

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148404.10 + phase: 0
(400 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
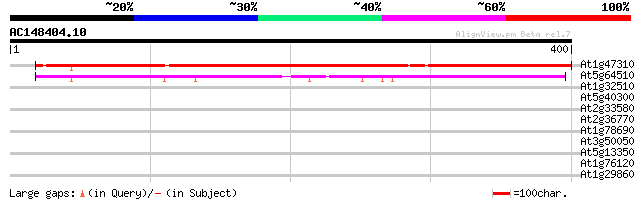
Score E
Sequences producing significant alignments: (bits) Value
At1g47310 unknown protein 286 2e-77
At5g64510 unknown protein 114 1e-25
At1g32510 unknown protein 31 1.1
At5g40300 unknown protein 30 1.8
At2g33580 putative protein kinase 30 2.3
At2g36770 putative glucosyl transferase 29 4.0
At1g78690 unknown protein 29 4.0
At3g50050 unknown protein 29 5.2
At5g13350 auxin-responsive - like protein 28 6.8
At1g76120 unknown protein 28 8.9
At1g29860 putative protein 28 8.9
>At1g47310 unknown protein
Length = 395
Score = 286 bits (731), Expect = 2e-77
Identities = 166/387 (42%), Positives = 241/387 (61%), Gaps = 11/387 (2%)
Query: 19 FLQLFTAFSNASSSSSSTHSNITH--IFQDILKAISSRQKWDLNDVRVFNFDVAKIRFGT 76
F+Q+ T + A S SNIT I QD+LK IS +QKW+L +VR +V KIR GT
Sbjct: 15 FIQVLT-LAVALDPSQPDESNITATPILQDVLKEISVKQKWNLEEVRFSKLEVKKIRIGT 73
Query: 77 SQNYLFRIGSSKNNFTVKFSDEISSWNHNKFTTTPKPDLASLVDQLSSIAFLDY-IKLEG 135
S+ + RI K+ F F DEI+ W + + +L LV +++S LD + L+G
Sbjct: 74 SRRFEIRIRLGKSRFVFIFPDEITDWRRSGGGSDV--ELQELVREVNSSKVLDPPLVLKG 131
Query: 136 PFELRVHESHHLSLSLPMNVSYNGLKHIIVGKGITVEVRRAREISFYYQSDLDLQRNGSV 195
PFEL V + LSLSLPMN+S++GLK ++V +GI+VE+R A+ +S ++ S
Sbjct: 132 PFELLVDGNDRLSLSLPMNISHSGLKRVLVSEGISVEIREAQAVSLFHSSHRRYAATVDP 191
Query: 196 ICSNQKNEFWPFLQSMCVPLIPIRIIGSASLIAYVARNPYVQIGTALISEDAVELLPEKC 255
+ + + W F S+CVPL PI+IIGSASL+A+ N QI T+ +S++A+ L EKC
Sbjct: 192 VNIKEGSSLWSFWGSVCVPLPPIQIIGSASLVAFRTSNATTQIKTSYLSDEAIHLYAEKC 251
Query: 256 YHGC-VFRKQACPVASLNLRLILLEKILRSLLGHKILQDRLSGLIKANIKAYAGVKFPLE 314
Y+ +R+ P L L++ LEK+L S LG+ Q S + A +KA V+F LE
Sbjct: 252 YYKAHTYRQHRFPNDLLGLKIHKLEKVLNS-LGNGTRQTVSS--VTAKLKASGMVRFQLE 308
Query: 315 LERDVG-NNATLSTLPDWRTRPSVERVWFEVMARVEDSRLKPLSIKKVKPFIESDSVSWA 373
+ER +G N + +S WRT+P +ERVWFEV A++E +LK + ++KV PFIE D+ +W+
Sbjct: 309 IERSIGKNESVISKKVAWRTKPKIERVWFEVTAKIEGDKLKAVRLRKVVPFIEVDTEAWS 368
Query: 374 NLMSNLSYTKLRPVLLPPEALTLDVKW 400
+LMSN+S+TK +L+P EALTLDVKW
Sbjct: 369 SLMSNMSFTKFPSLLVPQEALTLDVKW 395
>At5g64510 unknown protein
Length = 424
Score = 114 bits (284), Expect = 1e-25
Identities = 104/419 (24%), Positives = 188/419 (44%), Gaps = 48/419 (11%)
Query: 19 FLQLFTAFSNASSSSSSTHSNITH--IFQDILKAISSRQKWDLNDVRVFNFDVAKIRFGT 76
FL LFT F ++S+ SS +N + D+ ++I + +V+V FD+ G
Sbjct: 13 FLILFTIFPSSSALISSPDANPPYPKAISDLKESIVKGLGFQSEEVKVSGFDIRDALVGH 72
Query: 77 SQNYLFRIGSSKNNFTVKFSDEISSWNHNKFTT--TPKPDLASLVDQLSSIAFLDYI--- 131
S +Y F + K ++++ W + +P LV + D +
