

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148405.2 + phase: 0
(209 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
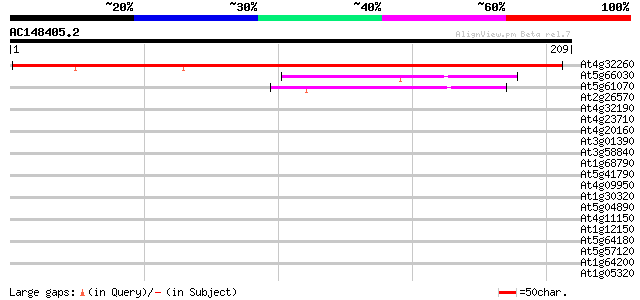
Score E
Sequences producing significant alignments: (bits) Value
At4g32260 H+-transporting ATP synthase chain 9 - like protein 251 1e-67
At5g66030 Golgi-localized protein GRIP 45 3e-05
At5g61070 HDA18 40 9e-04
At2g26570 unknown protein 39 0.002
At4g32190 unknown protein 39 0.002
At4g23710 vacuolar-type H+-ATPase (V-ATPase) subunit G2 (Vag2 / ... 39 0.003
At4g20160 Glu-rich protein 38 0.004
At3g01390 vacuolar-type H+-ATPase subunit G1 (VHA-G1 / AVMA10) 38 0.004
At3g58840 unknown protein 36 0.013
At1g68790 putative nuclear matrix constituent protein 1 (NMCP1) 35 0.028
At5g41790 myosin heavy chain-like protein 34 0.048
At4g09950 AIG1-like protein 34 0.048
At1g30320 unknown protein 34 0.048
At5g04890 unknown protein 34 0.063
At4g11150 vacuolar-type H+-ATPase subunit E1 (VHA-E1) 34 0.063
At1g12150 hypothetical protein 34 0.063
At5g64180 unknown protein 33 0.082
At5g57120 unknown protein 33 0.082
At1g64200 H+-transporting ATPase protein, putative 33 0.082
At1g05320 unknown protein 33 0.082
>At4g32260 H+-transporting ATP synthase chain 9 - like protein
Length = 219
Score = 251 bits (642), Expect = 1e-67
Identities = 139/216 (64%), Positives = 167/216 (76%), Gaps = 11/216 (5%)
Query: 2 ASMIMASSKPIIRVPTNSLPST---------PKLPSLQTITPNLKLQLSKLKSMTLAATS 52
A+ IMASSKP+I + +N P+ P++P T + +L+ + L S T + +
Sbjct: 3 ANSIMASSKPLISLSSNQQPNRVQIPKFAKLPQIPKSLTSSTDLRSKALSLSSATAKSLA 62
Query: 53 LSFVFAPPSLA--FEKAALFDFNLTLPIIVVEFLFLMFALDKLYFTPLGTFMDNRDAEIR 110
L FAPPS+A EKA LFDFNLTLPIIVVEFLFLMFALDK+Y++PLG FMD RDA I+
Sbjct: 63 LIAAFAPPSMAEAMEKAQLFDFNLTLPIIVVEFLFLMFALDKVYYSPLGNFMDQRDASIK 122
Query: 111 GKLNSVTNTSEEVKELEEQANAVLRAARAEIAVALNQMKKETQAEVEEKIAEGRKKVDAE 170
KL SV +TS EVKEL+EQA AV+RAARAEIA ALN+MKKETQ EVEEK+AEGRKKV+ E
Sbjct: 123 EKLASVKDTSTEVKELDEQAAAVMRAARAEIAAALNKMKKETQVEVEEKLAEGRKKVEEE 182
Query: 171 LQEALANLEKQKEETVKALDSQIAALSQDIVNKVLP 206
L+EALA+LE QKEET+KALDSQIAALS+DIV KVLP
Sbjct: 183 LKEALASLESQKEETIKALDSQIAALSEDIVKKVLP 218
>At5g66030 Golgi-localized protein GRIP
Length = 788
Score = 45.1 bits (105), Expect = 3e-05
Identities = 31/96 (32%), Positives = 48/96 (49%), Gaps = 9/96 (9%)
Query: 102 MDNRDAEIRGKLNSVTNTSEEVKELEEQANAVLRAARAEIAVA--------LNQMKKETQ 153
+ +DAE++G + E + +A+A+L+ E+A A L + KE +
Sbjct: 410 LSTKDAELKGAREEINRLQSEFSSYKIRAHALLQKKDMELAAAKDSEQIKSLEEALKEAE 469
