

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148405.1 + phase: 0
(149 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
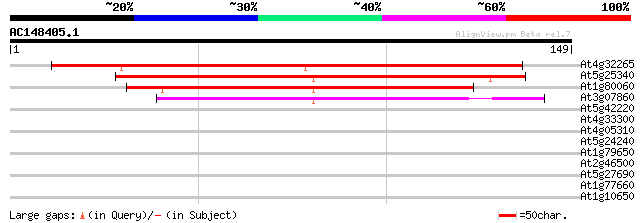
Score E
Sequences producing significant alignments: (bits) Value
At4g32265 unknown protein 144 2e-35
At5g25340 unknown protein 118 1e-27
At1g80060 hypothetical protein 93 5e-20
At3g07860 unknown protein 55 1e-08
At5g42220 unknown protein 30 0.48
At4g33300 putative protein 30 0.63
At4g05310 putative protein 29 0.82
At5g24240 ubiquitin 29 1.1
At1g79650 putative RAD23 protein 28 2.4
At2g46500 putative ubiquitin 27 3.1
At5g27690 unknown protein (At5g27690) 26 9.0
At1g77660 unknown protein 26 9.0
At1g10650 similar to putative inhibitor of apoptosis gi|3924605;... 26 9.0
>At4g32265 unknown protein
Length = 239
Score = 144 bits (362), Expect = 2e-35
Identities = 73/134 (54%), Positives = 94/134 (69%), Gaps = 9/134 (6%)
Query: 12 PRRSLSIQLSSMIIVDC-----TLPYDKLPSQNLTLTVLKLDSSSFQVEVAKTATVAVLK 66
PRRSL+ LS + ++D + Y+++P + + LTVLKLD SSF ++V KTATV LK
Sbjct: 5 PRRSLAASLSPLRLIDGLPRRRSFNYNQMPEEPIKLTVLKLDGSSFGIQVLKTATVGELK 64
Query: 67 QAVEAAFRHKP----EKISWPLVWGQFCLCYEGQKLVTETDYLRDYGIKDGDQLRFIRHV 122
AVEAAF H P KISWP VWGQFCL YE ++L+ E++YL ++GIKDGDQLRFIRH+
Sbjct: 65 MAVEAAFSHLPISGPGKISWPHVWGQFCLSYEDKRLINESEYLTEFGIKDGDQLRFIRHI 124
Query: 123 SNVCNFQRKPKKRT 136
SN C K K +T
Sbjct: 125 SNYCMLMVKHKSKT 138
>At5g25340 unknown protein
Length = 208
Score = 118 bits (296), Expect = 1e-27
Identities = 60/117 (51%), Positives = 79/117 (67%), Gaps = 8/117 (6%)
Query: 29 TLPYDKLPSQNLTLTVLKLDSSSFQVEVAKTATVAVLKQAVEAAFRHKPEK----ISWPL 84
T YDKLP++ + L+VLKLD SSF V V +ATV LK A+E AF H P+K ISW
Sbjct: 8 TFSYDKLPNEPIRLSVLKLDGSSFDVYVLTSATVGDLKVAIETAFSHVPKKGPSKISWSH 67
Query: 85 VWGQFCLCYEGQKLVTETDYLRDYGIKDGDQLRFIRHVSNVC----NFQRKPKKRTV 137
VWG FCLC+ GQKL+T+TD + +YG+KDGD++RF HVS + RK K++ +
Sbjct: 68 VWGHFCLCFGGQKLITDTDCIGNYGMKDGDEVRFKNHVSGNAVLSKGYSRKSKQKNL 124
>At1g80060 hypothetical protein
Length = 243
Score = 93.2 bits (230), Expect = 5e-20
Identities = 50/97 (51%), Positives = 63/97 (64%), Gaps = 5/97 (5%)
Query: 32 YDKLPSQN-LTLTVLKLDSSSFQVEVAKTATVAVLKQAVEAAFRHKPEK----ISWPLVW 86
Y KLP Q + L+V+KL+ S F VEVAK +VA LK+AVE F P + ISW VW
Sbjct: 44 YLKLPPQGRIKLSVVKLNGSLFDVEVAKDCSVAELKRAVEQVFTISPLEGHGMISWSHVW 103
Query: 87 GQFCLCYEGQKLVTETDYLRDYGIKDGDQLRFIRHVS 123
G FCLCY Q+LV + +R G+ DGDQL F+RH+S
Sbjct: 104 GHFCLCYRDQRLVNDKTSIRYLGLNDGDQLHFVRHLS 140
>At3g07860 unknown protein
Length = 165
Score = 55.5 bits (132), Expect = 1e-08
Identities = 34/109 (31%), Positives = 54/109 (49%), Gaps = 12/109 (11%)
Query: 40 LTLTVLKLDSSSFQVEVAKTATVAVLKQAVEAAFRHKPEK------ISWPLVWGQFCLCY 93
+ L+V+KLD SS V V +AT+ LK ++ + ISW VW FCL
Sbjct: 52 MRLSVVKLDGSSLDVAVMNSATLKDLKLLIKKKVNEMEQANMGHRHISWKHVWSNFCLSC 111
Query: 94 EGQKLVTETDYLRDYGIKDGDQLRFIRHVSNVCNFQRKPKKRTVNLKQH 142
+KL+ + L+D GI++ Q+ F+ +V +K + R K+H
Sbjct: 112 NNEKLLDDNAVLQDVGIRNNSQVTFMPYV------MKKGRGRHSKRKKH 154
>At5g42220 unknown protein
Length = 879
Score = 30.0 bits (66), Expect = 0.48
Identities = 22/87 (25%), Positives = 42/87 (47%), Gaps = 10/87 (11%)
Query: 33 DKLPSQNLTLTVLKLDSSSFQVEVAKTATVAVLKQAVEAAFRHKPEKISWPLVWGQFCLC 92
+K P L L + LDS ++ +V K TV + K+ + + + P+ GQ L
Sbjct: 17 EKTPESTLELNIKTLDSRTYTFQVNKNETVLLFKEKIAS-------ETGVPV--GQQRLI 67
Query: 93 YEGQKLVTETDYLRDYGIKDGDQLRFI 119
+ G +++ + L +Y +++G L I
