Sbjct: 610 ELEGLCQSSQSKHEDAEGRLKDLELLLQTEKYRIQELEEQVSSLEKKHGETEADSKGYLG 669
Query: 486 EQAEIAWRKQLEKCGLLLKQLQECSVELPYEDEDRTFLQSSSSTDAFNKLKTSDDQIDIL 545
+ AE+ + LE + L E ++ + E+E ++ T KL+ + D+ +
Sbjct: 670 QVAEL--QSTLEAFQVKSSSL-EAALNIATENEKELTENLNAVTSEKKKLEATVDEYSVK 726
Query: 546 LAEVENL 552
++E ENL
Sbjct: 727 ISESENL 733
Score = 45.4 bits (106), Expect = 9e-05
Identities = 47/221 (21%), Positives = 91/221 (40%), Gaps = 18/221 (8%)
Query: 305 LSTMQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEVELETSKQKNKENLQQAILI 364
L + L ++ E + +L +E+ K+ L + KD E +L T+ +K E L++ +
Sbjct: 342 LDAQTKGLQAKLSEQEGINSKLAEELKEKELLESLSKDQEEKLRTANEKLAEVLKEKEAL 401
Query: 365 ERERFTQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLTKESIVQQNDVLLQDLDAARE- 423
E ++ ++ + E+E KLK+ D + N L Q L + E
Sbjct: 402 EAN-VAEVTSNVATVTEVCNELEEKLKTSDENFSKTDALLSQALSNNSELEQKLKSLEEL 460
Query: 424 ----------------QLEILSKQYGELEAKSKADIKVLVKEVKSLRSSQTELKKELSES 467
+LE + + + ++K+ IK L + + EL+++L+
Sbjct: 461 HSEAGSAAAAATQKNLELEDVVRSSSQAAEEAKSQIKELETKFTAAEQKNAELEQQLNLL 520
Query: 468 IKEKYEAEKLLLHEREKSEQAEIAWRKQLEKCGLLLKQLQE 508
+ +AE+ L EKS + + A E+ Q+QE
Sbjct: 521 QLKSSDAERELKELSEKSSELQTAIEVAEEEKKQATTQMQE 561
Score = 43.9 bits (102), Expect = 3e-04
Identities = 73/331 (22%), Positives = 142/331 (42%), Gaps = 38/331 (11%)
Query: 271 VSRETDIVGHTKLKSSGDAQLVLPLDQRNK-----LNRILSTMQRRLVTAKTDMEDLIVR 325
+ +E D L+SS +++L+ + + K L ++ + TA +E+ I R
Sbjct: 874 LKQEFDQAQEKSLQSSSESELLAETNNQLKIKIQELEGLIGSGSVEKETALKRLEEAIER 933
Query: 326 LNQEIAAKDFLTTKVKDLEVELETSKQKNKE--NLQQAILIERE----RFTQMQWDMEEL 379
NQ+ L K+K E ++E K+ E + +E E + ++ +EEL
Sbjct: 934 FNQKETESSDLVEKLKTHENQIEEYKKLAHEASGVADTRKVELEDALSKLKNLESTIEEL 993
Query: 380 RRKS----------LEMEMKLKSE---SGGNVGQDLTKESIVQ-QNDVLLQDLDAAREQL 425
K E+ +KL E G + TK S ++ + + +L+A++ +
Sbjct: 994 GAKCQGLEKESGDLAEVNLKLNLELANHGSEANELQTKLSALEAEKEQTANELEASKTTI 1053
Query: 426 EILSKQYGELEAKSKADIKVLVKEVKSLRSSQTELKKELSESIKEKYEAEKLLLHEREKS 485
E L+KQ K ++ I +E + + K+EL +S+ K E + + + +
Sbjct: 1054 EDLTKQLTSEGEKLQSQISSHTEENNQVNAMFQSTKEEL-QSVIAKLEEQLTVESSKADT 1112
Query: 486 EQAEIAWRKQL--EKCGL------LLKQLQECSVELPYEDEDRTFLQSSSSTDAFNKLKT 537
+EI + + EK L L K L E +L E+ + S + +KL+
Sbjct: 1113 LVSEIEKLRAVAAEKSVLESHFEELEKTLSEVKAQLK-ENVENAATASVKVAELTSKLQE 1171
Query: 538 SD---DQIDILLAEVENLEKDAGSAASNVDK 565
+ + D+L +V L+K+ +A S++D+
Sbjct: 1172 HEHIAGERDVLNEQVLQLQKELQAAQSSIDE 1202
Score = 42.0 bits (97), Expect = 0.001
Identities = 58/298 (19%), Positives = 131/298 (43%), Gaps = 32/298 (10%)
Query: 321 DLIVRLNQEIAAKDFLTTKVKDLEVELETSKQKNKENLQQAILIERER------FTQMQW 374
DL+ +N E+ ++ + ++VE E K+ + +E ++ + Q
Sbjct: 27 DLLQAVNGEVPKEEKEEEDGEFIKVEKEAFDAKDDAEKADHVPVEEQKEVIERSSSGSQR 86
Query: 375 DMEELRRKSLEMEMKLKSESGGNVGQDLTKESIVQQNDVLLQDLDAAREQLEILSKQYGE 434
++ E + K+ E+E++L+ +G + +N L +L +A+E+LE K++G+
Sbjct: 87 ELHESQEKAKELELELERVAG-------ELKRYESENTHLKDELLSAKEKLEETEKKHGD 139
Query: 435 LEAKSKADIKVLVK-------EVKSLR---SSQTELKKELSESIKEKYEAEKLLLHEREK 484
LE K + +V+ ++KSL S KEL+E +KE ++A L E E
Sbjct: 140 LEVVQKKQQEKIVEGEERHSSQLKSLEDALQSHDAKDKELTE-VKEAFDA---LGIELES 195
Query: 485 SEQAEIAWRKQLEKCGLLLKQLQECSVELPYEDEDRTFLQSSSSTDAFNKLKTSDDQIDI 544
S + I + L++ ++ + EL + +S + + LK++ +
Sbjct: 196 SRKKLIELEEGLKRSAEEAQKFE----ELHKQSASHADSESQKALEFSELLKSTKESAKE 251
Query: 545 LLAEVENLEKDAGSAASNVDKTNDIKDGV-ICGDEMRKIISDLFLDNVRLRKQTNRVT 601
+ ++ +L+++ + + ++ + E+ + +L L RL + +V+
Sbjct: 252 MEEKMASLQQEIKELNEKMSENEKVEAALKSSAGELAAVQEELALSKSRLLETEQKVS 309
>At1g77580 unknown protein
Length = 779
Score = 54.3 bits (129), Expect = 2e-07
Identities = 70/298 (23%), Positives = 130/298 (43%), Gaps = 38/298 (12%)
Query: 320 EDLIVRLNQEIAAK-DFLTTKVKDLEVELETSKQKNKENLQQAILIERERFTQMQWDMEE 378
E+++V +A++ + LT+++K+LE +LE + K L+ + RE + E
Sbjct: 331 EEVVVPSENSLASEIEVLTSRIKELEEKLE-KLEAEKHELENEVKCNREEAVVHIENSEV 389
Query: 379 LRRKSLEMEMKL----------KSESGGNVGQDLT--KESIVQQNDVLLQDLDAAREQLE 426
L ++ E+E KL KSE N + + + S+ + +VL EQLE
Sbjct: 390 LTSRTKELEEKLEKLEAEKEELKSEVKCNREKAVVHVENSLAAEIEVLTSRTKELEEQLE 449
Query: 427 ILSKQYGELEAKSK---------------ADIKVLVKEVKSLRSSQTEL---KKELSESI 468
L + ELE++ K +I+VL +K L +L K EL +
Sbjct: 450 KLEAEKVELESEVKCNREEAVAQVENSLATEIEVLTCRIKQLEEKLEKLEVEKDELKSEV 509
Query: 469 KEKYEAEKLLLHEREKSEQAEIAWRKQLEKCGLLLKQLQECSVELPYEDEDRTFLQSSSS 528
K E E L E E ++ +LEK + +LQ S ++ + + + +
Sbjct: 510 KCNREVESTLRFELEAIACEKMELENKLEKLEVEKAELQ-ISFDIIKDKYEESQVCLQEI 568
Query: 529 TDAFNKLKTSDDQIDILLAEVEN----LEKDAGSAASNVDK-TNDIKDGVICGDEMRK 581
+++T ++ L AEVE+ +E DA + ++ ++ D++ DE+R+
Sbjct: 569 ETKLGEIQTEMKLVNELKAEVESQTIAMEADAKTKSAKIESLEEDMRKERFAFDELRR 626
Score = 35.0 bits (79), Expect = 0.12
Identities = 40/202 (19%), Positives = 94/202 (45%), Gaps = 15/202 (7%)
Query: 394 SGGNVGQDLTKESIVQQNDVLLQDLDAAREQLEILSKQYGELEAK----------SKADI 443
+G + + +T+E +V + L +++ +++ L ++ +LEA+ ++ +
Sbjct: 321 NGKSGPESVTEEVVVPSENSLASEIEVLTSRIKELEEKLEKLEAEKHELENEVKCNREEA 380
Query: 444 KVLVKEVKSLRSSQTELKKELSESIKEKYEAEKLLLHEREKS-EQAEIAWRKQLEKCGLL 502
V ++ + L S EL+++L + EK E + + REK+ E + ++E
Sbjct: 381 VVHIENSEVLTSRTKELEEKLEKLEAEKEELKSEVKCNREKAVVHVENSLAAEIEVLTSR 440
Query: 503 LKQLQECSVELPYEDEDRTFLQSSSSTDAFNKLKTS-DDQIDILLAEVENLEKDAGSAAS 561
K+L+E +L E + + +A +++ S +I++L ++ LE+
Sbjct: 441 TKELEEQLEKLEAEKVELESEVKCNREEAVAQVENSLATEIEVLTCRIKQLEEKLEKL-- 498
Query: 562 NVDKTNDIKDGVICGDEMRKII 583
V+K +++K V C E+ +
Sbjct: 499 EVEK-DELKSEVKCNREVESTL 519
Score = 32.7 bits (73), Expect = 0.60
Identities = 35/160 (21%), Positives = 71/160 (43%), Gaps = 33/160 (20%)