Query: 154 AEVEEKIAEGRKKVDAELQEALANLEKQKEETVKAL 189
EV AE R + +LQ ALA+LEK+ EE AL
Sbjct: 470 KEVYLVSAE-RDRAQQDLQSALASLEKELEERAGAL 504
>At5g61070 HDA18
Length = 682
Score = 40.0 bits (92), Expect = 9e-04
Identities = 29/90 (32%), Positives = 46/90 (50%), Gaps = 3/90 (3%)
Query: 98 LGTFMDNRDAEI--RGKLNSVTNTSEEVKELEEQANAVLRAARAEIAVALNQMKKETQAE 155
L + M RD E+ R K N E E E +A +L AR ++ L+ + Q E
Sbjct: 446 LKSLMAARDGELEARRKELKAKNKELEANEKELEAGLMLIRAREDVICGLHAKIESLQQE 505
Query: 156 VEEKIAEGRKKVDAELQEALANLEKQKEET 185
+E +A+ +++D ELQE A ++ KE+T
Sbjct: 506 RDEAVAKA-ERIDKELQEDRARSQEFKEDT 534
>At2g26570 unknown protein
Length = 807
Score = 39.3 bits (90), Expect = 0.002
Identities = 27/74 (36%), Positives = 42/74 (56%), Gaps = 5/74 (6%)
Query: 115 SVTNTSEEVKELEEQANAVLRAARAEIAVALNQMK--KETQAEVEEKIAEGRKKVDAE-- 170
SVT + EE EL ++A+ A A +A A+++++ KET+ EK+ E + +DA
Sbjct: 636 SVTLSLEEYYELSKRAHEAEELANARVAAAVSRIEEAKETEMRSLEKLEEVNRDMDARKK 695
Query: 171 -LQEALANLEKQKE 183
L+EA EK KE
Sbjct: 696 ALKEATEKAEKAKE 709
Score = 27.7 bits (60), Expect = 4.5
Identities = 20/87 (22%), Positives = 43/87 (48%), Gaps = 1/87 (1%)
Query: 112 KLNSVTNTSEEVKELEEQANAVLRAARAE-IAVALNQMKKETQAEVEEKIAEGRKKVDAE 170
+LN ++S+++K + A+A+L +AE +A +++K+E +
Sbjct: 398 RLNQQIHSSKDLKSKLDTASALLLDLKAELVAYMESKLKQEACDSTTNTDPSTENMSHPD 457
Query: 171 LQEALANLEKQKEETVKALDSQIAALS 197
L A+A+ +K+ EE ++ A +S
Sbjct: 458 LHAAVASAKKELEEVNVNIEKAAAEVS 484
>At4g32190 unknown protein
Length = 783
Score = 38.9 bits (89), Expect = 0.002
Identities = 33/111 (29%), Positives = 59/111 (52%), Gaps = 15/111 (13%)
Query: 105 RDAEIRGKLNSVTNTSEEVK----------ELEEQANAVLRAARAEIAVALNQMKKETQA 154
++ E+ + N S+EV +L +AN V++ EI AL + +E +
Sbjct: 208 KEEELEKMRQEIANRSKEVSMAISEFESKSQLLSKANEVVKRQEGEI-YALQRALEEKEE 266
Query: 155 EVEEKIAEGRKKVDAE-LQEALANLEKQKEETVKALDSQIAALSQDIVNKV 204
E+E I++ KK++ E L+E ANL+KQ EE + A D ++ L ++ V ++
Sbjct: 267 ELE--ISKATKKLEQEKLRETEANLKKQTEEWLIAQD-EVNKLKEETVKRL 314
>At4g23710 vacuolar-type H+-ATPase (V-ATPase) subunit G2 (Vag2 /
VHA-G2)
Length = 106
Score = 38.5 bits (88), Expect = 0.003
Identities = 24/92 (26%), Positives = 53/92 (57%), Gaps = 7/92 (7%)
Query: 121 EEVKELEEQANAVLRAARAEIAVALNQMKKETQAEVEE---KIAEG-RKKVDAELQEALA 176
+++ E +A ++ AAR L Q K+E + EV E +G ++K++A ++ A
Sbjct: 7 QQLLAAEREAQQIVNAARTAKMTRLKQAKEEAETEVAEHKTSTEQGFQRKLEATSGDSGA 66
Query: 177 NLEKQKEET---VKALDSQIAALSQDIVNKVL 205
N+++ ++ET ++ L ++ +S+D+V+ +L
Sbjct: 67 NVKRLEQETDAKIEQLKNEATRISKDVVDMLL 98
>At4g20160 Glu-rich protein
Length = 1188
