

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148360.3 - phase: 0
(301 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
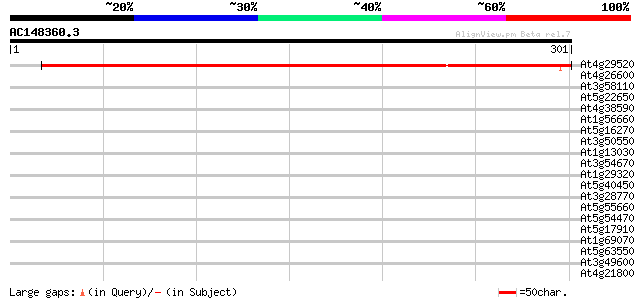
Score E
Sequences producing significant alignments: (bits) Value
At4g29520 unknown protein 384 e-107
At4g26600 unknown protein 41 0.001
At3g58110 putative protein 40 0.002
At5g22650 putative histone deacetylase (HD2B) 39 0.003
At4g38590 galactosidase like protein 39 0.004
At1g56660 hypothetical protein 39 0.004
At5g16270 RAD21-3 38 0.006
At3g50550 unknown protein 37 0.010
At1g13030 unknown protein 37 0.017
At3g54670 structural maintenance of chromosomes (SMC) - like pro... 36 0.022
At1g29320 unknown protein 36 0.022
At5g40450 unknown protein 36 0.029
At3g28770 hypothetical protein 36 0.029
At5g55660 putative protein 35 0.038
At5g54470 putative protein 35 0.038
At5g17910 putative protein 35 0.038
At1g69070 hypothetical protein 35 0.038
At5g63550 unknown protein 35 0.050
At3g49600 putative protein 35 0.050
At4g21800 unknown protein 34 0.085
>At4g29520 unknown protein
Length = 306
Score = 384 bits (987), Expect = e-107
Identities = 185/286 (64%), Positives = 228/286 (79%), Gaps = 3/286 (1%)
Query: 18 IPLSHCAKKPVGIARKEDVPYIKCQVCEILAKQLYQQVQSKKAEISPKKISEYQIIEIAE 77
+P+S AKKP RKEDVPYIKCQVCE L+ +L+Q V+ K+ +ISPKKISEY+IIEIAE
Sbjct: 22 LPVSDAAKKPSSTPRKEDVPYIKCQVCEKLSSRLHQLVKEKQQQISPKKISEYEIIEIAE 81
Query: 78 NVCNLKKVEADWILRIDIVEKADRLELEEHDSEGQCNSECKTVERACQEVMGYSDTDVAE 137
NVCNLKK EADW+L+IDIVEK D L L E EG CNS+CKT+E ACQ+V+GYSDTDVAE
Sbjct: 82 NVCNLKKEEADWMLKIDIVEKGDNLVLVEQQEEGMCNSKCKTIENACQKVIGYSDTDVAE 141
Query: 138 YLYSSKPDIDSLTNYLCKDLSKACNTKPPPVPKDRTPGEPFVAKSSKEAEMEKLLKSMEG 197
Y+Y SKPD+ SL N+LCKDL+ AC+ KPPPVPKDR PGEPFVAK SK+AEM+K+L+SM+G
Sbjct: 142 YIYKSKPDLVSLVNHLCKDLTDACSKKPPPVPKDRVPGEPFVAKPSKDAEMDKILRSMQG 201
Query: 198 MPGAPGMKMYSRDDLMKKNFGAENEDDDEDEDEEEDEADFPSKLGKILKSKENEKGDWKQ 257
MPGAPGMK+YSR+D+ K N G E++D D+DEDEEED+ FP LGK+LK KE++ + K+
Sbjct: 202 MPGAPGMKVYSREDIEKGNIGNEDDDGDDDEDEEEDD-KFPKNLGKVLKEKESKTEELKK 260
Query: 258 KIRKGIVDTSTTLKKHATKVSNHIQRWWKGKKTTSSK--KNSKSEL 301
I K LK+HA KVSN ++RWWKG ++SSK K+ KSEL
Sbjct: 261 TITKEFKKKGEALKRHAQKVSNRVRRWWKGLGSSSSKKPKSGKSEL 306
>At4g26600 unknown protein
Length = 671
Score = 40.8 bits (94), Expect = 0.001
Identities = 28/102 (27%), Positives = 50/102 (48%), Gaps = 10/102 (9%)
Query: 141 SSKPDIDSLTNYLCKDLSKACNTKPPPVPKDRT------PGEPFVAKSSKEAEMEKLLKS 194
S P ++ T L KA T+ PP+ K R P + + +E E E+ +S
Sbjct: 14 SQTPPLNKQTK--ASPLKKAAKTQKPPLKKQRKCISEKKPLKKPEVSTDEEEEEEENEQS 71
Query: 195 MEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDEDEEEDEAD 236
EG G ++S D N +++DDD+D+D+++++A+
Sbjct: 72 DEGSES--GSDLFSDGDEEGNNDSDDDDDDDDDDDDDDEDAE 111
Score = 32.7 bits (73), Expect = 0.25