Query: 320 EDLIVRLNQEIAAKD-----------FLTTKVKDLEVELETSKQKNKENLQQAILIERER 368
E+ +V L +++ A D L +K+ +L ++ + ++ +Q A++ ER
Sbjct: 101 ENEVVELKEKLEAADDKNRVLEDRVSHLDGALKECVRQLRQARDEQEQRIQDAVI---ER 157
Query: 369 FTQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLTKESIVQQNDVLLQDLDAAREQLEI- 427
++Q L + E K SE + + + KE+++ ++++L A E+LEI
Sbjct: 158 TQELQSSRTSLENQIFETATK--SEELSQMAESVAKENVMLRHELL-----ARCEELEIR 210
Query: 428 -----LSKQYGELEAKSKADIKVLVKEVKSLRSSQTELKK 462
LS Q E +K + D +K + + E +K
Sbjct: 211 TIERDLSTQAAETASKQQLD------SIKKVAKLEAECRK 244
Score = 29.3 bits (64), Expect = 6.7
Identities = 40/201 (19%), Positives = 88/201 (42%), Gaps = 17/201 (8%)
Query: 376 MEELRRKSLEMEMKLKSESGGNVGQ---DLTKESIVQQNDVLLQ------DLDAAREQLE 426
ME+ +R+S E +SES ++ + ++ ES ++ D +Q +++ +E+L+
Sbjct: 1 MEKRKRESSERSFG-ESESVSSLSEKDSEIQPESTMESRDDEIQSPTVSLEVETEKEELK 59
Query: 427 ILSKQYGELEAKSKADIKVLVKEVKSLRSSQTELKKELSESIKEKYEAEKLLLHEREKSE 486
K E + + A++ VK + K E++ +AE ++ +EK E
Sbjct: 60 DSMKTLAEKLSAALANVSAKDDLVK-------QHVKVAEEAVAGWEKAENEVVELKEKLE 112
Query: 487 QAEIAWRKQLEKCGLLLKQLQECSVELPYEDEDRTFLQSSSSTDAFNKLKTSDDQIDILL 546
A+ R ++ L L+EC +L +++ + + +L++S ++ +
Sbjct: 113 AADDKNRVLEDRVSHLDGALKECVRQLRQARDEQEQRIQDAVIERTQELQSSRTSLENQI 172
Query: 547 AEVENLEKDAGSAASNVDKTN 567
E ++ A +V K N
Sbjct: 173 FETATKSEELSQMAESVAKEN 193
>At5g41790 myosin heavy chain-like protein
Length = 1305
Score = 53.5 bits (127), Expect = 3e-07
Identities = 83/385 (21%), Positives = 162/385 (41%), Gaps = 68/385 (17%)
Query: 271 VSRETDIVGHTKLKSSGDAQLVLPLDQRNKLNRILSTMQRRLVTAKTDMEDLIVRLN--- 327
+ ET+I T + +AQ R + RI S +++ + T++ L +L
Sbjct: 769 IDLETEIASKTTVVEQLEAQ------NREMVARI-SELEKTMEERGTELSALTQKLEDND 821
Query: 328 -QEIAAKDFLTTKVKDLEVELETSKQKNKENLQQAILIERERFTQMQWDMEEL------- 379
Q ++ + LT ++ L EL++ + +E +Q + E +++ +E+
Sbjct: 822 KQSSSSIETLTAEIDGLRAELDSMSVQKEEVEKQMVCKSEEASVKIKRLDDEVNGLRQQV 881
Query: 380 -----RRKSLEMEMKLKSESGGNVGQDLTK------------ESIVQQNDVLLQDLDAAR 422
+R LE++++ KSE +T ESI+++ + L + +
Sbjct: 882 ASLDSQRAELEIQLEKKSEEISEYLSQITNLKEEIINKVKVHESILEEINGLSEKIKGRE 941
Query: 423 EQLEILSKQYGEL--EAKSKAD--------IKVLVKEVKSLRSSQTELKKELSESIKEKY 472
+LE L KQ EL E ++K + I V E+ +L LK EL +K
Sbjct: 942 LELETLGKQRSELDEELRTKKEENVQMHDKINVASSEIMALTELINNLKNELDSLQVQKS 1001
Query: 473 EAEKLLLHEREKSEQAEIAWRKQLEKCGLLLKQLQECSVELPYEDEDRTFLQSSSSTDAF 532
E E L EREK E++E++ Q+ L + QE + E+ + + F
Sbjct: 1002 ETEAEL--EREKQEKSELS--NQITDVQKALVE-QEAAYNTLEEEHKQI-------NELF 1049
Query: 533 NKLKTSDDQIDILLAEVENLEKDAGSAASNVDKTNDIKDGV---------ICGDEMRKII 583
+ + + +++ + E + L ++ G ++ D T + + + GDE+ ++
Sbjct: 1050 KETEATLNKVTVDYKEAQRLLEERGKEVTSRDSTIGVHEETMESLRNELEMKGDEIETLM 1109
Query: 584 SDLFLDNVRLR--KQTNRVTRHMLS 606
+ V+LR Q RVT +L+
Sbjct: 1110 EKISNIEVKLRLSNQKLRVTEQVLT 1134
Score = 47.4 bits (111), Expect = 2e-05
Identities = 73/356 (20%), Positives = 157/356 (43%), Gaps = 61/356 (17%)
Query: 299 NKLNRILSTMQRRLVTAKTDMEDLIVRLNQEIAAKDF-------LTTKVKDLEVELETSK 351
N++ +TMQ + + E V+ + + +D +T+ +LE +LE+SK
Sbjct: 108 NEIQEAQNTMQELMSESGQLKESHSVKERELFSLRDIHEIHQRDSSTRASELEAQLESSK 167
Query: 352 QK------------------NKENLQQAILIERERFT--QMQWDMEELRRKSLEMEMKLK 391
Q+ + +N++ +E+ + T ++ ++ +L+ E E +L
Sbjct: 168 QQVSDLSASLKAAEEENKAISSKNVETMNKLEQTQNTIQELMAELGKLKDSHREKESELS 227
Query: 392 S--------ESGGNVGQDLTKESIVQQNDV---LLQDLDAAREQLEILSKQYGELEAKSK 440
S + ++ +E + + L Q L+ A E+ ++LS++ EL + K
Sbjct: 228 SLVEVHETHQRDSSIHVKELEEQVESSKKLVAELNQTLNNAEEEKKVLSQKIAELSNEIK 287
Query: 441 A---DIKVLVKEVKSLRSSQTELKKELSESIKEKYEAEKLLLHEREKSEQAEIAWRKQLE 497
I+ LV E L+ S + ++L S+++ +E H+RE S + QLE
Sbjct: 288 EAQNTIQELVSESGQLKESHSVKDRDLF-SLRDIHET-----HQRESSTRVS-ELEAQLE 340
Query: 498 KCGLLLKQLQECSVELPYEDEDRTFLQSSSSTDAFNKLKTSDDQIDILLAEVENL----- 552
+++ + +V+L +E+ + SS + + +KL+ + + I L+ E+ L
Sbjct: 341 SSE---QRISDLTVDLKDAEEENKAI-SSKNLEIMDKLEQAQNTIKELMDELGELKDRHK 396
Query: 553 --EKDAGSAASNVD-KTNDIKDGVICGDEMRKIISDLFLD-NVRLRKQTNRVTRHM 604
E + S + D + D+K + +E +K++S LD + +++ + HM
Sbjct: 397 EKESELSSLVKSADQQVADMKQSLDNAEEEKKMLSQRILDISNEIQEAQKTIQEHM 452
Score = 45.4 bits (106), Expect = 9e-05
Identities = 54/220 (24%), Positives = 104/220 (46%), Gaps = 34/220 (15%)
Query: 338 TKVKDLEVELETSKQKNKENLQQAILIERERFTQMQWDMEELRRKSLEM-EMKLKSESGG 396
T +++L EL K+K KE E E + ++ R S ++ E++ ES
Sbjct: 27 TTIQELISELGEMKEKYKEK-------ESEHSSLVELHKTHERESSSQVKELEAHIESSE 79
Query: 397 NVGQDLTKESIVQQNDVLLQDLDAAREQLEILSKQYGELE---AKSKADIKVLVKEVKSL 453
+ D T Q L+ A E+ ++LS++ EL +++ ++ L+ E L
Sbjct: 80 KLVADFT------------QSLNNAEEEKKLLSQKIAELSNEIQEAQNTMQELMSESGQL 127
Query: 454 RSSQTELKKELSESIKEKYEAEKLLLHEREKSEQAEIAWRKQLEKCGLLLKQLQECSVEL 513
+ S + ++EL S+++ +E +H+R+ S +A QLE +Q+ + S L
Sbjct: 128 KESHSVKERELF-SLRDIHE-----IHQRDSSTRAS-ELEAQLESSK---QQVSDLSASL 177
Query: 514 PYEDEDRTFLQSSSSTDAFNKLKTSDDQIDILLAEVENLE 553
+E+ + SS + + NKL+ + + I L+AE+ L+
Sbjct: 178 KAAEEENKAI-SSKNVETMNKLEQTQNTIQELMAELGKLK 216
Score = 40.4 bits (93), Expect = 0.003
Identities = 59/318 (18%), Positives = 145/318 (45%), Gaps = 30/318 (9%)
Query: 308 MQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEVELETSKQKNKENLQQAILI--- 364
++ ++ ++K + +L LN K L+ K+ +L E++ ++ +E + ++ +
Sbjct: 247 LEEQVESSKKLVAELNQTLNNAEEEKKVLSQKIAELSNEIKEAQNTIQELVSESGQLKES 306
Query: 365 ----ERERFTQMQWDMEELRRKSLEM-EMKLKSESGGNVGQDLT---------KESIVQQ 410
+R+ F+ R S + E++ + ES DLT ++I +
Sbjct: 307 HSVKDRDLFSLRDIHETHQRESSTRVSELEAQLESSEQRISDLTVDLKDAEEENKAISSK 366
Query: 411 NDVLLQDLDAAREQLEILSKQYGELEAKSKADIKVLVKEVKSLRSSQTELKKELSESIKE 470