Score = 37.7 bits (86), Expect = 0.004
Identities = 27/88 (30%), Positives = 39/88 (43%), Gaps = 8/88 (9%)
Query: 102 MDNRDAEIRGKLNSVTNTS--EEVKELEEQANAVLRAARAEIAVALNQMKKETQAEVEEK 159
M+ R +GK TNT E LEE +L AE + + T+ EV EK
Sbjct: 497 MNRRKVVEKGKTEGNTNTERVERNDVLEEATRRILSVESAERSSTTTSKETMTRCEVAEK 556
Query: 160 IAEGRKKVDAELQEALANLEKQKEETVK 187
+ +G+KK D +LE K + V+
Sbjct: 557 VVKGKKKED------FVSLESNKSKEVE 578
>At3g01390 vacuolar-type H+-ATPase subunit G1 (VHA-G1 / AVMA10)
Length = 110
Score = 37.7 bits (86), Expect = 0.004
Identities = 23/92 (25%), Positives = 50/92 (54%), Gaps = 7/92 (7%)
Query: 121 EEVKELEEQANAVLRAARAEIAVALNQMKKETQAEVEEKIAEGRKKVDAELQEAL----A 176
+++ E +A ++ AAR L Q K+E + E+ E A+ + +L+E A
Sbjct: 11 QQLLAAEVEAQHIVNAARTAKMARLKQAKEEAEKEIAEYKAQTEQDFQRKLEETSGDSGA 70
Query: 177 NLEKQKEET---VKALDSQIAALSQDIVNKVL 205
N+++ ++ET ++ L ++ + +S+D+V +L
Sbjct: 71 NVKRLEQETDTKIEQLKNEASRISKDVVEMLL 102
>At3g58840 unknown protein
Length = 318
Score = 36.2 bits (82), Expect = 0.013
Identities = 30/126 (23%), Positives = 56/126 (43%), Gaps = 27/126 (21%)
Query: 98 LGTFMDNRDAEIRGKLNSVTNTSEEVKELEEQANAVLRAARAEIAVALNQMKKETQ---- 153
L T + N ++ LN V T+EEV EL++ A AEI L +KE +
Sbjct: 98 LETEVSNLHDDLITSLNGVDKTAEEVAELKK--------ALAEIVEKLEGCEKEAEGLRK 149
Query: 154 --AEVEEKIAEGRKKVDA-------------ELQEALANLEKQKEETVKALDSQIAALSQ 198
AEVE+++ + +K+ +E + ++ +K+ ++ L + L+
Sbjct: 150 DRAEVEKRVRDLERKIGVLEVREMEEKSKKLRSEEEMREIDDEKKREIEELQKTVIVLNL 209
Query: 199 DIVNKV 204
++V V
Sbjct: 210 ELVKNV 215
>At1g68790 putative nuclear matrix constituent protein 1 (NMCP1)
Length = 1085
Score = 35.0 bits (79), Expect = 0.028
Identities = 28/96 (29%), Positives = 48/96 (49%), Gaps = 7/96 (7%)
Query: 114 NSVTNTSEEVKELEEQANAVLRAARAEIAVALNQMKKETQAEVEEKIAEGRKKVDAEL-- 171
N + E+ +L + AVL + R E + L QM++ E+E K AE +++ E+
Sbjct: 340 NLIEREQMEIGKLLDDQKAVLDSRRREFEMELEQMRRSLDEELEGKKAE-IEQLQVEISH 398
Query: 172 -QEALANLE---KQKEETVKALDSQIAALSQDIVNK 203
+E LA E ++KEE VK + + A + + K
Sbjct: 399 KEEKLAKREAALEKKEEGVKKKEKDLDARLKTVKEK 434
Score = 27.3 bits (59), Expect = 5.9
Identities = 22/76 (28%), Positives = 34/76 (43%), Gaps = 10/76 (13%)
Query: 106 DAEIRGKLNSVTNTSEEVKELEEQANAVLRAARAEIAVALNQMKKETQAEVEEKIAEGRK 165
D E+ GK + E+ EE + A+ E A+ KKE + +EK + R
Sbjct: 379 DEELEGKKAEIEQLQVEISHKEE------KLAKREAALE----KKEEGVKKKEKDLDARL 428
Query: 166 KVDAELQEALANLEKQ 181
K E ++AL EK+
Sbjct: 429 KTVKEKEKALKAEEKK 444
Score = 27.3 bits (59), Expect = 5.9
Identities = 24/96 (25%), Positives = 46/96 (47%), Gaps = 5/96 (5%)