Identities = 27/102 (26%), Positives = 49/102 (47%), Gaps = 15/102 (14%)
Query: 153 LCKDLSKACNTKPPPVPKDRTPGEPFVAKSSKEAEMEKLLKSMEGMPGAPGMKMYSRDDL 212
L ++ K N++ PP+ K +K + ++K K+ + P K S
Sbjct: 4 LTRNKKKGSNSQTPPLNKQ-----------TKASPLKKAAKTQKP-PLKKQRKCISEKKP 51
Query: 213 MKKNFGAENEDDDEDEDEEEDEADFPSKLGKILKSKENEKGD 254
+KK E D+E+E+EE +++D S+ G L S +E+G+
Sbjct: 52 LKK---PEVSTDEEEEEEENEQSDEGSESGSDLFSDGDEEGN 90
>At3g58110 putative protein
Length = 784
Score = 40.0 bits (92), Expect = 0.002
Identities = 27/86 (31%), Positives = 44/86 (50%), Gaps = 7/86 (8%)
Query: 209 RDDLMKKNFGAENEDDDEDEDEEEDEADFPSKLGK--ILKSKENEKGDWKQKIRKGIVDT 266
++DL+K+ A++E+DD+D+D++ E DF K K L+ KE + G + G VD
Sbjct: 304 KEDLLKEKCKADDEEDDDDDDDDVKEVDFLVKSPKEDCLEVKEEDVGAADSRKDDGAVDL 363
Query: 267 STTLKKHATKVSNHIQRWWKGKKTTS 292
K V H+ G++T S
Sbjct: 364 -----KEDKYVEEHMLELNLGQETVS 384
>At5g22650 putative histone deacetylase (HD2B)
Length = 306
Score = 38.9 bits (89), Expect = 0.003
Identities = 43/170 (25%), Positives = 66/170 (38%), Gaps = 16/170 (9%)
Query: 132 DTDVAEYLYSSKPDIDSLTNYLCKDLSKACNTKPPPVPKDRTPGEPFVAKSSKEAEMEKL 191
D DV E + + P + + KA +TKP P + P E +S +E E +
Sbjct: 106 DEDVPEAVPAPAPTAVTANGNAGAAVVKA-DTKPKAKPAEVKPAEE-KPESDEEDESDDE 163
Query: 192 LKSMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDEDEEEDEADFPSKLGKILKSKENE 251
+S E GM + D ++DD+E++ E+E+E + P K I K + NE
Sbjct: 164 DESEEDDDSEKGMDVDEDD----------SDDDEEEDSEDEEEEETPKKPEPINKKRPNE 213
Query: 252 KGDWK----QKIRKGIVDTSTTLKKHATKVSNHIQRWWKGKKTTSSKKNS 297
+K + ST K K H KK S N+
Sbjct: 214 SVSKTPVSGKKAKPAAAPASTPQKTEEKKKGGHTATPHPAKKGGKSPVNA 263
>At4g38590 galactosidase like protein
Length = 1036
Score = 38.5 bits (88), Expect = 0.004
Identities = 18/43 (41%), Positives = 28/43 (64%)
Query: 209 RDDLMKKNFGAENEDDDEDEDEEEDEADFPSKLGKILKSKENE 251
+D+ KK E E+DDED+DEEE+E D +K K +++K +
Sbjct: 822 QDEKKKKEDKDEEEEDDEDDDEEEEEEDKENKDTKDMENKNQD 864
Score = 31.6 bits (70), Expect = 0.55
Identities = 16/50 (32%), Positives = 27/50 (54%), Gaps = 2/50 (4%)
Query: 205 KMYSRDDLMKKNFGAENEDDDEDEDEEEDEADFPSKLGKILKSKENEKGD 254
K + D KK + E++D+++D+EE+E + K K K EN+ D
Sbjct: 817 KKEGKQDEKKKKEDKDEEEEDDEDDDEEEEEE--DKENKDTKDMENKNQD 864
>At1g56660 hypothetical protein
Length = 522
Score = 38.5 bits (88), Expect = 0.004
Identities = 38/185 (20%), Positives = 70/185 (37%), Gaps = 4/185 (2%)
Query: 97 EKADRLELEEHDSEGQCNSECKTVERACQEVMGYSDTDVAEYLYSSKPDIDSLTNYLCKD 156
E AD E ++ ++ + + ++ C++ D D E + + KD
Sbjct: 317 EAADHKEGKKKKNKDKAKKKETVIDEVCEKETKDKDDDEGETKQKKNKKKEKKSEKGEKD 376
Query: 157 LSKACNTKPP----PVPKDRTPGEPFVAKSSKEAEMEKLLKSMEGMPGAPGMKMYSRDDL 212
+ + + P + +D EP K ++ EK +EG G K +D
Sbjct: 377 VKEDKKKENPLETEVMSRDIKLEEPEAEKKEEDDTEEKKKSKVEGGESEEGKKKKKKDKK 436
Query: 213 MKKNFGAENEDDDEDEDEEEDEADFPSKLGKILKSKENEKGDWKQKIRKGIVDTSTTLKK 272
K + EDE+E++D++ G K ++ +K K+K I T L K