N ++ L+ A+ ++ L + GEL+ + K L VKS ++K+ L + +E
Sbjct: 367 NLEIMDKLEQAQNTIKELMDELGELKDRHKEKESELSSLVKSADQQVADMKQSLDNAEEE 426
Query: 471 KYEAEKLLL------HEREKSEQAEIAWRKQLEKCGLLLKQLQECSVELPYEDEDRTFLQ 524
K + +L E +K+ Q ++ +QL++ +K+ + + +E R
Sbjct: 427 KKMLSQRILDISNEIQEAQKTIQEHMSESEQLKE-SHGVKERELTGLRDIHETHQRE--S 483
Query: 525 SSSSTDAFNKLKTSDDQIDILLAEVENLEKDAGSAASNV-DKTNDIKDGVICGDEMRKII 583
S+ ++ +LK + ++ L A + E++ S +S + + T+++K ++++++
Sbjct: 484 STRLSELETQLKLLEQRVVDLSASLNAAEEEKKSLSSMILEITDELKQ---AQSKVQELV 540
Query: 584 SDLFLDNVRLRKQTNRVT 601
++L L ++ N ++
Sbjct: 541 TELAESKDTLTQKENELS 558
Score = 39.3 bits (90), Expect = 0.006
Identities = 67/337 (19%), Positives = 145/337 (42%), Gaps = 44/337 (13%)
Query: 297 QRNKLNRILST-------MQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEVELET 349
QR++L+ L T M ++ A +++ L +N D L + + E ELE
Sbjct: 950 QRSELDEELRTKKEENVQMHDKINVASSEIMALTELINNLKNELDSLQVQKSETEAELER 1009
Query: 350 SKQKNKE------NLQQAILIERERFTQMQWDMEELRRKSLEME-------------MKL 390
KQ+ E ++Q+A++ + + ++ + +++ E E +L
Sbjct: 1010 EKQEKSELSNQITDVQKALVEQEAAYNTLEEEHKQINELFKETEATLNKVTVDYKEAQRL 1069
Query: 391 KSESGGNV-GQDLTKESIVQQNDVLLQDLDAAREQLEILSKQYGELEAKSKADIKVLVKE 449
E G V +D T + + L +L+ +++E L ++ +E K + + L
Sbjct: 1070 LEERGKEVTSRDSTIGVHEETMESLRNELEMKGDEIETLMEKISNIEVKLRLSNQKLRVT 1129
Query: 450 VKSLRSSQTELKKELSESIKEKYEAEK--LLLHEREKSEQAEIAWRKQLEKCGLLLKQLQ 507
+ L + +KE ++ ++E+ EK + HE + EIA +K + + Q
Sbjct: 1130 EQVLTEKEEAFRKEEAKHLEEQALLEKNLTMTHETYRGMIKEIA-----DKVNITVDGFQ 1184
Query: 508 ECSVELPYED--EDRTFLQSSSST-DAFNKLKTSDDQIDILLAEV----ENLEKDAGSAA 560
S +L + ++T +++S A N + + + + + E+ E ++K G
Sbjct: 1185 SMSEKLTEKQGRYEKTVMEASKILWTATNWVIERNHEKEKMNKEIEKKDEEIKKLGGKVR 1244
Query: 561 SNVDKTNDIKDGVI-CGDEMRKIISDL--FLDNVRLR 594
+ + +K+ ++ G+E R+ I L ++D+ R R
Sbjct: 1245 EDEKEKEMMKETLMGLGEEKREAIRQLCVWIDHHRSR 1281
Score = 35.8 bits (81), Expect = 0.071
Identities = 59/311 (18%), Positives = 133/311 (41%), Gaps = 29/311 (9%)
Query: 295 LDQRNKLNRILSTMQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEVELETSKQKN 354
LD N++ T+Q + ++ E V+ + +D T ++ L + +
Sbjct: 435 LDISNEIQEAQKTIQEHMSESEQLKESHGVKERELTGLRDIHETHQRESSTRLSELETQL 494
Query: 355 KENLQQAILIERERFTQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLTKESIVQQNDVL 414
K L+Q ++ + + + L LE+ +LK + Q+L E + + D L
Sbjct: 495 KL-LEQRVVDLSASLNAAEEEKKSLSSMILEITDELKQAQ--SKVQELVTE-LAESKDTL 550
Query: 415 LQDLDAAREQLEILSKQYGELEAKSKADIKVLVKEVKSLRSSQTELKKELSESIKEKYEA 474
Q ++ E+ S + E+ K D VKE+++ S E KEL++++ E
Sbjct: 551 TQ------KENELSS--FVEVHEAHKRDSSSQVKELEARVESAEEQVKELNQNLNSSEEE 602
Query: 475 EKLLLHE---------REKSEQAEIAWRKQLEKCGLLLKQLQECSVELPYEDEDRTFLQS 525
+K+L + R +S E++ + K K + S+ +E R
Sbjct: 603 KKILSQQISEMSIKIKRAESTIQELSSESERLKGSHAEKDNELFSLRDIHETHQRELSTQ 662
Query: 526 SSSTDAFNKLKTSDDQIDILLAEVENLEKDAGSAASNVDKTNDIKDGVICGDEMRKIISD 585
+A +L++S+ ++ L ++ E+++ + ++ + +T+D + + ++ +
Sbjct: 663 LRGLEA--QLESSEHRVLELSESLKAAEEESRTMSTKISETSDEL------ERTQIMVQE 714
Query: 586 LFLDNVRLRKQ 596
L D+ +L++Q
Sbjct: 715 LTADSSKLKEQ 725
>At1g68790 putative nuclear matrix constituent protein 1 (NMCP1)
Length = 1085
Score = 51.6 bits (122), Expect = 1e-06
Identities = 62/265 (23%), Positives = 121/265 (45%), Gaps = 33/265 (12%)
Query: 320 EDLIVRLNQEIAA-----KDFLTTKVKDLEVELET----------SKQKNKENLQQAILI 364
E+LI R EI K L ++ ++ E+ELE K+ E LQ I
Sbjct: 339 ENLIEREQMEIGKLLDDQKAVLDSRRREFEMELEQMRRSLDEELEGKKAEIEQLQVEISH 398
Query: 365 ERERFTQMQWDMEE----LRRKSLEMEMKLKSESGGNVGQDLTKESIVQQNDVLLQDLDA 420
+ E+ + + +E+ +++K +++ +LK+ ++ + +N+ LL+D +
Sbjct: 399 KEEKLAKREAALEKKEEGVKKKEKDLDARLKTVKEKEKALKAEEKKLHMENERLLEDKEC 458
Query: 421 AREQLEILSKQYGELEAKSKADIKVLVKEVKSLRSSQTE------LKKELSESIKEKYEA 474
R+ + + ++ G K ++ I+ +E +SLR ++ E L+ EL + I + +
Sbjct: 459 LRKLKDEI-EEIGTETTKQESRIR---EEHESLRITKEERVEFLRLQSELKQQIDKVKQE 514
Query: 475 EKLLLHEREKSEQAEIAWRKQLEKCGLLLKQLQECSVELPYEDEDRTFLQSSSSTDAFNK 534
E+LLL ERE+ +Q + + K+ E + E+ E+E LQ S +
Sbjct: 515 EELLLKEREELKQDKERFEKEWEALDKKRANITREQNEVAEENEKLRNLQISEKHRLKRE 574
Query: 535 LKTSDD----QIDILLAEVENLEKD 555
TS D ++D + + E+ E D
Sbjct: 575 EMTSRDNLKRELDGVKMQKESFEAD 599
Score = 40.4 bits (93), Expect = 0.003
Identities = 57/230 (24%), Positives = 106/230 (45%), Gaps = 37/230 (16%)
Query: 351 KQKNKENLQQAILIERERFTQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLT------- 403
K+K ENLQQ I + + T+ + E ++ K ++ +K K D+
Sbjct: 281 KEKILENLQQKISVAKSELTEKE---ESIKIKLNDISLKEKDFEAMKAKVDIKEKELHEF 337
Query: 404 KESIVQQNDV----LLQD----LDAAREQLEILSKQY-----GELEAKSKADIKVLVKEV 450
+E+++++ + LL D LD+ R + E+ +Q ELE K KA+I+ L E+
Sbjct: 338 EENLIEREQMEIGKLLDDQKAVLDSRRREFEMELEQMRRSLDEELEGK-KAEIEQLQVEI 396
Query: 451 K------SLRSSQTELKKELSESIKEKYEAEKLLLHEREKSEQAEIAWRKQLEKCGLLLK 504
+ R + E K+E + ++ +A + E+EK+ +AE K+L L
Sbjct: 397 SHKEEKLAKREAALEKKEEGVKKKEKDLDARLKTVKEKEKALKAE---EKKLHMENERLL 453
Query: 505 QLQECSVELPYEDED---RTFLQSSSSTDAFNKLK-TSDDQIDILLAEVE 550
+ +EC +L E E+ T Q S + L+ T +++++ L + E
Sbjct: 454 EDKECLRKLKDEIEEIGTETTKQESRIREEHESLRITKEERVEFLRLQSE 503
Score = 30.8 bits (68), Expect = 2.3
Identities = 31/125 (24%), Positives = 60/125 (47%), Gaps = 11/125 (8%)
Query: 364 IERERFTQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLTKESIVQQNDVLLQDLDAARE 423
I E+F+ M EL RK E+E + K ++ S+V + + RE
Sbjct: 189 IAEEKFSVMNRKSSELERKLKEVETREKVHQREHL-------SLVTEREAHEAVFYKQRE 241
Query: 424 QLEILSKQYGELEAKSKADIKVLV--KEVKSLRSSQT-ELKKELSESIKEKYEAEKLLLH 480
L+ K+ LE +++K + +E + + + +T E K+++ E++++K K L
Sbjct: 242 DLQEWEKKL-TLEEDRLSEVKRSINHREERVMENERTIEKKEKILENLQQKISVAKSELT 300