Query: 117 TNTSEEVKELEEQANAVLRAARAEIAVALNQMKKETQAEVEEKIAEGRKKVDAELQEALA 176
T+T+ E+++ ++A +L+ + A+ LN+ K + E K K+ AEL+ L
Sbjct: 92 TSTNNELQQAYDEAMEMLKREKTSNAITLNEADK--REENLRKALIDEKQFVAELENDLK 149
Query: 177 NLEKQK---EETVKALDSQIAALSQDIVNKVLPVSR 209
+++ + T +A + AL + K L V R
Sbjct: 150 YWQREHSVVKSTSEAKLEEANALVIGMKEKALEVDR 185
>At5g41790 myosin heavy chain-like protein
Length = 1305
Score = 34.3 bits (77), Expect = 0.048
Identities = 29/105 (27%), Positives = 53/105 (49%), Gaps = 11/105 (10%)
Query: 102 MDNRDAEIRGKLNSVTNTSEEVKELEEQANAVLRAARAEIA--VALNQMKKETQA----E 155
+ +++ E KL NT +E+ + R +E++ V +++ + + E
Sbjct: 187 ISSKNVETMNKLEQTQNTIQELMAELGKLKDSHREKESELSSLVEVHETHQRDSSIHVKE 246
Query: 156 VEEKIAEGRKKVDAELQEALANLEKQKEETVKALDSQIAALSQDI 200
+EE++ E KK+ AEL + L N E++K K L +IA LS +I
Sbjct: 247 LEEQV-ESSKKLVAELNQTLNNAEEEK----KVLSQKIAELSNEI 286
Score = 32.7 bits (73), Expect = 0.14
Identities = 27/98 (27%), Positives = 50/98 (50%), Gaps = 10/98 (10%)
Query: 107 AEIRGKLNSVTNTSEEVKELEEQANAVLRAARAEIAVALNQMKKETQAEVEEKIAEGRKK 166
AEI G + + S + +E+E+Q + E +V + ++ E ++ + ++
Sbjct: 833 AEIDGLRAELDSMSVQKEEVEKQ----MVCKSEEASVKIKRLDDEVNGLRQQVASLDSQR 888
Query: 167 VDAELQEALANLEKQKEETVKALDSQIAALSQDIVNKV 204
+ E+Q LEK+ EE + L SQI L ++I+NKV
Sbjct: 889 AELEIQ-----LEKKSEEISEYL-SQITNLKEEIINKV 920
Score = 27.3 bits (59), Expect = 5.9
Identities = 24/81 (29%), Positives = 38/81 (46%), Gaps = 14/81 (17%)
Query: 118 NTSEEVKELEEQANAVLRAARAEIAVALNQMKKETQAEVEEKIAEGRKKVDAELQEALAN 177
++S VKELEEQ + + AE+ LN AE KKV ++ L+N
Sbjct: 239 DSSIHVKELEEQVESSKKLV-AELNQTLNN-------------AEEEKKVLSQKIAELSN 284
Query: 178 LEKQKEETVKALDSQIAALSQ 198
K+ + T++ L S+ L +
Sbjct: 285 EIKEAQNTIQELVSESGQLKE 305
>At4g09950 AIG1-like protein
Length = 336
Score = 34.3 bits (77), Expect = 0.048
Identities = 21/84 (25%), Positives = 45/84 (53%), Gaps = 1/84 (1%)
Query: 121 EEVKELEEQANAVLRAARAEIAVALNQMKKETQAEVEEKIAEGRKKVDAELQEALANLEK 180
E+ K++EE + +++ L + E ++EKI+ K+ +++E LA +
Sbjct: 229 EKQKQIEEMKGWSSKQEISQMKKELEKSHNEMLEGIKEKISNQLKESLEDVKEQLAKAQA 288
Query: 181 QKEETVKALDSQIAALSQDIVNKV 204
++EET K + ++I LS D + ++
Sbjct: 289 EREETEKKM-NEIQKLSSDEIRRL 311
>At1g30320 unknown protein
Length = 509
Score = 34.3 bits (77), Expect = 0.048
Identities = 43/186 (23%), Positives = 79/186 (42%), Gaps = 18/186 (9%)
Query: 13 IRVPTNSLPSTPK--LPSLQTITPNLKLQLSKLKSMTLAA---TSLSFVFAPPSLAFEKA 67
+R PT+SLPSTP+ P +++ N + +LS+ + +L ++A +
Sbjct: 315 LRSPTSSLPSTPRGGQPEESSMSKNTRRELSEEEEKAKTRREIVALGVQLGKMNIAAWAS 374