Sbjct: 437 KNKKKDTKEPKMTEDEEEKKDDSKDVKIEGSKAKEEKKDKDVKKKKGGNDIGKLKTKLAK 496
Query: 273 HATKV 277
K+
Sbjct: 497 IDEKI 501
Score = 34.3 bits (77), Expect = 0.085
Identities = 63/279 (22%), Positives = 109/279 (38%), Gaps = 37/279 (13%)
Query: 45 EILAKQLYQQVQSKKAEISP--------KKISEYQIIEIAENVCNLKKVEADWILRI--- 93
E+ AK + ++V++KK E S KK + E+ E+ + KK + + +
Sbjct: 36 EVKAKSI-EKVKAKKDEESSGKSKKDKEKKKGKNVDSEVKEDKDDDKKKDGKMVSKKHEE 94
Query: 94 ---DIVEKADRLELEEHDSEGQCNSECKTVERACQEVMGYSDTDVAEYLYSSKPDIDSLT 150
D+ K +++EEH+ E + E K E +E G + E S + +
Sbjct: 95 GHGDLEVKESDVKVEEHEKEHKKGKE-KKHEELEEEKEGKKKKNKKEKDESGPEEKNKKA 153
Query: 151 NYLCK---------DLSKACNTKPPPVPKDRTPGEPFVAKSSKEAEMEKLLKSMEGMPGA 201
+ K +L + K KD + E K KE + ++ KS E
Sbjct: 154 DKEKKHEDVSQEKEELEEEDGKKNKKKEKDESGTEEKKKKPKKEKKQKEESKSNEDKKVK 213
Query: 202 PGMKMYSRDDLMKKNFGAENEDDDEDEDEEEDEADFPSKLGKILKSKENEKGDWKQKIRK 261
+ + DL E +DE++ +E DE D K K+K+ EK + + +K
Sbjct: 214 GKKEKGEKGDL---------EKEDEEKKKEHDETDQEMKEKDSKKNKKKEKDESCAEEKK 264
Query: 262 GIVDTSTTLKKHATKVSNHIQRWWKGKKTTSSKKNSKSE 300
D K +T+ + + KGKK K + E
Sbjct: 265 KKPDKEKKEKDESTEKED---KKLKGKKGKGEKPEKEDE 300
Score = 33.9 bits (76), Expect = 0.11
Identities = 58/267 (21%), Positives = 99/267 (36%), Gaps = 31/267 (11%)
Query: 48 AKQLYQQVQSKKAEISPKKISEYQIIEIAENVCNLKKVEADWILRIDIVEKADRLELE-- 105
AK+ V+ K E+ PK+ E +E+ +++KV+A K D+ + +
Sbjct: 8 AKEEKLHVKIKTQELDPKEKGENVEVEMEVKAKSIEKVKAKKDEESSGKSKKDKEKKKGK 67
Query: 106 ------EHDSEGQCNSECKTVERACQEVMG-----YSDTDVAEYLYSSKPDIDSLTNYLC 154
+ D + + K V + +E G SD V E+ K + L
Sbjct: 68 NVDSEVKEDKDDDKKKDGKMVSKKHEEGHGDLEVKESDVKVEEHEKEHKKGKEKKHEEL- 126
Query: 155 KDLSKACNTKPPPVPKDRTPGEPFVAKSSKEAEMEKLLKSMEGMPGAPGMKMYSRDDLMK 214
+ K K KD + E K+ KE + E + + E ++
Sbjct: 127 -EEEKEGKKKKNKKEKDESGPEEKNKKADKEKKHEDVSQEKEE---------------LE 170
Query: 215 KNFGAENEDDDEDEDEEEDEADFPSKLGKIL-KSKENEKGDWKQKIRKGIVDTSTTLKKH 273
+ G +N+ ++DE E++ P K K +SK NE K K KG +
Sbjct: 171 EEDGKKNKKKEKDESGTEEKKKKPKKEKKQKEESKSNEDKKVKGKKEKGEKGDLEKEDEE 230
Query: 274 ATKVSNHIQRWWKGKKTTSSKKNSKSE 300
K + + K K + +KK K E
Sbjct: 231 KKKEHDETDQEMKEKDSKKNKKKEKDE 257
Score = 30.4 bits (67), Expect = 1.2
Identities = 51/253 (20%), Positives = 94/253 (36%), Gaps = 22/253 (8%)
Query: 53 QQVQSKKAEISPKKISEYQIIEIAENVCNLKKVEADWILRIDIVEKADRL--------EL 104
Q+++ K ++ + KK + E + + +K E D + EK D+ E
Sbjct: 240 QEMKEKDSKKNKKKEKDESCAEEKKKKPDKEKKEKD-----ESTEKEDKKLKGKKGKGEK 294
Query: 105 EEHDSEGQCNSECKTVERACQEVMGYSDTDVAEYLYSSKPDIDSLTNYLCKDLSKACN-- 162
E + EG+ E E QE+ D + A++ K + + C
Sbjct: 295 PEKEDEGKKTKEHDATE---QEM----DDEAADHKEGKKKKNKDKAKKKETVIDEVCEKE 347
Query: 163 TKPPPVPKDRTPGEPFVAKSSKEAEMEKLLKSMEGMPGAPGMKMYSRDDLMKKNFGAENE 222
TK + T + K K + EK +K + ++ SRD +++ + E
Sbjct: 348 TKDKDDDEGETKQKKNKKKEKKSEKGEKDVKEDKKKENPLETEVMSRDIKLEEPEAEKKE 407