Query: 481 EREKS 485
E+E+S
Sbjct: 301 EKEES 305
>At5g16730 putative protein
Length = 853
Score = 51.2 bits (121), Expect = 2e-06
Identities = 58/285 (20%), Positives = 125/285 (43%), Gaps = 44/285 (15%)
Query: 302 NRILSTMQRRLVTAKTDMED-------------LIVRLNQEIAAKDFLTTKVKDLEVELE 348
N +++ ++ +V K D+E ++ +LN ++ A + L E +
Sbjct: 264 NEMVAKLEDEIVVLKRDLESARGFEAEVKEKEMIVEKLNVDLEAAKMAESNAHSLSNEWQ 323
Query: 349 TSKQKNKENLQQAILIERERFTQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLTKESIV 408
+ ++ +E L++A +ER SLE MK S + T+ + +
Sbjct: 324 SKAKELEEQLEEANKLERSASV------------SLESVMKQLEGSNDKLHDTETEITDL 371
Query: 409 QQNDVLLQDLDA-AREQLEILSKQYGELE---AKSKADIKVLVKEVKSLRSSQTE-LKKE 463
++ V L+ A +E LE+ ++ G +E +K++ +++ L E+++++ + LKKE
Sbjct: 372 KERIVTLETTVAKQKEDLEVSEQRLGSVEEEVSKNEKEVEKLKSELETVKEEKNRALKKE 431
Query: 464 --LSESIKEKYEAEKLLLHEREKSEQAEIAWRKQLEKCGLLL------------KQLQEC 509
+ ++ E + LL + E S++ E +K +E L K L +
Sbjct: 432 QDATSRVQRLSEEKSKLLSDLESSKEEEEKSKKAMESLASALHEVSSEGRELKEKLLSQG 491
Query: 510 SVELPYEDEDRTFLQSSSSTDAFNKLKTSDDQIDILLAEVENLEK 554
E + +D + +++ N L + +ID+L++ VE +K
Sbjct: 492 DHEYETQIDDLKLVIKATNEKYENMLDEARHEIDVLVSAVEQTKK 536
Score = 39.7 bits (91), Expect = 0.005
Identities = 69/339 (20%), Positives = 152/339 (44%), Gaps = 35/339 (10%)
Query: 265 LPGSASVSRETDIVGHTKLKSSGDAQLVLPLDQRNKLNRILSTMQRRLVTAKTDMEDLIV 324
L SASVS E+ + +L+ S D + + RI+ T++ + K D+E
Sbjct: 339 LERSASVSLESVM---KQLEGSNDKLHDTETEITDLKERIV-TLETTVAKQKEDLEVSEQ 394
Query: 325 RLNQEIAAKDFLTTKVKDLEVELETSKQKNKENLQQ----AILIER--ERFTQMQWDMEE 378
RL +V+ L+ ELET K++ L++ ++R E +++ D+E
Sbjct: 395 RLGSVEEEVSKNEKEVEKLKSELETVKEEKNRALKKEQDATSRVQRLSEEKSKLLSDLES 454
Query: 379 LRRKSLEMEMKLKSESGG-----NVGQDLTKESIVQQNDVLLQDLDAAREQLEILSKQYG 433
+ + + + ++S + + G++L ++ + Q + +D + ++ +++Y
Sbjct: 455 SKEEEEKSKKAMESLASALHEVSSEGRELKEKLLSQGDHEYETQIDDLKLVIKATNEKYE 514
Query: 434 ELEAKSKADIKVLVKEVKSLRSSQTELKKELSESIKEKYEAEKLLLHEREKSEQAEIAWR 493
+ +++ +I VLV V+ + KK S K+ E L++ +K E+ +
Sbjct: 515 NMLDEARHEIDVLVSAVE-------QTKKHFESSKKDWEMKEANLVNYVKKMEEDVASMG 567
Query: 494 KQLEKCGLLLKQLQECSVELPYEDEDRTFLQSSSSTDAFNKLKTSDDQIDILLAEVENLE 553
K++ + LLK+ + E+ D + + + + D+ LK +++I L + +
Sbjct: 568 KEMNRLDNLLKRTE--------EEADAAWKKEAQTKDS---LKEVEEEIVYLQETLGEAK 616
Query: 554 KDAGSAASN-VDKTNDIKDGVICGDEMRKIISDLFLDNV 591
++ N +DK + ++ VI +E K D+ L +
Sbjct: 617 AESMKLKENLLDKETEFQN-VIHENEDLKAKEDVSLKKI 654
>At3g10170 hypothetical protein
Length = 634
Score = 50.8 bits (120), Expect = 2e-06
Identities = 76/361 (21%), Positives = 144/361 (39%), Gaps = 65/361 (18%)
Query: 272 SRETDIVGHTKLKSSGDAQLVLPLDQRNKLNRILSTMQRRLVTAKTDMEDLIVRLNQEIA 331
S +D++ H + L + L + +++ KT ++D +L +
Sbjct: 188 SERSDLLSHIECLEKDIGSL-----SSSSLAKEKENLRKDFEKTKTKLKDTESKLKNSMQ 242
Query: 332 AKDFLTTKVKDLEVEL------------ETSKQKNKENLQQ-AILIERERFTQMQWDMEE 378
K L + E EL + SKQ++ ++ ++L+ER +Q + ++
Sbjct: 243 DKTKLEAEKASAERELKRLHSQKALLERDISKQESFAGKRRDSLLVERSANQSLQEEFKQ 302
Query: 379 LRRKSLEMEMKLKSESGGNVGQDLTKESIVQQND-------VLLQDLDAAREQLEILSKQ 431
L + EME + S + KE + +ND L + L+ + +LE L
Sbjct: 303 LEVLAFEMETTIASLEEELAAERGEKEEALCRNDGLGSEITDLTEKLEHSNTKLEHLQND 362
Query: 432 YGELEAK---SKADIKVLVKEVKSLRSSQTEL---------------------KKELSES 467
EL+ + S +D + L VK L + EL +K L+E+
Sbjct: 363 VTELKTRLEVSSSDQQQLETNVKQLLEEKEELAMHLANSLLEMEEEKAIWSSKEKALTEA 422
Query: 468 IKEKYEAEKLLLHEREKSEQAEIAWRKQLEKCGL---LLKQLQECSVELPYEDEDRTFLQ 524
++EK K + E E +E +K+LE C L L CS E +D++ + +
Sbjct: 423 VEEKIRLYKNIQIESLSKEMSE--EKKELESCRLECVTLADRLRCSEENAKQDKESSLEK 480
Query: 525 SSSSTDAFNKLKTSD-----------DQIDILLAEVENLEKDAGSAASNVDKTNDIKDGV 573
S ++L+++D IDIL +EV++ K + + +D + G+
Sbjct: 481 SLEIDRLGDELRSADAVSKQSQEVLKSDIDILKSEVQHACKMSDTFQREMDYVTSERQGL 540
Query: 574 I 574
+
Sbjct: 541 L 541
>At1g79280 hypothetical protein
Length = 2111
Score = 50.8 bits (120), Expect = 2e-06
Identities = 65/280 (23%), Positives = 136/280 (48%), Gaps = 38/280 (13%)
Query: 288 DAQLVLPLDQRNKLNRILSTMQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEVEL 347
D + VL L+++NK+ S +++ L K D+ + RL+ A+ + ++ + EL
Sbjct: 1400 DCKKVL-LEKQNKI----SLLEKELTNCKKDLSEREKRLDDAQQAQATMQSEFNKQKQEL 1454
Query: 348 ETSKQKNKENLQQAILIERERFTQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLTKESI 407
E +K+ + + + + ++ + + +EL +++ + +L+ E+ G+ T +++
Sbjct: 1455 EKNKK-----IHYTLNMTKRKYEK---EKDELSKQNQSLAKQLE-EAKEEAGKRTTTDAV 1505
Query: 408 VQQNDVLLQDLDAAREQLEILSKQYGELE---AKSKADIKVLVKEVKSLRSSQTELKKEL 464
V+Q+ +++ + ++++IL K +L+ K D+K +E+ RS + ++KE+
Sbjct: 1506 VEQS---VKEREEKEKRIQILDKYVHQLKDEVRKKTEDLKKKDEELTKERSERKSVEKEV 1562
Query: 465 SESIKEKYEAEKLLLHEREKSEQAEIAWRKQLEKCGLLLKQLQECSVELPYEDEDRTFLQ 524
+S+ K + EK + E E+A +LE+ L L E +L + D +
Sbjct: 1563 GDSL-TKIKKEKTKVDE-------ELA---KLERYQTALTHLSEELEKLKHADGN----- 1606
Query: 525 SSSSTDAFNKLKTS--DDQIDILLAEVENLEKDAGSAASN 562
T A L S +DQ ++ VE E+ A S ASN
Sbjct: 1607 LPEGTSAVQVLSGSILNDQAAAYVSAVEYFERVARSIASN 1646
Score = 39.7 bits (91), Expect = 0.005
Identities = 78/373 (20%), Positives = 144/373 (37%), Gaps = 75/373 (20%)
Query: 282 KLKSSGDAQLVLPLDQRNKLNRILSTMQRRLVTAKTDMEDLIVRLNQEIAA--------- 332
K + S +A+LV ++ ++L +L TA ED ++ + EIA+
Sbjct: 1018 KRQRSLEAELVSLRERVSELENDCIQKSEQLATAAAGKEDALLSASAEIASLREENLVKK 1077
Query: 333 ------KDFLTTKVKDLEVELETSKQKNKENLQQAILIER---------ERFTQMQWDME 377
++T DLE E E + + +Q IL+ + +Q +
Sbjct: 1078 SQIEAMNIQMSTLKNDLETEHEKWRVAQRNYERQVILLSETIQELTKTSQALAALQEEAS 1137
Query: 378 ELRR----KSLE--------MEMKLKSESGGNVGQDLTKESIVQQNDVLLQDLDAAREQL 425
ELR+ + +E E KL E N+ + E + +QN +L L+A