Query: 68 ALFDFNLTLPIIVVEFLFLMFALDKLYFTPLGTFMDNRDAEIRGKLNSVTNTSE-EVKEL 126
+ N E K+ F T + + + K N+ E ++
Sbjct: 375 KEEEENKKNNGDAEE-------AQKIEFEKRATAWEEAE---KSKHNARYKREEIRIQAW 424
Query: 127 EEQANAVLRAARAEIAVALNQMKKETQAEVEEKIAEGRKKVDAELQEALANLEKQKEETV 186
E Q A L A I + QMK E +A++ +KIA +++ +E + ALA K ++
Sbjct: 425 ESQEKAKLEAEMRRIEAKVEQMKAEAEAKIMKKIALAKQR--SEEKRALAEARKTRDAEK 482
Query: 187 KALDSQ 192
++Q
Sbjct: 483 AVAEAQ 488
>At5g04890 unknown protein
Length = 366
Score = 33.9 bits (76), Expect = 0.063
Identities = 26/102 (25%), Positives = 51/102 (49%), Gaps = 3/102 (2%)
Query: 102 MDNRDAEIRGKLNSVTNTSEE--VKELEEQANAVLRAARAEIAVALNQMKKETQAEVEEK 159
++ ++A IR KL EE +++L+E+A A AA ++ + +K + ++EE+
Sbjct: 156 LEEKEALIR-KLQEEAKAKEEAEMRKLQEEAKAKEEAAAKKLQEEIEAKEKLEERKLEER 214
Query: 160 IAEGRKKVDAELQEALANLEKQKEETVKALDSQIAALSQDIV 201
E RK D +L E + Q+ ++V + L ++V
Sbjct: 215 RLEERKLEDMKLAEEAKLKKIQERKSVDESGEKEKILKPEVV 256
>At4g11150 vacuolar-type H+-ATPase subunit E1 (VHA-E1)
Length = 230
Score = 33.9 bits (76), Expect = 0.063
Identities = 27/92 (29%), Positives = 46/92 (49%), Gaps = 11/92 (11%)
Query: 124 KELEEQANAVLRAARAEIAVALNQM----KKETQAEVE--EKIAEGRKKVDAELQEALAN 177
+E EE+AN + +A E + Q+ KK+ + + E EK A+ RKK+D +Q
Sbjct: 19 QEAEEKANEISVSAEEEFNIEKLQLVEAEKKKIRQDYEKKEKQADVRKKIDYSMQ----- 73
Query: 178 LEKQKEETVKALDSQIAALSQDIVNKVLPVSR 209
L + + ++A D + A+ +L VSR
Sbjct: 74 LNASRIKVLQAQDDIVNAMKDQAAKDLLNVSR 105
>At1g12150 hypothetical protein
Length = 548
Score = 33.9 bits (76), Expect = 0.063
Identities = 23/80 (28%), Positives = 40/80 (49%), Gaps = 1/80 (1%)
Query: 109 IRGKLNSVTNTSEEVKELEEQANAVLRAARAEIAVALNQMKKETQAEVEEKIAEGRKKVD 168
I +LN T +E + E +++ + R E+ + ++ Q E E E KK++
Sbjct: 303 ITNELNEATMRLQEAADDECSLRSLVNSLRMELEDLRREREELQQKEAERLEIEETKKLE 362
Query: 169 AELQEALANLEKQKEETVKA 188
A QE+L LE+ K E ++A
Sbjct: 363 ALKQESL-KLEQMKTEAIEA 381
>At5g64180 unknown protein
Length = 158
Score = 33.5 bits (75), Expect = 0.082
Identities = 21/85 (24%), Positives = 39/85 (45%)
Query: 116 VTNTSEEVKELEEQANAVLRAARAEIAVALNQMKKETQAEVEEKIAEGRKKVDAELQEAL 175
V+ + +K LE++ + R A I A + ++ QAE +K AE R + + E
Sbjct: 51 VSEQARTIKVLEQRVQTLERELDAAITAAAHARSEKRQAESSQKAAESRAQDVTKELENT 110
Query: 176 ANLEKQKEETVKALDSQIAALSQDI 200
+ K E ++ + QI+ +I
Sbjct: 111 TKVFKLHMEELRGMQEQISKRDNEI 135
>At5g57120 unknown protein
Length = 330
Score = 33.5 bits (75), Expect = 0.082
Identities = 23/93 (24%), Positives = 44/93 (46%), Gaps = 8/93 (8%)
Query: 101 FMDNRDAEIRGKLNSVTNTSEEVKELEEQANAVLRAARAEIAVALNQMKKETQAEVEEKI 160