Query: 223 DDDEDEDEEEDEADFPSKLGKILKSKENEKGDWKQKIRKGIVDTSTTLKKHATKVSNHIQ 282
+DD +E ++ S+ GK K K+ +K K + + K + V
Sbjct: 408 EDDTEEKKKSKVEGGESEEGKKKKKKDKKKNKKKDTKEPKMTEDEEEKKDDSKDVKIEGS 467
Query: 283 RWWKGKKTTSSKK 295
+ + KK KK
Sbjct: 468 KAKEEKKDKDVKK 480
>At5g16270 RAD21-3
Length = 1031
Score = 38.1 bits (87), Expect = 0.006
Identities = 50/211 (23%), Positives = 83/211 (38%), Gaps = 39/211 (18%)
Query: 64 PKKISEYQIIEIAENVCNLKKVEADWILRIDIVEKADRL--ELEEHDSEGQCNSECKTVE 121
P + S +IEIAE ++ + + L+++ D L E E+ D+ + + + +
Sbjct: 771 PNEKSCTDVIEIAEGDTDINPIFNEMDLKVE-----DELPHEDEKTDASAEVSELGRDDQ 825
Query: 122 RACQEVMGYSDTDVAEYLYSSKPDIDSLTNYLCKDLSKACNTKPPPVPKDRTPGEPFVAK 181
C +G ++T E S +++ L + S N P + EP
Sbjct: 826 TPCDNTVGSTETGCLEAGDLSNMALENCNEPLVEANSDGLN----PETESYNKYEPHNEM 881
Query: 182 SSKEAEMEKLLKSMEGMPGAPGMKMYSRDDLMKKN-----------FGAENEDDDE-DED 229
S++EA M+ L + SRD LM N G N DDDE DED
Sbjct: 882 SNEEASMQNALDG----------EHTSRDGLMGDNDEMDTMENAHDTGFLNVDDDEVDED 931
Query: 230 EEEDEADFPSKLGKILKSKENEKGDWKQKIR 260
EED+ + +++ E W + R
Sbjct: 932 HEEDDIQYDD------ETRLLENSGWSSRTR 956
>At3g50550 unknown protein
Length = 95
Score = 37.4 bits (85), Expect = 0.010
Identities = 17/45 (37%), Positives = 29/45 (63%), Gaps = 2/45 (4%)
Query: 208 SRDDLMKKNFGAENED--DDEDEDEEEDEADFPSKLGKILKSKEN 250
S DD N ++++D DD+DE+EEE+E K+ ++LK K++
Sbjct: 51 SEDDYTDSNSDSDDDDEEDDDDEEEEEEEDSLVDKVTRLLKGKQD 95
>At1g13030 unknown protein
Length = 608
Score = 36.6 bits (83), Expect = 0.017
Identities = 20/99 (20%), Positives = 46/99 (46%)
Query: 202 PGMKMYSRDDLMKKNFGAENEDDDEDEDEEEDEADFPSKLGKILKSKENEKGDWKQKIRK 261
PG + + ++ K+ G E+E ++++ +EE +E K K K+ + ++K +
Sbjct: 122 PGEMLLANEEFQKETGGYESESEEDELEEEAEEFVPEKKASKKRKTSSKNQSTKRKKCKL 181
Query: 262 GIVDTSTTLKKHATKVSNHIQRWWKGKKTTSSKKNSKSE 300
+ S +++ VSN +++ K K N+ +
Sbjct: 182 DTTEESPDERENTAVVSNVVKKKKKKKSLDVQSANNDEQ 220
>At3g54670 structural maintenance of chromosomes (SMC) - like
protein
Length = 1265
Score = 36.2 bits (82), Expect = 0.022
Identities = 45/219 (20%), Positives = 86/219 (38%), Gaps = 32/219 (14%)
Query: 82 LKKVEADWILRIDIVEKADRLELEEHDSEGQCNSECKTVERACQEVMGYSDTDVAEYLYS 141
LKK + D+ +++ + ++++E + G+ + K ++ A E+ S D L
Sbjct: 707 LKKNKEDFEQQLENIGSIREMQMKESEISGKISGLEKKIQYA--EIEKKSIKDKLPQLEQ 764
Query: 142 SKPDIDSLTNYLCKDLSKACNTKPPPVPKDRTPGEPFVAKSSKEAEMEKLLKSMEGMPGA 201
+ +I + + +LSKA D+ E + ++++ K G
Sbjct: 765 EERNIIEEIDRIKPELSKAIAR----TEVDKRKTEMNKLEKRMNEIVDRIYKDFSQSVGV 820
Query: 202 PGMKMYSRDDLMKKNFGAENEDDDEDEDEEEDEADFPSKLGKILKSKENEKG-DWKQKIR 260
P +++Y L E E E+ + ++L K+ E E+ D +IR
Sbjct: 821 PNIRVYEETQLKTA------------EKEAEERLELSNQLAKLKYQLEYEQNRDVGSRIR 868
Query: 261 K-------------GIVDTSTTLKKHATKVSNHIQRWWK 286
K GI T + K+ A K++N I W K
Sbjct: 869 KIESSISSLETDLEGIQKTMSERKETAVKITNEINNWKK 907