Sbjct: 1138 ELRKLADARGIENSELNAKWSEEKLMLEQQKNLAEKKYHE-LNEQNKLLHSRLEAKHLNS 1196
Query: 426 EILSKQYGELEA-----------------------KSKADIKVLVKEVKSLR-SSQTELK 461
+ + G + + K A+ ++ + + LR SQ+ LK
Sbjct: 1197 AEKNSRSGTISSGSTDSDHLEDSGLQRVVHYLRRTKEIAETEISLMRQEKLRLQSQSALK 1256
Query: 462 KELSESIKEKYEAEK------LLLHEREKSEQAEIAWRKQLEKCGLLLKQLQECSVELPY 515
++ES + AE+ LL + KS Q +++ L + + L++ + + E
Sbjct: 1257 --MAESARGSLTAERASTRASLLTDDGIKSLQLQVSEMNLLRESNMQLREENKHNFEKCQ 1314
Query: 516 EDEDRTFLQSSSSTDAFNKLKTSDDQIDILLAEVENLEKDAGSAASNVDKTN------DI 569
E + S + N LKT ++D+ + E+E L + VD+ DI
Sbjct: 1315 EMREVAQKARMESENFENLLKTKQTELDLCMKEMEKLRMETDLHKKRVDELRETYRNIDI 1374
Query: 570 KDGVICGDEMRKI 582
D DE+R++
Sbjct: 1375 ADYNRLKDEVRQL 1387
Score = 36.2 bits (82), Expect = 0.055
Identities = 53/225 (23%), Positives = 97/225 (42%), Gaps = 25/225 (11%)
Query: 295 LDQRNKLNRILSTMQRRLVTAKTDM---EDLIVRLNQEIAAKDFLTTKVKDLEVELETS- 350
LD+ KL S + RL A ++ + + RL+QE K+ K L+ EL
Sbjct: 149 LDKIVKLTDTSSEKEARLAEATAELARSQAMCSRLSQE---KELTERHAKWLDEELTAKV 205
Query: 351 --------KQKNKENLQQAILIERER-----FTQMQWDMEELRRKSLEMEMKLKSESGGN 397
+ + E+ A L++ E+ + + W E LR ++ + S
Sbjct: 206 DSYAELRRRHSDLESEMSAKLVDVEKNYIECSSSLNWHKERLRELETKIGSLQEDLSSCK 265
Query: 398 VGQDLTKESIVQQNDVLLQDLDAAREQLEILSKQYGELEAKSKADIKVLVKEVKSLRSSQ 457
T+E + + +D +E E S++ GELE KA ++ + +V+S S +
Sbjct: 266 DAATTTEEQYTAELFTANKLVDLYKESSEEWSRKAGELEGVIKA-LEARLSQVES--SYK 322
Query: 458 TELKKELSESIKEKYEAEKLLLHEREKSEQAEIAWRKQLEKCGLL 502
L KE+ S K+ E E L ++ + +AEI ++ ++ L+
Sbjct: 323 ERLDKEV--STKQLLEKENGDLKQKLEKCEAEIEKTRKTDELNLI 365
>At1g67230 unknown protein
Length = 1132
Score = 50.8 bits (120), Expect = 2e-06
Identities = 75/329 (22%), Positives = 139/329 (41%), Gaps = 40/329 (12%)
Query: 282 KLKSSGDAQLVLPLDQRNKLNRILSTMQRRLVTAKTDMEDLI--VRLNQEIAAKDFLTTK 339
K+ AQL ++++L R+ + + +T++++ I R QE+ K+ K
Sbjct: 457 KVSGENQAQLSEINKEKDEL-RVTEEERSEYLRLQTELKEQIEKCRSQQELLQKEAEDLK 515
Query: 340 V------KDLEVELETSKQK----------NKENLQQAILIERERFTQMQWDMEELRRKS 383
K+ E EL+ K K KE L++ I +E ER + + E +
Sbjct: 516 AQRESFEKEWE-ELDERKAKIGNELKNITDQKEKLERHIHLEEERLKKEKQAANENMERE 574
Query: 384 LEMEMKLKSESGGNVGQD--LTKESIVQQNDVLLQDLDAAREQLE-----ILSKQYGELE 436
LE K+ + + + + + LL D++ + +LE IL ++ EL+
Sbjct: 575 LETLEVAKASFAETMEYERSMLSKKAESERSQLLHDIEMRKRKLESDMQTILEEKERELQ 634
Query: 437 AKSKADIKVLVKEVKSLRSSQTELKKELSESIKEKYEAEKLLLH---EREKSEQAEIAWR 493
AK K + KE+ ++ + ++E+ + E+ EK L + E+ + R
Sbjct: 635 AKKKLFEEEREKELSNINYLRDVARREMMDMQNERQRIEKEKLEVDSSKNHLEEQQTEIR 694
Query: 494 KQLEKCGLLLKQLQECSVELPYEDEDRTFLQSSSSTDAFNKLKTSDDQIDILLAEVENLE 553
K ++ L K+L+E + + E FL S S N + + +++L E++NLE
Sbjct: 695 KDVDDLVALTKKLKEQREQ--FISERSRFLSSMESNR--NCSRCGELLSELVLPEIDNLE 750
Query: 554 KDAGSAASNVDKTNDIKDGVICGDEMRKI 582
N+ K +I D EMR I
Sbjct: 751 ------MPNMSKLANILDNEAPRQEMRDI 773
Score = 46.6 bits (109), Expect = 4e-05
Identities = 66/280 (23%), Positives = 128/280 (45%), Gaps = 38/280 (13%)
Query: 319 MEDLIVRLNQEIAAKDFLTTKVKDLEVELETSKQKNKENLQQAILIERERF--TQMQWDM 376
++DL +R + K + TK ++L+ E + + K +QQ + + + TQ ++++
Sbjct: 299 IKDLALREQETDVLKKSIETKARELQALQEKLEAREKMAVQQLVDEHQAKLDSTQREFEL 358
Query: 377 E-ELRRKSLEMEMKLKSESGGNVGQDLT--KESIVQQNDVLLQDLDAAREQ---LEILSK 430
E E +RKS++ +K K + +E + ++ L + L+ +E+ ++ K
Sbjct: 359 EMEQKRKSIDDSLKSKVAEVEKREAEWKHMEEKVAKREQALDRKLEKHKEKENDFDLRLK 418
Query: 431 QYGELEAKSKADIKVLVKEVKSLRSSQT--------------ELKKELSESIKEKYEAEK 476
E K++ K L E K L + E + +LSE KEK ++
Sbjct: 419 GISGREKALKSEEKALETEKKKLLEDKEIILNLKALVEKVSGENQAQLSEINKEK---DE 475
Query: 477 LLLHEREKSE--QAEIAWRKQLEKCGLLLKQLQECSVELPYEDEDRTFLQSSSSTD---- 530
L + E E+SE + + ++Q+EKC + LQ+ + +L + E +F + D
Sbjct: 476 LRVTEEERSEYLRLQTELKEQIEKCRSQQELLQKEAEDLKAQRE--SFEKEWEELDERKA 533
Query: 531 -AFNKLKTSDDQIDIL----LAEVENLEKDAGSAASNVDK 565
N+LK DQ + L E E L+K+ +A N+++
Sbjct: 534 KIGNELKNITDQKEKLERHIHLEEERLKKEKQAANENMER 573
Score = 38.1 bits (87), Expect = 0.014
Identities = 46/218 (21%), Positives = 99/218 (45%), Gaps = 44/218 (20%)
Query: 352 QKNKENLQQAILIERERFTQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLTKESIVQQN 411
+K +E L++A+ IE++ ++ ++ELR ++ E++ T +S + +
Sbjct: 113 EKREEGLRKALGIEKQCALDLEKALKELRAENAEIK--------------FTADSKLTEA 158
Query: 412 DVLLQDLD----AAREQLEILSKQYGELEAKSKADIKVLVKEVKSLRSSQTELKKELSES 467
+ L++ ++ +L + + E+ KS +D++ KEV++ SS L++E
Sbjct: 159 NALVRSVEEKSLEVEAKLRAVDAKLAEVSRKS-SDVERKAKEVEARESS---LQRERFSY 214
Query: 468 IKEKYEAEKLLLHEREKSEQAEIAWRKQLE-------KCGLLLKQLQECSVELPYEDEDR 520
I E+ E L +RE + W ++L+ K +++KQ ++ + + D+
Sbjct: 215 IAEREADEATLSKQREDLRE----WERKLQEGEERVAKSQMIVKQREDRA-----NESDK 265
Query: 521 TFLQSSSSTDAFNKLKTSDDQIDILLAEVENLEKDAGS 558
Q +L+ + +ID V+ LE D S
Sbjct: 266 IIKQKG------KELEEAQKKIDAANLAVKKLEDDVSS 297
>At1g12150 hypothetical protein
Length = 548
Score = 49.7 bits (117), Expect = 5e-06
Identities = 65/311 (20%), Positives = 140/311 (44%), Gaps = 42/311 (13%)
Query: 281 TKLKSSGDAQLVLPLDQR--NKLNRILSTMQRRL----VTAKTDMEDLIVRLNQEIAAKD 334
T L + +AQ L ++ N+L++ +S M+ + + A ++++ + ++ ++
Sbjct: 184 TALNQAAEAQRALQVNSAKVNELSKEISDMKDAIHQLKLAAAQNLQEHANIVKEKDDLRE 243
Query: 335 FLTTKVKDLEVELETSKQKNKENLQQAILIERERFTQMQWDMEELRRKSLEMEMKLKSES 394
T V++ E +L +++ + L + + + + ++E LR EMK ES
Sbjct: 244 CYRTAVEEAEKKLLVLRKEYEPELSRTL---EAKLLETTSEIEVLRE-----EMKKAHES 295
Query: 395 GGNVGQDLTKE-------------------SIVQQNDVLLQDLDAAREQLEILSKQYGEL 435
N + +T E S+V + L+DL RE+ E+ K+ L
Sbjct: 296 EMNTVKIITNELNEATMRLQEAADDECSLRSLVNSLRMELEDL--RREREELQQKEAERL 353
Query: 436 EAKSKADIKVLVKEVKSLRSSQTELKKELSESIKEKYEAEKLLLHEREKSEQAEIAWRKQ 495
