

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148360.15 - phase: 0
(422 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
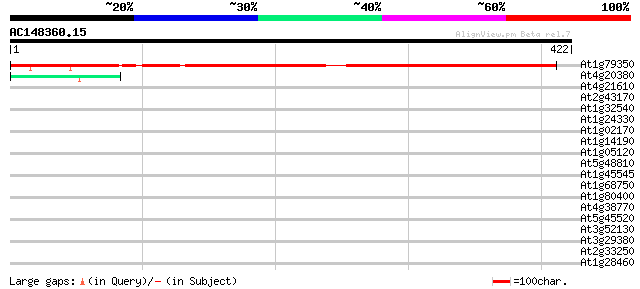
Score E
Sequences producing significant alignments: (bits) Value
At1g79350 hypothetical protein 494 e-140
At4g20380 zinc-finger protein Lsd1 45 6e-05
At4g21610 Lsd1 like protein 41 0.001
At2g43170 unknown protein 39 0.005
At1g32540 zinc-finger containing protein 37 0.016
At1g24330 hypothetical protein 33 0.39
At1g02170 hypothetical protein 32 0.50
At1g14190 putative mandelonitrile lyase 32 0.66
At1g05120 hypothetical protein, 3' partial 32 0.66
At5g48810 cytochrome b5 (dbj|BAA74840.1) 30 1.9
At1g45545 hypothetical protein 30 2.5
At1g68750 putative phosphoenolpyruvate carboxylase 30 3.3
At1g80400 putative RING zinc finger protein 29 5.6
At4g38770 extensin - like protein 28 7.3
At5g45520 putative protein 28 9.5
At3g52130 5B protein like protein 28 9.5
At3g29380 transcription initiation factor IIB, putative 28 9.5
At2g33250 unknown protein 28 9.5
At1g28460 hypothetical protein 28 9.5
>At1g79350 hypothetical protein
Length = 1318
Score = 494 bits (1273), Expect = e-140
Identities = 255/473 (53%), Positives = 316/473 (65%), Gaps = 85/473 (17%)
Query: 1 MNRQP-QPPPPSTAR--------------ARCAGCRTYFSAAQGVAELPCPNCQMPHVF- 44
M + P QPPPP A+ RCAGCR GV E CP CQ+P +
Sbjct: 1 MTQSPVQPPPPLPAQPHSAAGGVIRGDVQVRCAGCRVILRVKTGVVEFSCPTCQLPQMLP 60
Query: 45 ----------------------------------------------FVDSSAVKIRCSSC 58
+D + +++ C++C
Sbjct: 61 PELLSRARPQFPQSPQQPPQPIQTLPPPIQQQLKPLNLPRPPVPAHGIDPTKMQLPCANC 120
Query: 59 KAVVNAPSNLSKFPCPQCHVRIDVHADVEEVNEKIDSEPVISCFHKFSQTVSQELDMLQN 118
+A++N P L++F CPQCHV + V DV ++N + + S H T +
Sbjct: 121 QAILNVPHGLTRFSCPQCHVELAV--DVSKLNRSLTA----SQSHSNPPTPAAPTVPPPP 174
Query: 119 PRETVGKQSVLVNEVEQEEGDGGIAGETFTDYRPSKLSIGSPHPDPIVETSSLSAVQPPE 178
P E V ++++ EVE+EE +GG AGETF DYRP KLSIG PHPDPIVETSSLSAVQPPE
Sbjct: 175 PPEEVNEEAI---EVEREEDEGGTAGETFMDYRPPKLSIGPPHPDPIVETSSLSAVQPPE 231
Query: 179 PTYDPKIKNDLERSKALSCLQIETLVYACQRHLQHVPSGPRAGFFIGDGAGVGKGRTVAG 238
PTYD KIK +LERSKALSCLQIETLVYACQRHLQH+ G RAGFF+GDGAGVGKGRT+AG
Sbjct: 232 PTYDLKIKEELERSKALSCLQIETLVYACQRHLQHLADGTRAGFFVGDGAGVGKGRTIAG 291
Query: 239 LIWENWHHGRRKTLWISVGSDLKFDARRDLDDMGASCISVHALNKLPYTKLDTKSVGVRE 298
WIS+GSDLK+DARRDLDD+GA+C+ V+ LNKLPY+KLD+K+VG++E
Sbjct: 292 --------------WISIGSDLKYDARRDLDDVGATCVGVNPLNKLPYSKLDSKNVGIKE 337
Query: 299 GVIFSTYSSLIASSDRGRTRMQQLVQWCGPKFDGLIIFDECHKAKNLVPEKDKKPTKTGQ 358
GV+F TY+SLIASS++GR+R+QQLVQWCGP+FDGL+IFDECHKAKNLVPE +PT+ GQ
Sbjct: 338 GVVFLTYNSLIASSEKGRSRLQQLVQWCGPEFDGLLIFDECHKAKNLVPEAGSQPTRIGQ 397
Query: 359 AVLDIQAQLPEARVVYCSATGASEPRNMAYMVRLGLWGAGTFFPDFGEFLGSV 411
AV+DIQ ++P+ARV+YCSATGASEPRNM YMVRLGLWGAGT F DF +FLG++
Sbjct: 398 AVVDIQDKIPQARVIYCSATGASEPRNMGYMVRLGLWGAGTSFSDFNKFLGAL 450
>At4g20380 zinc-finger protein Lsd1
Length = 184