>At1g29320 unknown protein
Length = 468
Score = 36.2 bits (82), Expect = 0.022
Identities = 29/105 (27%), Positives = 49/105 (46%), Gaps = 5/105 (4%)
Query: 179 VAKSSKEAEMEKLLKSMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDEDEEEDEADFP 238
VA+++ E +M + + + AP + S+ + + E EDD+E+EDE E
Sbjct: 366 VAEAATEEKMTIMDQEDDETEKAPVKRKKSKKEKRSREIVFEGEDDEENEDEIEKAPVKT 425
Query: 239 SKLGKILKSKEN-EKGDWKQKIRKGIVDTSTTLKKHATKVSNHIQ 282
K K +S+E +G+ K ++R T K TK+ H Q
Sbjct: 426 KKSKKEKRSREKVSEGEEKDELR----SKKTREHKKKTKIVKHNQ 466
>At5g40450 unknown protein
Length = 2910
Score = 35.8 bits (81), Expect = 0.029
Identities = 45/223 (20%), Positives = 91/223 (40%), Gaps = 33/223 (14%)
Query: 97 EKADRLELEEHDSEGQCNSECKTVERACQEVMGYSDTDVAEYLYSSKPDIDSLTNYLCKD 156
E A +E EE E + E+ + + + + + ++P+I + +
Sbjct: 2697 EAAKEIEQEEGKQTNIVKEEIREEEKEINQESFNNVKETDDAIDKTQPEIRDI-----ES 2751
Query: 157 LSKACNTKPPPVPKDRTPGE---------PFVAKSSKEAEMEK-------------LLKS 194
LS T+ P P+ P + P + S E E++K ++K
Sbjct: 2752 LSSVSKTQDKPEPEYEVPNQQKREITNEVPSLENSKIEEELQKKDEESENTKDLFSVVKE 2811
Query: 195 MEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDEDEEEDEADFPSKLGKILKSKENEKGD 254
E P K S D + K+ E+E+DDED + ++D+ S + ++++K+
Sbjct: 2812 TEPTLKEPARKSLS-DHIQKEPKTEEDENDDEDHEHKDDKTSPDSIV--MVEAKDTVSIV 2868
Query: 255 WKQKIRKGIVDTSTTLKKHATKVSNHIQRWWKGKKTTSSKKNS 297
K GI+ + KH+ + +++ GK + ++K +S
Sbjct: 2869 KTHKKSHGILSGVGSKVKHSI---SKVKKVLTGKSSHTTKPSS 2908
Score = 31.6 bits (70), Expect = 0.55
Identities = 38/185 (20%), Positives = 72/185 (38%), Gaps = 17/185 (9%)
Query: 53 QQVQSKKAEISPKKISEYQIIEIAENVCNLKKVEADWILRIDIVEKADRLELEEHDSEGQ 112
++V +K I + +E E++ N+ KV D ++++E+D E
Sbjct: 143 EEVSVEKPVIEEDQTEAKHSLEQEEDIGNISKVLTD----------TTPVKVDEYDIEKS 192
Query: 113 CNSECKTVERACQEVMGYSDTDVAEYLYSSKPDIDSLTNYLCKDLSKACNTKPPPVPKDR 172
NS C+ + EV +D+ E + + T + + NT V +D+
Sbjct: 193 LNSVCEEIPIKTDEVREETDSRTVETSVNGTEAEHNATVSVEEISRNGDNTVNETVSEDQ 252
Query: 173 --TPGEPFVAKSSKEAEMEKLLKSMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDEDE 230
T GEP + + E E K++ K+ + ++ ED+ ++E
Sbjct: 253 TATDGEPLHDVETIKREAEPFYKTV-----VEDAKIVNTEETTAHESKILKEDNHQEEYA 307
Query: 231 EEDEA 235
E EA
Sbjct: 308 ESVEA 312
>At3g28770 hypothetical protein
Length = 2081
Score = 35.8 bits (81), Expect = 0.029
Identities = 46/217 (21%), Positives = 89/217 (40%), Gaps = 23/217 (10%)
Query: 94 DIVEKADRLELEEHDSEG-QCNSECKTVERACQEVMGYSDTDVAEYLYSSKPDIDSLTNY 152
++ K D+ EL++ +S G + N+E E+ Q G+ ++ ++ + + + DS
Sbjct: 601 NLENKEDKKELKDDESVGAKTNNETSLEEKREQTQKGHDNSINSKIVDNKGGNADSNKE- 659
Query: 153 LCKDLSKACNTKPPPVP-KDRTPGEPFVAKSSKEAEMEKLLKSMEGMPGAPGMKMYSRDD 211
K++ +T + K+ T E V K+ +E +G G K D
Sbjct: 660 --KEVHVGDSTNDNNMESKEDTKSEVEVKKNDGSSE--------KGEEGKENNKDSMEDK 709
Query: 212 LMKKNFGAENEDDDEDEDEEEDEADF----------PSKLGKILKSKENEKGDWKQKIRK 261