E + ++ L +E L +TE + +E+ + E L ++++E A IA +
Sbjct: 354 EIEETKKLEALKQESLKLEQMKTEAIEARNEAANMNRKIESL----KKETEAAMIAAEEA 409
Query: 496 LEKCGLLLKQLQEC-SVELPYEDEDRTFLQSSSSTDAFNKLKTSDDQIDILLAEVENLEK 554
++ L++++++E S E +E + Q S ++S +I I + E E+L++
Sbjct: 410 EKRLELVIREVEEAKSAEEKVREEMKMISQKQESKK--QDEESSGSKIKITIQEFESLKR 467
Query: 555 DAGSAASNVDK 565
AG + ++K
Sbjct: 468 GAGETEAAIEK 478
>At1g64330 unknown protein
Length = 555
Score = 49.3 bits (116), Expect = 6e-06
Identities = 47/186 (25%), Positives = 85/186 (45%), Gaps = 30/186 (16%)
Query: 314 TAKTDMEDLIVRLNQEIAAK----DFLTTKVKDLEVELETSKQKNKENLQQAILIERERF 369
+A D+E+ + L E+ K + L K+ ++EV+L S QK + + +
Sbjct: 335 SAIVDLEETVESLRNEVERKGDEIESLMEKMSNIEVKLRLSNQK--------LRVTEQVL 386
Query: 370 TQMQWDMEELRRKSLEMEMKLKSE--SGGNVGQDLTKESIVQQNDVLLQDLDAAREQLEI 427
T+ + +++ + K LE + L+ + + + L KE + + +L + E+LE
Sbjct: 387 TEKEGELKRIEAKHLEEQALLEEKIATTHETYRGLIKEISERVDSTILNRFQSLSEKLEE 446
Query: 428 LSKQYGELEAKSKADIKVLVKEVKSLRSSQ---TELKKELSESIKEKYEAEKLL---LHE 481
K Y K +V+ K L +++ E+KKE E KEK E EK L + E
Sbjct: 447 KHKSYE----------KTVVEATKMLLTAKKCVVEMKKEKDEMAKEKEEVEKKLEGQVRE 496
Query: 482 REKSEQ 487
EK ++
Sbjct: 497 EEKEKE 502
Score = 38.5 bits (88), Expect = 0.011
Identities = 51/251 (20%), Positives = 109/251 (43%), Gaps = 38/251 (15%)
Query: 297 QRNKLNRILSTMQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLE---VELETSKQK 353
Q+N+ L ++ + D+ L ++ AA + L+ + K + E E + +K
Sbjct: 238 QKNETEAELEREKQEKPALLNQINDVQKALLEQEAAYNTLSQEHKQINGLFEEREATIKK 297
Query: 354 NKENLQQAILIERERFTQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLTKESIVQQNDV 413
++ +QA RE + MEE R+ E + S V + T ES+ + +
Sbjct: 298 LTDDYKQA----REMLEEYMSKMEETERRMQETGKDVASRESAIVDLEETVESLRNEVER 353
Query: 414 LLQDLDAAREQL------------------EILSKQYGEL---EAKSKADIKVLVKEVKS 452
++++ E++ ++L+++ GEL EAK + +L +++ +
Sbjct: 354 KGDEIESLMEKMSNIEVKLRLSNQKLRVTEQVLTEKEGELKRIEAKHLEEQALLEEKIAT 413
Query: 453 LRSSQTELKKELSE----SIKEKYEAEKLLLHEREKSEQAEIAWRKQLEKCGLLLKQLQE 508
+ L KE+SE +I ++++ L E+ KS + K + + +L ++
Sbjct: 414 THETYRGLIKEISERVDSTILNRFQSLSEKLEEKHKS------YEKTVVEATKMLLTAKK 467
Query: 509 CSVELPYEDED 519
C VE+ E ++
Sbjct: 468 CVVEMKKEKDE 478
>At4g38065 hypothetical protein
Length = 1050
Score = 48.1 bits (113), Expect = 1e-05
Identities = 77/352 (21%), Positives = 151/352 (42%), Gaps = 47/352 (13%)
Query: 283 LKSSGDAQLVLP---LDQRNKLNRILSTMQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTK 339
L+ S QL+L +D N R L+ + L A +++ D + Q +
Sbjct: 673 LEESTKTQLLLQEKVVDVENDSKRKLADVSEALEIANSELSDKTSEVFQIEFQLWVWKSI 732
Query: 340 VKDLEVELETSKQKNKENLQQAILIERERFTQMQWDMEELRRKSLEMEMKLKSESGGNVG 399
K L+ ELE ++ K +A L+E+ E ++++ E+ KLK S
Sbjct: 733 AKRLKAELEQNQNLRKR--VEASLLEQVGVG------EAIKQEKNELVHKLKVISHARSS 784
Query: 400 QDLTKESIVQQNDVLLQDL---------DAAREQLE--ILSKQYGELEAKSKADIKVLVK 448
KES+++ D +L+ L D+ R +LE +L+ GE E +++ +I L +
Sbjct: 785 DSEKKESLMRDKDEMLESLQREVELLEQDSLRRELEDVVLAHMIGERELQNEREICALQQ 844
Query: 449 EVKSLRSSQTELK---KELSESIKEKYEAEKLLLHEREKSEQAEIAWRKQLEKCGLLLKQ 505
+ + L + EL+ K +S +++K +L EK +I + E +++ +
Sbjct: 845 KDQDLCEVKHELEGSLKSVSLLLQQKQNEVNMLRKTWEKLTARQILTAVETESKKMMIIE 904
Query: 506 LQ----ECSVELPYEDED-RTFLQSSSSTDAFNKLKTSDDQIDILLAEVENLEKDAGSAA 560
L+ S +L +E F Q ++ + A +L+T ++ + +++ +
Sbjct: 905 LEGEISSLSQKLETSNESVSCFRQEATKSRA--ELETKQTELKEVTTQMQEKLR-----T 957
Query: 561 SNVDKTNDIKDGVICGDEMRKIIS----------DLFLDNVRLRKQTNRVTR 602
S +KT +K+ E R ++S L+ + +L K RVT+
Sbjct: 958 SEAEKTELVKEVASLSTEKRNLLSFISEMEDGMLKLYDGDTKLMKTLERVTQ 1009
Score = 41.2 bits (95), Expect = 0.002
Identities = 62/284 (21%), Positives = 118/284 (40%), Gaps = 41/284 (14%)
Query: 293 LPLDQRNKLNRILSTMQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEVELETSKQ 352
L +D RNK L K + ++ + + + ++++ E+ K+
Sbjct: 18 LRIDYRNKTEL--------LENLKKVQNEQLIEIREARLVNEKHGFEIEEKSREIAELKR 69
Query: 353 KNKENLQQAILIERERFTQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLTKES-----I 407
N+E Q L E++ + D+ + R + E + + E N+ L + S +
Sbjct: 70 ANEE--LQRCLREKDSVVKRVNDVNDKLRANGEDKYREFEEEKRNMMSGLDEASEKNIDL 127
Query: 408 VQQNDVLLQDLD--------AAREQLEILSKQYGELEAKSKADIKVLVKEVKSLRSSQTE 459
Q+N+V +++ A +++E G E + + D+ V ++E KS + +
Sbjct: 128 EQKNNVYRAEIEGLKGLLAVAETKRIEAEKTVKGMKEMRGRDDVVVKMEEEKSQVEEKLK 187
Query: 460 LKKELSESIKEKYEAEKLLLH------EREKSEQAEIAWRKQLEKCGL------LLKQLQ 507
KKE + ++E YE K L E EKS+ + + Q + + L K+LQ
Sbjct: 188 WKKEQFKHLEEAYEKLKNLFKDSKKEWEEEKSKLLDEIYSLQTKLDSVTRISEDLQKKLQ 247
Query: 508 ECSVELPYEDEDRTFLQSSSS------TDAFNKLKTSDDQIDIL 545
C+ L E+ R L+ S DAF + + + Q+D L
Sbjct: 248 MCNGALTQEETRRKHLEIQVSEFKAKYEDAFAECQDARTQLDDL 291
>At5g66030 Golgi-localized protein GRIP
Length = 788
Score = 47.4 bits (111), Expect = 2e-05
Identities = 50/234 (21%), Positives = 106/234 (44%), Gaps = 18/234 (7%)
Query: 280 HTKLKSSGDAQLVLPLDQRNKLNRILSTMQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTK 339
H K +AQ+V L +R+K +S++Q L ++ + ++ E A
Sbjct: 289 HQKNLEGLEAQVVDALSERDKAAETISSLQVLLAEKESKIAEMEAAATGEAARLRAAAET 348
Query: 340 VKDLEVELETSKQKNKENLQQAILIERERFTQMQWDMEELRRKSLEMEM-KLKSESGGNV 398
+K L++ +K KE + + + + ++ E E+E+ K++S+ G +
Sbjct: 349 LKGELAHLKSENEKEKETWEASCDALKSKL-----EIAESNYLQAEIEVAKMRSQLGSEM 403
Query: 399 GQDLTKESIVQQNDVLLQDLDAAREQLEILSKQYGELEAKSKADIKVLVKEVKSLRSSQT 458
+ I+ D +L ARE++ L ++ + ++ A ++ K+++ + +
Sbjct: 404 SM---QTQILSTKDA---ELKGAREEINRLQSEFSSYKIRAHALLQ--KKDMELAAAKDS 455
Query: 459 ELKKELSESIKEKYEAEKLLLHEREKSEQ----AEIAWRKQLEKCGLLLKQLQE 508
E K L E++KE + L+ ER++++Q A + K+LE+ LK E