Sbjct: 68 FRG-RVLKDDHPLSEYHLENGHTLHLI 93
>At4g33300 putative protein
Length = 855
Score = 29.6 bits (65), Expect = 0.63
Identities = 32/111 (28%), Positives = 47/111 (41%), Gaps = 10/111 (9%)
Query: 25 IVDCTLPYD---KLPSQNLTLTVLKLDS-SSFQVEVAKTATVAVLKQAVEAAFRHKPEKI 80
+ DC P+D KL + T LD +SF+ T V+ K E F + E +
Sbjct: 265 VPDCNFPFDGARKLVILDDVWTTQALDRLTSFKFPGCTTLVVSRSK-LTEPKFTYDVEVL 323
Query: 81 SWPLVWGQFCLCYEGQKLVTE---TDYLRDYGIKDGDQLRF--IRHVSNVC 126
S FCLC GQK + D ++ + I+ LR + V+N C
Sbjct: 324 SEDEAISLFCLCAFGQKSIPLGFCKDLVKQHNIQSFSILRVLCLAQVANEC 374
>At4g05310 putative protein
Length = 415
Score = 29.3 bits (64), Expect = 0.82
Identities = 13/28 (46%), Positives = 20/28 (71%)
Query: 47 LDSSSFQVEVAKTATVAVLKQAVEAAFR 74
L SSF++EV +T T+ V+KQ +E + R
Sbjct: 8 LGGSSFEIEVERTDTLLVVKQKIEKSQR 35
>At5g24240 ubiquitin
Length = 574
Score = 28.9 bits (63), Expect = 1.1
Identities = 11/35 (31%), Positives = 21/35 (59%)
Query: 91 LCYEGQKLVTETDYLRDYGIKDGDQLRFIRHVSNV 125
L Y+G+++ +RDYG+ DG L + +S++
Sbjct: 74 LLYDGREVSRNDSQIRDYGLADGKLLHLVIRLSDL 108
>At1g79650 putative RAD23 protein
Length = 371
Score = 27.7 bits (60), Expect = 2.4
Identities = 20/74 (27%), Positives = 36/74 (48%), Gaps = 6/74 (8%)
Query: 40 LTLTVLKLDSSSFQVEVAKTATVAVLKQAVEAAFRHKPEKISWPLVWGQFCLCYEGQKLV 99
+ LTV L S F++ V + T+ +K+ +E K ++P GQ L + G+ L
Sbjct: 1 MKLTVKTLKGSHFEIRVLPSDTIMAVKKNIE----DSQGKDNYPC--GQQLLIHNGKVLK 54
Query: 100 TETDYLRDYGIKDG 113
ET + + ++G
Sbjct: 55 DETSLVENKVTEEG 68
>At2g46500 putative ubiquitin
Length = 566
Score = 27.3 bits (59), Expect = 3.1
Identities = 10/35 (28%), Positives = 21/35 (59%)
Query: 91 LCYEGQKLVTETDYLRDYGIKDGDQLRFIRHVSNV 125
L + G++L +RDYG+ +G+ L + +S++
Sbjct: 76 LVFGGRELARSNSNMRDYGVSEGNILHLVLKLSDL 110
>At5g27690 unknown protein (At5g27690)
Length = 352
Score = 25.8 bits (55), Expect = 9.0
Identities = 8/14 (57%), Positives = 13/14 (92%)
Query: 121 HVSNVCNFQRKPKK 134
+++N CN+Q+KPKK
Sbjct: 101 NINNDCNYQKKPKK 114
>At1g77660 unknown protein
Length = 421
Score = 25.8 bits (55), Expect = 9.0
Identities = 14/50 (28%), Positives = 24/50 (48%), Gaps = 4/50 (8%)
Query: 37 SQNLTL----TVLKLDSSSFQVEVAKTATVAVLKQAVEAAFRHKPEKISW 82
S+NL L L L +S + V +A+ + V + +HKP+ + W
Sbjct: 102 SENLLLGLIFVALALFFASRNMAVINQTVIAIKQIRVRSRIKHKPKPVQW 151
>At1g10650 similar to putative inhibitor of apoptosis gi|3924605;
similar to ESTs gb|AI994578.1, gb|T04171, gb|AA728525
Length = 339
Score = 25.8 bits (55), Expect = 9.0
Identities = 14/45 (31%), Positives = 24/45 (53%)
Query: 3 ELTSVSNESPRRSLSIQLSSMIIVDCTLPYDKLPSQNLTLTVLKL 47
E S+SN R+ +Q+S + T +++P +NL T L+L
Sbjct: 66 EAESISNNIQRQQQKLQMSLNYNYNNTSVREEVPKENLVSTGLRL 110
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.320 0.133 0.390
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,038,474
Number of Sequences: 26719
Number of extensions: 110268
Number of successful extensions: 262
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 10
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 248
Number of HSP's gapped (non-prelim): 13
length of query: 149
length of database: 11,318,596
effective HSP length: 90
effective length of query: 59
effective length of database: 8,913,886
effective search space: 525919274
effective search space used: 525919274
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 55 (25.8 bits)
Medicago: description of AC148405.1