++ + DD+ D++++EA GK +SKEN+K + +
Sbjct: 710 KLENKESQTDSKDDKSVDDKQEEAQIYGGESKDDKSVEAKGKKKESKENKKTKTNENRVR 769
Query: 262 GIVDTSTTLKKHATKVSNHIQRWWKGKKTTSSKKNSK 298
+ KK + KV ++ K K+ +K N K
Sbjct: 770 NKEENVQGNKKESEKVEKGEKKESKDAKSVETKDNKK 806
Score = 29.6 bits (65), Expect = 2.1
Identities = 51/222 (22%), Positives = 84/222 (36%), Gaps = 28/222 (12%)
Query: 98 KADRLELEEHDSEGQCNSECKTVERACQEVMGYSDTDVAEYLYSSKPDIDS--------- 148
K ++ +E D + N+E K E +E ++ + E +K +S
Sbjct: 952 KNSNMKKKEEDKKEYVNNELKKQEDNKKETTKSENSKLKEENKDNKEKKESEDSASKNRE 1011
Query: 149 ---LTNYLCKDLSKACNTKPPPVPKDRTPGEPFVAKSSKEAEMEKLLKSMEGMPGAPGMK 205
K +A K K R + KS KE E + LK+ + K
Sbjct: 1012 KKEYEEKKSKTKEEAKKEKKKSQDKKREEKDSEERKSKKEKEESRDLKAKKKEEETKEKK 1071
Query: 206 MYSRDDLMKKNFGAENEDD---DEDEDEEEDEADFPSKLGKILKSKENEKGDWKQKIRKG 262
KK E+ED+ ++ED++E + SK K KE +K D ++
Sbjct: 1072 ESENHKSKKKEDKKEHEDNKSMKKEEDKKEKKKHEESKSRK----KEEDKKDMEK----- 1122
Query: 263 IVDTSTTLK---KHATKVSNHIQRWWKGKKTTSSKKN-SKSE 300
+ D ++ K K+ K S H++ K K+N KSE
Sbjct: 1123 LEDQNSNKKKEDKNEKKKSQHVKLVKKESDKKEKKENEEKSE 1164
Score = 27.7 bits (60), Expect = 7.9
Identities = 28/120 (23%), Positives = 56/120 (46%), Gaps = 15/120 (12%)
Query: 175 GEPFVAKSSKEAEME-KLLKSMEGMPGAPGMKMYSRDD------LMKKNFGAENEDDDED 227
G+ ++SK E++ + +S +G G K S ++ ++++N G E+ +
Sbjct: 1595 GDEVGKENSKTIEVKGRHEESKDGKTNENGGKEVSTEEGSKDSNIVERNGGKEDSIKEGS 1654
Query: 228 EDE--------EEDEADFPSKLGKILKSKENEKGDWKQKIRKGIVDTSTTLKKHATKVSN 279
ED EE + SK GKI + KE ++ K+ + ++ K+++TK S+
Sbjct: 1655 EDGKTVEINGGEELSTEEGSKDGKIEEGKEGKENSTKEGSKDDKIEEGMEGKENSTKESS 1714
Score = 27.7 bits (60), Expect = 7.9
Identities = 19/84 (22%), Positives = 41/84 (48%), Gaps = 4/84 (4%)
Query: 181 KSSKEAEMEKLLKSMEGMPGAPGMKMYSRDDLMKKNFGAEN---EDDDEDEDEEEDEADF 237
K KE++ + K E ++ ++D K+ +EN +++++D E+++ D
Sbjct: 946 KKKKESKNSNMKKKEEDKKEYVNNELKKQEDNKKETTKSENSKLKEENKDNKEKKESEDS 1005
Query: 238 PSKLGKILKSKENEKGDWKQKIRK 261
SK + K E +K K++ +K
Sbjct: 1006 ASK-NREKKEYEEKKSKTKEEAKK 1028
>At5g55660 putative protein
Length = 759
Score = 35.4 bits (80), Expect = 0.038
Identities = 46/175 (26%), Positives = 74/175 (42%), Gaps = 15/175 (8%)
Query: 86 EADWILRIDIVEKADRLELEEHDSEGQCNSECKTVERACQEVMGYSDTDVAEYLYSSKP- 144
E D +++ + VE AD + E + EG+ E K + + G + D E L
Sbjct: 118 EEDAVMK-ESVESADNKDAE--NPEGEQEKESKEEKLEGGKANGNEEGDTEEKLVGGDKG 174
Query: 145 -DIDS---LTNYLCKDLSKACNTKPPP--VPKDRTPGEPFVAKSSKEAEMEKLLKSMEGM 198
D+D + N D +A K ++ T V +++KE ++E K E
Sbjct: 175 DDVDEAEKVENVDEDDKEEALKEKNEAELAEEEETNKGEEVKEANKEDDVEADTKVAE-- 232
Query: 199 PGAPGMKMYSRDDLMKKNFGAENEDDDEDEDEEEDEADFPSKLGKILKSKENEKG 253
P K S+D+ K E+E +DE E+ +D+ D K I KS + KG
Sbjct: 233 PEVEDKKTESKDENEDKEEEKEDEKEDEKEESNDDKED---KKEDIKKSNKRGKG 284
Score = 30.4 bits (67), Expect = 1.2
Identities = 43/183 (23%), Positives = 73/183 (39%), Gaps = 27/183 (14%)