Sbjct: 456 EQIKSLEEALKEAEKEVYLVSAERDRAQQDLQSALASLEKELEERAGALKDASE 509
Score = 43.9 bits (102), Expect = 3e-04
Identities = 72/324 (22%), Positives = 135/324 (41%), Gaps = 44/324 (13%)
Query: 270 SVSRETDIVGHTKL-------KSSGDAQLVLPLDQRNKLNRILSTMQRRLVTAKTDMEDL 322
S +E+D+VG + K DA + ++L ++++ ++ ++ E L
Sbjct: 2 SEDKESDVVGEEEESHVIKEDKELNDASNETLTENGDQLLQMIAELRLENDFLRSQFEGL 61
Query: 323 IVRLNQEIAAKDFLTTKVKDLEVELETSKQKNKE--NLQQAILIERERFTQMQWDMEELR 380
++ A+ K + +E + KQ ++ +L + I +E++ + +E LR
Sbjct: 62 -----KDEVAQGRSLQKAEQVEADSAQLKQLQEQVASLSREIDVEKQTRVAAEQALEHLR 116
Query: 381 RKSLEMEMKLKSESG--GNVGQDLTKESIVQQNDVLLQDLDAAREQLEILSKQYGELEAK 438
E + K + S V Q L +E +++ D DLDA +L +KQ + K
Sbjct: 117 EAYSEADAKSQEYSSKFSQVEQKLDQE--IKERDEKYADLDAKFTRLHKRAKQRIQEIQK 174
Query: 439 SKADIKVLVKEV----KSLRSSQTELKKELSESIKEKYEAEKLLLHEREKSEQAEIAWRK 494
K D+ +EV + S + +++EL + ++ EA K + ER++ A R
Sbjct: 175 EKDDLDARFREVNETAERASSQHSSMQQELERTRQQANEALKAMDAERQQLRSANNKLRD 234
Query: 495 QLEKCGLLLKQLQECSVELPYEDEDRTFLQSSSSTDAF-----NKLKTSDDQIDILLAE- 548
+E+ L LQ P E++ T QS D +L+ +++ I + E
Sbjct: 235 TIEE---LRGSLQ------PKENKIETLQQSLLDKDQILEDLKKQLQAVEERKQIAVTEL 285
Query: 549 -------VENLEKDAGSAASNVDK 565
+E LE A S DK
Sbjct: 286 SAKHQKNLEGLEAQVVDALSERDK 309
>At1g63300 hypothetical protein
Length = 1029
Score = 47.4 bits (111), Expect = 2e-05
Identities = 62/261 (23%), Positives = 121/261 (45%), Gaps = 31/261 (11%)
Query: 327 NQEIAAKDFLTTKVKDLEVELETSKQKNKENLQQAIL------IERERFTQMQWDM-EEL 379
N +A K L + K L ++++ N++ +A+ +++ + +M D +EL
Sbjct: 616 NASVAGK--LQDEFKRLSEQMDSMFTSNEKMAMKAMTEANELRMQKRQLEEMIKDANDEL 673
Query: 380 RRKSLEMEMKLKSESGGNVGQDLTKESIVQQNDVLLQDLDAAREQLEILSKQYGELEAKS 439
R E E KL S + L+ ++ Q + +L++LD +++ + ++ A
Sbjct: 674 RANQAEYEAKLHELS-----EKLSFKT--SQMERMLENLDEKSNEIDNQKRHEEDVTANL 726
Query: 440 KADIKVLVKEVKSLRSSQTE--LKKELSESIKEKYEAEKLLLHEREKSEQAEIAWRKQLE 497
+IK+L +E+++L+ +Q L+ E +E+++ E K + E E S Q E + +LE
Sbjct: 727 NQEIKILKEEIENLKKNQDSLMLQAEQAENLRVDLEKTKKSVMEAEASLQRENMKKIELE 786
Query: 498 -KCGLLLKQLQECSVELPY----EDEDRT---FLQSSSST--DAFNKLKTSDDQIDILLA 547
K L+ K+ + + EL +DE T LQ+ T + LK S + D+
Sbjct: 787 SKISLMRKESESLAAELQVIKLAKDEKETAISLLQTELETVRSQCDDLKHSLSENDL--- 843
Query: 548 EVENLEKDAGSAASNVDKTND 568
E+E +K S + K +
Sbjct: 844 EMEKHKKQVAHVKSELKKKEE 864
Score = 39.7 bits (91), Expect = 0.005
Identities = 60/314 (19%), Positives = 133/314 (42%), Gaps = 45/314 (14%)
Query: 275 TDIVGHTKLKSSGDAQLVLPLDQRNKLNRILSTMQRRLVTAKTDMEDLIVRLNQEIAAKD 334
TD+ ++ +L + ++Q IL Q ++ K + L +L +
Sbjct: 472 TDLYNEIEIYKRDKDELEIQMEQLALDYEILK-QQNHDISYKLEQSQLQEQLKIQYECSS 530
Query: 335 FLT--TKVKDLEVELETSKQKNKENLQQAILIERERFTQMQWDMEELRRKSLEMEMKLKS 392
L T++++ LE +K E +++ +E +QM+ EE+ +++ E + +
Sbjct: 531 SLVDVTELENQVESLEAELKKQSEEFSESLCRIKELESQMETLEEEMEKQAQVFEADIDA 590
Query: 393 ESGGNVGQDLTKESIVQQNDVLLQD-------LDAAREQLEILSKQYGELEAKSKADIKV 445
+ G V Q+ + +Q + L + +++ + LS+Q + ++
Sbjct: 591 VTRGKVEQE---QRAIQAEETLRKTRWKNASVAGKLQDEFKRLSEQMDSMFTSNEKMAMK 647
Query: 446 LVKEVKSLRSSQTELKKELSESIKE------KYEAEKLLLHEREKSEQAEIAWRKQLEKC 499
+ E LR + +L++ + ++ E +YEA+ LHE + + + Q+E+
Sbjct: 648 AMTEANELRMQKRQLEEMIKDANDELRANQAEYEAK---LHELSEKLSFKTS---QMER- 700
Query: 500 GLLLKQLQECSVELPYE---DEDRTFLQSSSSTDAFNKLKTSDDQIDILLAEVENLEKDA 556
+L+ L E S E+ + +ED T + +I IL E+ENL+K+
Sbjct: 701 --MLENLDEKSNEIDNQKRHEEDVT--------------ANLNQEIKILKEEIENLKKNQ 744
Query: 557 GSAASNVDKTNDIK 570
S ++ +++
Sbjct: 745 DSLMLQAEQAENLR 758
Score = 38.1 bits (87), Expect = 0.014
Identities = 54/253 (21%), Positives = 105/253 (41%), Gaps = 35/253 (13%)
Query: 260 DGSVDLPGSASVSRETDIVGHTKLKSSGDAQLVLPLDQRNKLNRILSTMQRRLVTAKTDM 319
+ S+ + E+ I K S A+L + +++ +S +Q L T ++
Sbjct: 772 EASLQRENMKKIELESKISLMRKESESLAAELQVIKLAKDEKETAISLLQTELETVRSQC 831
Query: 320 EDLIVRLNQEIAAKDFLTTKVKDLEVELETSKQKNKENLQQAILIERERFTQM------- 372
+DL L++ + +V ++ EL+ K++ NL++ + R T+
Sbjct: 832 DDLKHSLSENDLEMEKHKKQVAHVKSELK-KKEETMANLEKKLKESRTAITKTAQRNNIN 890
Query: 373 ------------QWDMEELRRKSLEMEMKLKS---ESGGNVGQDLTKESIVQQNDVLLQD 417
+ + + + K LE ++KLK ES N+ +++ L
Sbjct: 891 KGSPVGAHGGSKEVAVMKDKIKLLEGQIKLKETALESSSNM--------FIEKEKNLKNR 942
Query: 418 LDAAREQLEILSKQYGE---LEAKSKADIKVLVKEVKSLRSSQTELKKELSESIKEKYEA 474
++ +L+ S++ E L + DI VLV E++SLR ++ EL E ++E+Y
Sbjct: 943 IEELETKLDQNSQEMSENELLNGQENEDIGVLVAEIESLRECNGSMEMELKE-MRERYSE 1001
Query: 475 EKLLLHEREKSEQ 487
L E E Q
Sbjct: 1002 ISLRFAEVEGERQ 1014
>At1g13220 putative nuclear matrix constituent protein
Length = 1128
Score = 47.4 bits (111), Expect = 2e-05
Identities = 60/250 (24%), Positives = 109/250 (43%), Gaps = 41/250 (16%)
Query: 263 VDLPGSASVSRETDIVGH-----TKLKSSGDAQLVLPLDQRNKLNRILSTMQRRLVTAKT 317
VDL S S E DI TK K + Q+ L K N + + ++ + T
Sbjct: 314 VDLSMSKSKETEEDITKRLEELTTKEKEAHTLQITLLA----KENELRAFEEKLIAREGT 369
Query: 318 DMEDLIVRLNQEIAAKDFLTTKVKDLEVELETSKQKNKENLQQAIL-IERE--------- 367
+++ LI K+ L +K+ + E+E E ++ + LQ+ I +ER+
Sbjct: 370 EIQKLIDD------QKEVLGSKMLEFELECEEIRKSLDKELQRKIEELERQKVEIDHSEE 423
Query: 368 ----RFTQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLTKESIVQQNDVLL----QDLD 419
R M + + K +++E KLK+ +E I+Q + L Q L
Sbjct: 424 KLEKRNQAMNKKFDRVNEKEMDLEAKLKTIK--------EREKIIQAEEKRLSLEKQQLL 475
Query: 420 AAREQLEILSKQYGELEAKSKADIKVLVKEVKSLRSSQTELKKELSESIKEKYEAEKLLL 479
+ +E LE L ++ ++ A+ +++ +E KSL + E ++ L + K + EK +
Sbjct: 476 SDKESLEDLQQEIEKIRAEMTKKEEMIEEECKSLEIKKEEREEYLRLQSELKSQIEKSRV 535
Query: 480 HEREKSEQAE 489
HE S++ E
Sbjct: 536 HEEFLSKEVE 545
Score = 47.4 bits (111), Expect = 2e-05