Sbjct: 73 SVSYEFDLEIDNKVLPFKLLEDVNRWEYVDLPIFQVEQPSENGLVPMRNKKTSSDDVLPV 132
Query: 132 ----KLEGPFELRVHESHHLSLSLPMNVSYNGLKHIIVGKGITVEVRRAREISFYYQSDL 187
+L GP EL + +++++ LSLP +V LK +I+ G V V+ AR +S + DL
Sbjct: 133 LAPFQLSGPMELWIQDANNMRLSLPYDVDAGVLKKVILADGAVVTVKGARSVSLRHPIDL 192
Query: 188 DLQRNGSVICSNQKNEFWPFLQSMC-----------VPLIPIRIIGSASLIAYVARNPYV 236
L N S NEF L S+ P++ +RI+G SL A +++P
Sbjct: 193 PLPLNQS------SNEFASGLLSLAEQLRRASTDQESPVLSLRIVGPTSL-ASTSQSPDN 245
Query: 237 QIGTALISEDAVEL------------LPEKCYHGCVFRKQ---ACPVASL---NLRLILL 278
++ ++ VEL + + ++ P+ S+ N L+
Sbjct: 246 KLKLKRLAPGLVELSSMSKDKRSLSTIGANAMTTVLTPREFTTMWPITSINGSNANLLGF 305
Query: 279 EKILRSLLGHKILQDRLSGLIKANIKAYAGVKFPLELERDVGN-NATLSTLPDWRTRPSV 337
EK+L S+LG K + ++KA + A +K +E+ + + + P+WRT+P
Sbjct: 306 EKLLTSVLGPKAQEKGSFKVLKAKVAAQTFMKIGFGIEKKLKEADVEGLSFPEWRTKPET 365
Query: 338 ERVWFEVMARVEDSRLKPLSIKKVKPFIESDSVSWANLMSNLSYTKLRPVLLPPEALTL 396
R+ FEV+A+V+ + P ++ +V P D+++ + N++ +KL + PP TL
Sbjct: 366 MRMHFEVLAKVDGENVIPENVMRVDPIPLEDTIAQNVITGNVTMSKLPIIESPPSPFTL 424
>At1g32510 unknown protein
Length = 472
Score = 31.2 bits (69), Expect = 1.1
Identities = 31/124 (25%), Positives = 52/124 (41%), Gaps = 9/124 (7%)
Query: 27 SNASSSSSSTHSNITHIFQDILKAISSRQKWDLNDVRVFNFDVAKIRFGTSQNYLFRIGS 86
S++SSSSS T T +F R +W +++ R+F+ D ++ ++
Sbjct: 292 SSSSSSSSITGCRKTLVFYMGRAPFGGRTEWVMHEYRLFDNDTSQGSLNFKGDFALCRVI 351
Query: 87 SKNNFTVK--------FSDEISSWNHNKFTTTPKPDLASLVDQLSSIAFLDYIKLEGPFE 138
+N T+K SDE S N N F + S + ++ D+I LE F+
Sbjct: 352 KRNEHTLKKCEIISPEVSDESLSNNVNNFCQASDLEKGSCDASNTRLSSPDFI-LESSFQ 410
Query: 139 LRVH 142
H
Sbjct: 411 GNSH 414
>At5g40300 unknown protein
Length = 270
Score = 30.4 bits (67), Expect = 1.8
Identities = 16/59 (27%), Positives = 30/59 (50%), Gaps = 5/59 (8%)
Query: 72 IRFGTSQNYLFRIGSSKNNFTVKFSDEISSWNHNKFTTTPKPDLASLVDQLSSIAFLDY 130
+ FG Q + + S+ + +++ D S+W +KF PDLA LS ++F+ +
Sbjct: 200 LEFGLDQMLAYLLASASTSASIRVDDWQSNWGADKF-----PDLARASVALSYVSFVAF 253
>At2g33580 putative protein kinase
Length = 664
Score = 30.0 bits (66), Expect = 2.3
Identities = 18/64 (28%), Positives = 35/64 (54%), Gaps = 4/64 (6%)
Query: 277 LLEKILRSLLGHKILQDRLSGLIKANIKAYAGVKFPLELERDVGNNATLSTLPDWRTRPS 336
+L K++ S+LG + ++++L + ++ G ++PLEL + A D +RPS
Sbjct: 577 MLCKVINSVLGGENVREKLKEFMDPSL----GNEYPLELAYTMAQLAKSCVATDLNSRPS 632
Query: 337 VERV 340
V +V
Sbjct: 633 VTQV 636
>At2g36770 putative glucosyl transferase
Length = 496
Score = 29.3 bits (64), Expect = 4.0
Identities = 11/26 (42%), Positives = 18/26 (68%)
Query: 29 ASSSSSSTHSNITHIFQDILKAISSR 54
A S+HSNIT++ QDI++ + S+