Query: 132 DTDVAEYLYSSKPDIDSLTNYLCKDLSKA---CNTKPPPVPKDRTP--GEPFVAKSSKEA 186
D VA+ + + L +L K + N K V + RTP P SS +
Sbjct: 425 DISVAKATTKKEDIVTKLVEFLEKPHATTDVLVNEKEKGVKRKRTPKKSSPAAGSSSSKR 484
Query: 187 EMEKLLKSMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDEDEEE------------DE 234
+ K+ E +S D+ ++ E E+ +++ +EEE DE
Sbjct: 485 SAKSQKKTEEATRTNKKSVAHSDDESEEEKEDDEEEEKEQEVEEEEEENENGIPDKSEDE 544
Query: 235 ADFPSKLGKILKSKENEKGDWKQKIRKGIVDTSTTLKKHATKVSNHIQRWWKGKKTTSSK 294
A S+ + ++S+E + + K+K R TS+ K+ A K + KKT
Sbjct: 545 APQLSESEENVESEEESEEETKKKKRGS--RTSSDKKESAGKS--------RSKKTAVPT 594
Query: 295 KNS 297
K+S
Sbjct: 595 KSS 597
>At5g54470 putative protein
Length = 229
Score = 35.4 bits (80), Expect = 0.038
Identities = 38/175 (21%), Positives = 62/175 (34%), Gaps = 15/175 (8%)
Query: 118 KTVERACQEVMGYSDTDVAEYLYSSKPDIDSLTNYLCKD----LSKACNTKPPPVPKDRT 173
K E C Y ++D A + + + K L AC + P
Sbjct: 4 KKCELCCGVARMYCESDQASLCWDCDGKVHGANFLVAKHMRCLLCSACQSHTPWKASGLN 63
Query: 174 PGEPFVAKSSKEAEMEKLLKSMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDEDEEED 233
G S A + S+ G + ++++ N GAE+ D++ DEDEEE+
Sbjct: 64 LGPTVSICESCLARKKNNNSSLAGRD----QNLNQEEEIIGCNDGAESYDEESDEDEEEE 119
Query: 234 E-------ADFPSKLGKILKSKENEKGDWKQKIRKGIVDTSTTLKKHATKVSNHI 281
E A +L + S G+ Q +++ +D L K S I
Sbjct: 120 EVENQVVPAAVEQELPVVSSSSSVSSGEGDQVVKRTRLDLDLNLSDEMIKKSETI 174
>At5g17910 putative protein
Length = 1342
Score = 35.4 bits (80), Expect = 0.038
Identities = 20/64 (31%), Positives = 35/64 (54%), Gaps = 2/64 (3%)
Query: 220 ENEDDDEDEDEEEDEADFPSKLGKILKSKENEKGDWKQKIRKGIVDTSTTLKKHATKVSN 279
ENED++EDE+EE++E K K +SK K W + ++ ++D + + ++ N
Sbjct: 324 ENEDEEEDEEEEDEEEKQEKKEDKDDESKSAIK--WTEADQRNVMDLGSLELERNQRLEN 381
Query: 280 HIQR 283
I R
Sbjct: 382 LIAR 385
>At1g69070 hypothetical protein
Length = 901
Score = 35.4 bits (80), Expect = 0.038
Identities = 20/55 (36%), Positives = 32/55 (57%), Gaps = 1/55 (1%)
Query: 218 GAENEDDDEDEDEEEDEADFPSKLGKILKSKENEKGDWKQKIRKGIVDTSTTLKK 272
GAE ED++ED+DEE+D+ + ++L K K + +K + G V T +KK
Sbjct: 401 GAELEDEEEDDDEEDDDEE-DAELRVHKKLKNDYAAPYKGEGLSGTVKEKTNMKK 454
Score = 33.1 bits (74), Expect = 0.19
Identities = 17/51 (33%), Positives = 28/51 (54%), Gaps = 3/51 (5%)
Query: 210 DDLMKKNFGAENEDDDED---EDEEEDEADFPSKLGKILKSKENEKGDWKQ 257
DD++++ +N + DED E EEE++ D S G + K + DW+Q
Sbjct: 345 DDVLEREDNVDNSESDEDEDSESEEEEDDDGESDGGDEKQRKGHHLEDWEQ 395
>At5g63550 unknown protein
Length = 530
Score = 35.0 bits (79), Expect = 0.050
Identities = 41/175 (23%), Positives = 64/175 (36%), Gaps = 13/175 (7%)
Query: 135 VAEYLYSSKPDIDSLTNYLCKDLSKACNTKPPPVPKDRTPGEPFVAKSSKEAEMEKLLKS 194
V E+L S K D + D KA K P AK ++ + L
Sbjct: 232 VLEFLESPKETRD----VIIADQEKAKKRKSTPKRGKSGESSDTPAKRKRQTKKRDLPSD 287