F++ RD E N+ N E V+ +++ + ++ V + KET A +E+ +
Sbjct: 63 FLNKRDHEAAANGNTEANVVEAVENVKKDKKKK-KNKETKVEVTEEEKVKETDAVIEDGV 121
Query: 161 AEGRKKVDAELQEALANLEKQKEETVKALDSQI 193
E +KK + +++ +EE VK D+ I
Sbjct: 122 KEKKKKKETKVKVT-------EEEKVKETDAVI 147
>At1g64200 H+-transporting ATPase protein, putative
Length = 237
Score = 33.5 bits (75), Expect = 0.082
Identities = 25/92 (27%), Positives = 48/92 (52%), Gaps = 11/92 (11%)
Query: 124 KELEEQANAVLRAARAEIAVALNQM----KKETQAEVE--EKIAEGRKKVDAELQEALAN 177
+E EE+AN + ++ E + Q+ KK+ + E E EK + RKK+D +Q
Sbjct: 19 QEAEEKANEISISSEEEFNIEKLQLVEAEKKKIRQEYEKKEKQVDVRKKIDYSMQ----- 73
Query: 178 LEKQKEETVKALDSQIAALSQDIVNKVLPVSR 209
L + + ++A D + A+ ++ ++L VS+
Sbjct: 74 LNASRIKVLQAQDDIVNAMKEEAAKQLLKVSQ 105
>At1g05320 unknown protein
Length = 841
Score = 33.5 bits (75), Expect = 0.082
Identities = 27/103 (26%), Positives = 47/103 (45%), Gaps = 11/103 (10%)
Query: 103 DNRDAEIRGKLNSVTNTSEEVKELEEQANAVLRAARAEIAVALNQMKKETQAEVEE---- 158
D R E+ L + ++ELE++ + AE+ + LNQ +E ++
Sbjct: 470 DTRKVEVEEALLKLNTLESTIEELEKENGDL-----AEVNIKLNQKLANQGSETDDFQAK 524
Query: 159 -KIAEGRKKVDA-ELQEALANLEKQKEETVKALDSQIAALSQD 199
+ E K A ELQ + +L KQ + L SQI++L ++
Sbjct: 525 LSVLEAEKYQQAKELQITIEDLTKQLTSERERLRSQISSLEEE 567
Score = 26.9 bits (58), Expect = 7.7
Identities = 19/86 (22%), Positives = 44/86 (51%), Gaps = 2/86 (2%)
Query: 120 SEEVKELEEQANAVLRAARAEIAVALNQMKKETQAEVE-EKIAEGRKKVDAELQEALANL 178
+E+ K+LEE+ +L+ ++ Q+ E E +A+ ++ ++QE L
Sbjct: 364 TEKSKDLEEKIRVYEGKLAEACGQSLSLQEELDQSSAENELLADTNNQLKIKIQELEGYL 423
Query: 179 EKQKEETVKALDSQIAALSQDIVNKV 204
+ +KE ++ L +Q ++D++ K+
Sbjct: 424 DSEKETAIEKL-NQKDTEAKDLITKL 448
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.314 0.128 0.329
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,738,324
Number of Sequences: 26719
Number of extensions: 141457
Number of successful extensions: 1196
Number of sequences better than 10.0: 171
Number of HSP's better than 10.0 without gapping: 38
Number of HSP's successfully gapped in prelim test: 134
Number of HSP's that attempted gapping in prelim test: 1067
Number of HSP's gapped (non-prelim): 248
length of query: 209
length of database: 11,318,596
effective HSP length: 95
effective length of query: 114
effective length of database: 8,780,291
effective search space: 1000953174
effective search space used: 1000953174
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 58 (26.9 bits)
Medicago: description of AC148405.2