Sbjct: 470 AVEEGGSSHSNITYLLQDIMQQVKSK 495
>At1g78690 unknown protein
Length = 284
Score = 29.3 bits (64), Expect = 4.0
Identities = 22/68 (32%), Positives = 38/68 (55%), Gaps = 8/68 (11%)
Query: 216 IPIRII--GSASLIAYVARNPYVQIGTALISEDAVELLPEKCYHGCVFRKQACPVASLNL 273
+PIR + G+ASLIA R+P I +I E++PE +G R+ P+ + +L
Sbjct: 156 VPIRRLKWGTASLIA---RSPVTPIVLPIIHRGFEEMMPENYNNG---RRPLVPLPNKHL 209
Query: 274 RLILLEKI 281
++++ E I
Sbjct: 210 KVVVGEPI 217
>At3g50050 unknown protein
Length = 632
Score = 28.9 bits (63), Expect = 5.2
Identities = 21/64 (32%), Positives = 32/64 (49%), Gaps = 5/64 (7%)
Query: 337 VERVWFEVMARVEDSRLKPLSIKKVKPF-----IESDSVSWANLMSNLSYTKLRPVLLPP 391
V ++ ++ V S LKP K F ++S VS +NL S + + +R V+LPP
Sbjct: 475 VGQINLDIQLTVNSSYLKPRIEDLSKIFSKELDVKSSQVSLSNLTSKGNESLVRMVVLPP 534
Query: 392 EALT 395
E T
Sbjct: 535 EPST 538
>At5g13350 auxin-responsive - like protein
Length = 587
Score = 28.5 bits (62), Expect = 6.8
Identities = 11/17 (64%), Positives = 14/17 (81%)
Query: 105 NKFTTTPKPDLASLVDQ 121
+KF T P PDLASL++Q
Sbjct: 260 SKFLTAPNPDLASLIEQ 276
>At1g76120 unknown protein
Length = 463
Score = 28.1 bits (61), Expect = 8.9
Identities = 26/107 (24%), Positives = 43/107 (39%), Gaps = 12/107 (11%)
Query: 88 KNNFTVKFSDEISSWNHNKFTTTPKPD-----------LASLVDQLSSIAFLDYIKLEGP 136
K F+ S + S+N + FTT K D A+ V L I F+ L
Sbjct: 265 KERFSRILSCYVGSYNFHNFTTRTKADDPTANRQIISFTANTVINLDGIDFIKCEVLGKS 324
Query: 137 FEL-RVHESHHLSLSLPMNVSYNGLKHIIVGKGITVEVRRAREISFY 182
F L ++ + L++++ N + L K + + V A E+ Y
Sbjct: 325 FMLHQIRKMMGLAVAIMRNCASESLIQSAFSKDVNITVPMAPEVGLY 371
>At1g29860 putative protein
Length = 282
Score = 28.1 bits (61), Expect = 8.9
Identities = 19/57 (33%), Positives = 26/57 (45%)
Query: 104 HNKFTTTPKPDLASLVDQLSSIAFLDYIKLEGPFELRVHESHHLSLSLPMNVSYNGL 160
HN T+ + SL D L S +DY LE F+ + S S+S +N Y L
Sbjct: 7 HNYNTSLEEVHFKSLSDCLQSSLVMDYNSLEKVFKFSPYSSPFQSVSPSVNNPYLNL 63
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.322 0.137 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,699,917
Number of Sequences: 26719
Number of extensions: 360871
Number of successful extensions: 994
Number of sequences better than 10.0: 11
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 981
Number of HSP's gapped (non-prelim): 15
length of query: 400
length of database: 11,318,596
effective HSP length: 101
effective length of query: 299
effective length of database: 8,619,977
effective search space: 2577373123
effective search space used: 2577373123
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 61 (28.1 bits)
Medicago: description of AC148404.10