Identities = 63/263 (23%), Positives = 128/263 (47%), Gaps = 25/263 (9%)
Query: 300 KLNRILSTMQRRLVTAKTDMEDLIVRLNQEIAAK-DFLTTKVKDLEVELETSKQKNK--E 356
+L R + ++R+ V E L R NQ + K D + K DLE +L+T K++ K +
Sbjct: 403 ELQRKIEELERQKVEIDHSEEKLEKR-NQAMNKKFDRVNEKEMDLEAKLKTIKEREKIIQ 461
Query: 357 NLQQAILIERERFTQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLTKESIVQQNDVLLQ 416
++ + +E+++ + +E+L++ E+E K+++E KE ++++ L+
Sbjct: 462 AEEKRLSLEKQQLLSDKESLEDLQQ---EIE-KIRAEM-------TKKEEMIEEECKSLE 510
Query: 417 DLDAAREQLEILSKQYGELEAKSKADIKVLVKEVKSLRSSQTELKKELSESIKEK---YE 473
RE+ L + KS+ + L KEV++L+ + +KE E + EK Y
Sbjct: 511 IKKEEREEYLRLQSELKSQIEKSRVHEEFLSKEVENLKQEKERFEKEW-EILDEKQAVYN 569
Query: 474 AEKL-LLHEREKSEQAEIAWRKQL--EKCGLLLKQLQEC-SVELPYEDEDRTFLQSSSST 529
E++ + E+EK E+ ++ ++L E+ L ++ +QE + L E + S+
Sbjct: 570 KERIRISEEKEKFERFQLLEGERLKKEESALRVQIMQELDDIRLQRESFEANMEHERSAL 629
Query: 530 DAFNKLKTSD--DQIDILLAEVE 550
KL+ S D ++++ +E
Sbjct: 630 QEKVKLEQSKVIDDLEMMRRNLE 652
Score = 42.7 bits (99), Expect = 6e-04
Identities = 64/292 (21%), Positives = 126/292 (42%), Gaps = 29/292 (9%)
Query: 288 DAQLVLPLDQRNKLNRILSTMQRR------LVTAKTDMEDL--IVRLNQEIAAKDFLTTK 339
+AQ +L +Q + L + + QR L K +++L +R QE +K L+++
Sbjct: 126 EAQEILKREQSSHLYALTTVEQREENLRKALGLEKQCVQELEKALREIQEENSKIRLSSE 185
Query: 340 VKDLEVE-LETSKQKNKENLQQAILIERERFTQMQWDMEELRRKSLEMEMKLKSESGGNV 398
K +E L S +++ I + + E RKS E++++LK
Sbjct: 186 AKLVEANALVASVNGRSSDVENKIYSAESK-------LAEATRKSSELKLRLKEVE---- 234
Query: 399 GQDLTKESIVQQNDVLLQDLDAAREQLEILSKQY-GELEAKSKADIKVLVKEVKSLRSSQ 457
T+ES++QQ + + E ++Y E E K + + + ++ ++L +
Sbjct: 235 ----TRESVLQQERLSFTKERESYEGTFQKQREYLNEWEKKLQGKEESITEQKRNLNQRE 290
Query: 458 ---TELKKELSESIKEKYEAEKLLLHEREKSEQAEIAWRKQLEKCGLLLKQLQECSVEL- 513
E++K+L KE E + + KS++ E K+LE+ K+ + L
Sbjct: 291 EKVNEIEKKLKLKEKELEEWNRKVDLSMSKSKETEEDITKRLEELTTKEKEAHTLQITLL 350
Query: 514 PYEDEDRTFLQSSSSTDAFNKLKTSDDQIDILLAEVENLEKDAGSAASNVDK 565
E+E R F + + + K DDQ ++L +++ E + ++DK
Sbjct: 351 AKENELRAFEEKLIAREGTEIQKLIDDQKEVLGSKMLEFELECEEIRKSLDK 402
Score = 33.5 bits (75), Expect = 0.35
Identities = 43/216 (19%), Positives = 86/216 (38%), Gaps = 56/216 (25%)
Query: 353 KNKENLQQAILIERERF--------TQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLTK 404
+ KE ++ L+E ER Q+ +++++R + E ++ E + +
Sbjct: 577 EEKEKFERFQLLEGERLKKEESALRVQIMQELDDIRLQRESFEANMEHE------RSALQ 630
Query: 405 ESIVQQNDVLLQDLDAAREQLEI------------LSKQYGELEAKSKADI-------KV 445
E + + ++ DL+ R LEI L + + E K A++ +
Sbjct: 631 EKVKLEQSKVIDDLEMMRRNLEIELQERKEQDEKDLLDRMAQFEDKRMAELSDINHQKQA 690
Query: 446 LVKEVKSLRSSQTELKKELSE--SIKEKYEAEKLLLHE--------------------RE 483
L +E++ + S ++ L+KE E K+K + +++ +H RE
Sbjct: 691 LNREMEEMMSKRSALQKESEEIAKHKDKLKEQQVEMHNDISELSTLSINLKKRREVFGRE 750
Query: 484 KSE-QAEIAWRKQLEKCGLLLKQLQECSVELPYEDE 518
+S A + K CG L+ ++LP DE
Sbjct: 751 RSRFLAFVQKLKDCGSCGQLVNDFVLSDLQLPSNDE 786
>At5g41140 putative protein
Length = 983
Score = 46.6 bits (109), Expect = 4e-05
Identities = 49/228 (21%), Positives = 101/228 (43%), Gaps = 26/228 (11%)
Query: 297 QRNKLNRILSTMQRRLVTAKTDMEDLIVRLNQEIAAKDFLTTKVKDLEVELETSKQKNKE 356
Q+ +L +L L + + E +LN+ D T ++K + +LE K++ ++
Sbjct: 668 QKRQLEELLMNANDELRVNRVEYE---AKLNELSGKTDLKTKEMKRMSADLEYQKRQKED 724
Query: 357 ---NLQQAILIERERFTQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLTKESIVQQNDV 413
+L I ++ ++ D+EE R+ S+E E L E + I+ + +
Sbjct: 725 VNADLTHEITRRKDEIEILRLDLEETRKSSMETEASLSEE----------LQRIIDEKEA 774
Query: 414 LLQDLDAAREQLEILSKQYGELE---AKSKADIKVLVKEVKSLRSSQTELKKELSESIKE 470
+ + A + QLE L+ + ++++I+ L K+V +RS + ++E++
Sbjct: 775 V---ITALKSQLETAIAPCDNLKHSLSNNESEIENLRKQVVQVRSELEKKEEEMANLENR 831
Query: 471 KYEAEKLLLHEREKSEQAEIAWRKQLEKCGLLLKQLQECSVELPYEDE 518
+ A+ + E+ +E KQLE L + E S ++ E E
Sbjct: 832 EASADNITKTEQRSNEDR----IKQLEGQIKLKENALEASSKIFIEKE 875
Score = 36.6 bits (83), Expect = 0.042
Identities = 53/238 (22%), Positives = 106/238 (44%), Gaps = 35/238 (14%)
Query: 309 QRRLVTAKTDME------DLIVRLNQEIAAKDFLTTKVKDLEVELETSKQKNKENLQQAI 362
+R T++TD + D +V+ + + L ++ DL E+E K ++KE+L+ I
Sbjct: 444 RRMSCTSETDDDEDQKALDELVKGHMDAKEAHVLERRITDLYNEIEIYK-RDKEDLE--I 500
Query: 363 LIERERFTQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLTKESIVQQNDVLLQDLDAAR 422
+E Q+ D E L++++ ++ KL+ +S VQ+ + + ++
Sbjct: 501 QVE-----QLSLDYEILKQENHDISYKLE-------------QSQVQEQLKMQYECSSSL 542
Query: 423 EQLEILSKQYGELEAKSKADIKVLVKEVKSLRSSQTELK--KELSESIKEKYEAE-KLLL 479
+ L LEAK K K + + ++ +T++K +E E + +E + + +
Sbjct: 543 VNVNELENHVESLEAKLKKQYKECSESLYRIKELETQIKGMEEELEKQAQIFEGDIEAVT 602
Query: 480 HEREKSEQAEIAWRKQLEKCGLLLKQLQECSVELPYEDEDRTFLQSSSSTDAFNKLKT 537
+ + EQ I + L K + + SV +DE + + SST A N+ T
Sbjct: 603 RAKVEQEQRAIEAEEALRK-----TRWKNASVAGKIQDEFKRISEQMSSTLAANEKVT 655
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.310 0.129 0.356
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,522,002
Number of Sequences: 26719
Number of extensions: 611150
Number of successful extensions: 2932
Number of sequences better than 10.0: 296
Number of HSP's better than 10.0 without gapping: 38
Number of HSP's successfully gapped in prelim test: 260
Number of HSP's that attempted gapping in prelim test: 2482
Number of HSP's gapped (non-prelim): 546
length of query: 609
length of database: 11,318,596
effective HSP length: 105
effective length of query: 504
effective length of database: 8,513,101
effective search space: 4290602904
effective search space used: 4290602904
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 63 (28.9 bits)
Medicago: description of AC148405.4