Query: 195 ME-----GMPGAPGMKMYSRDDLMKKNFGAENEDDDEDEDEEEDEADFPSK----LGKIL 245
E G + G +D +++E D D++++E E + PSK K +
Sbjct: 288 TEEGKDEGDADSEGTNDPHEEDDAAPEEESDHEKTDTDDEKDEVEVEKPSKKKSSSKKTV 347
Query: 246 KSKENEKGDWKQKIRKGIVDTSTTLKKHATKVSNHIQRWWKGKKTTSSKKNSKSE 300
+ KG KQ KG + K K ++ + K SSK+ SK +
Sbjct: 348 EESSGSKGKDKQPSAKGSARSGEKSSKQIAKSTSSPAKKQKVDHVESSKEKSKKQ 402
Score = 30.0 bits (66), Expect = 1.6
Identities = 26/95 (27%), Positives = 39/95 (40%), Gaps = 6/95 (6%)
Query: 158 SKACNTKPPPVPKDRTPGEPFVAKSSKEAEMEKLLKSMEGMPGAPGMKMYSRDDLMKKNF 217
++ + K P V V K KE E EK+ G + + +
Sbjct: 3 TETLDEKTPEVNSPAKEEIDVVPKEEKEVEKEKVDSPRIGEAEEEKKEDEEEGEAKEGEL 62
Query: 218 GAENEDDD-EDEDEEEDEADFPSKLGKILKSKENE 251
G ++++DD E E+EEE+E SK KS E E
Sbjct: 63 GEKDKEDDVESEEEEEEEEGSGSK-----KSSEKE 92
>At3g49600 putative protein
Length = 1672
Score = 35.0 bits (79), Expect = 0.050
Identities = 49/200 (24%), Positives = 78/200 (38%), Gaps = 18/200 (9%)
Query: 100 DRLELEEHDSEGQCNSECKTVERACQEVMGYSDTDVAEYLYSSKPDIDSLTNYLCKDLSK 159
D E +EH + + S+ K R + S +D S+ D DS D K
Sbjct: 225 DSSESDEHGRDRRRRSKKKAKGRKQESESDSSSSD-------SESDSDS------DDGKK 271
Query: 160 ACNTKPPPVPKDRTPGEPFVAKSSKEAEME---KLLKS-MEGMPGAPGMKMYSRDDLMKK 215
KP K R+ + V+ S+E E + KL KS + +P RD ++
Sbjct: 272 RGRKKPTKTTKKRSRRKRSVSSESEEVESDDSKKLRKSHKKSLPSNRSGSKELRDKHDEQ 331
Query: 216 N-FGAENEDDDEDEDEEEDEADFPSKLGKILKSKENEKGDWKQKIRKGIVDTSTTLKKHA 274
+ G + D D E E ED K + + + +K D + + D T K A
Sbjct: 332 SRAGRKRHDSDVSEPESEDNKQPLRKKEEAYRGGQKQKRDDEDVEADHLKDRYTRDDKKA 391
Query: 275 TKVSNHIQRWWKGKKTTSSK 294
+ S+ + ++ KK SK
Sbjct: 392 ARDSDDSEIEYQNKKQLRSK 411
>At4g21800 unknown protein
Length = 379
Score = 34.3 bits (77), Expect = 0.085
Identities = 24/66 (36%), Positives = 33/66 (49%), Gaps = 16/66 (24%)
Query: 184 KEAEMEKLLKSMEGMPGAP-----GMK--------MYSRDDLMKKNFGAENEDDDEDEDE 230
K+ EMEKL K ME G G+K M DD ++F E+E+D +D +
Sbjct: 310 KKHEMEKLRKDMESSQGGTVVLNTGLKDRDATEKMMLEEDD---EDFQVEDEEDSDDAID 366
Query: 231 EEDEAD 236
E+DE D
Sbjct: 367 EDDEDD 372
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.311 0.129 0.376
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,370,356
Number of Sequences: 26719
Number of extensions: 349913
Number of successful extensions: 5784
Number of sequences better than 10.0: 293
Number of HSP's better than 10.0 without gapping: 152
Number of HSP's successfully gapped in prelim test: 147
Number of HSP's that attempted gapping in prelim test: 3572
Number of HSP's gapped (non-prelim): 1050
length of query: 301
length of database: 11,318,596
effective HSP length: 99
effective length of query: 202
effective length of database: 8,673,415
effective search space: 1752029830
effective search space used: 1752029830
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 60 (27.7 bits)
Medicago: description of AC148360.3