Score = 45.4 bits (106), Expect = 6e-05
Identities = 26/93 (27%), Positives = 38/93 (39%), Gaps = 10/93 (10%)
Query: 1 MNRQPQPPPP-STARARCAGCRTYFSAAQGVAELPCPNCQMPHVFFVDSSAV-------- 51
+N P PPPP A C GCRT +G + + C CQ ++ S+ V
Sbjct: 31 INMVPPPPPPHDMAHIICGGCRTMLMYTRGASSVRCSCCQTTNLVPAHSNQVAHAPSSQV 90
Query: 52 -KIRCSSCKAVVNAPSNLSKFPCPQCHVRIDVH 83
+I C C+ + P S C C +V+
Sbjct: 91 AQINCGHCRTTLMYPYGASSVKCAVCQFVTNVN 123
>At4g21610 Lsd1 like protein
Length = 155
Score = 40.8 bits (94), Expect = 0.001
Identities = 17/73 (23%), Positives = 32/73 (43%)
Query: 10 PSTARARCAGCRTYFSAAQGVAELPCPNCQMPHVFFVDSSAVKIRCSSCKAVVNAPSNLS 69
P A+ C CR S +G + C +CQ ++ + ++ C++CK ++ P
Sbjct: 56 PEKAQMVCGSCRRLLSYLRGSKHVKCSSCQTVNLVLEANQVGQVNCNNCKLLLMYPYGAP 115
Query: 70 KFPCPQCHVRIDV 82
C C+ D+
Sbjct: 116 AVRCSSCNSVTDI 128
Score = 32.3 bits (72), Expect = 0.50
Identities = 17/59 (28%), Positives = 29/59 (48%), Gaps = 4/59 (6%)
Query: 16 RCAGCRTY--FSAAQGVAELPCPNCQMPHVFFVDSSAVKIRCSSCKAVVNAPSNLSKFP 72
+C+ C+T A V ++ C NC++ + A +RCSSC +V + N + P
Sbjct: 80 KCSSCQTVNLVLEANQVGQVNCNNCKL--LLMYPYGAPAVRCSSCNSVTDISENNKRPP 136
>At2g43170 unknown protein
Length = 504
Score = 38.9 bits (89), Expect = 0.005
Identities = 15/48 (31%), Positives = 27/48 (56%)
Query: 137 EGDGGIAGETFTDYRPSKLSIGSPHPDPIVETSSLSAVQPPEPTYDPK 184
+G+ + G+ F+ P+ L+ S HPD T ++ V PP+ ++PK
Sbjct: 276 QGNNNVMGDMFSQATPNSLTSSSSHPDLTPLTGAIEIVPPPQKKFEPK 323
>At1g32540 zinc-finger containing protein
Length = 154
Score = 37.4 bits (85), Expect = 0.016
Identities = 18/90 (20%), Positives = 32/90 (35%), Gaps = 2/90 (2%)
Query: 8 PPPSTARAR--CAGCRTYFSAAQGVAELPCPNCQMPHVFFVDSSAVKIRCSSCKAVVNAP 65
PPP T A+ C GC T +G + C C ++ + + C +C ++
Sbjct: 65 PPPGTEMAQLVCGGCHTLLMYIRGATSVQCSCCHTVNLALEANQVAHVNCGNCMMLLMYQ 124
Query: 66 SNLSKFPCPQCHVRIDVHADVEEVNEKIDS 95
C C+ V + K ++
Sbjct: 125 YGARSVKCAVCNFVTSVGGSTSTTDSKFNN 154
Score = 33.9 bits (76), Expect = 0.17
Identities = 19/76 (25%), Positives = 32/76 (42%), Gaps = 5/76 (6%)
Query: 5 PQPPPPSTARARCAGCRTYFSAAQGVAELPCPNCQMPHVFFVDSSAVKIRCSSCKAVVNA 64
P P PP+ A A + G ++L C C+ ++ A + C+ C AV
Sbjct: 7 PYPTPPAPALAPSYNTPPANGSTSGQSQLVCSGCR--NLLMYPVGATSVCCAVCNAVTAV 64
Query: 65 P---SNLSKFPCPQCH 77
P + +++ C CH
Sbjct: 65 PPPGTEMAQLVCGGCH 80
>At1g24330 hypothetical protein
Length = 709
Score = 32.7 bits (73), Expect = 0.39
Identities = 26/79 (32%), Positives = 38/79 (47%), Gaps = 12/79 (15%)
Query: 296 VREGVIFSTYSSLIASSDRGRTRMQQLVQWCGPKFDGLIIFDECHKAKNLVPEKDKKPTK 355
++EGVI S S + S RGR + Q+L L++F E P K++ P K
Sbjct: 612 LQEGVIPSLVSISVNGSPRGRDKSQKL----------LMLFREQRHRDQPSPNKEEAPRK 661
Query: 356 TGQAVLDIQAQL--PEARV 372
T A + I A + PE+ V
Sbjct: 662 TVSAPMAIPAPVSAPESEV 680
>At1g02170 hypothetical protein
Length = 379
Score = 32.3 bits (72), Expect = 0.50
Identities = 15/38 (39%), Positives = 18/38 (46%), Gaps = 5/38 (13%)
Query: 7 PPPPSTARA-----RCAGCRTYFSAAQGVAELPCPNCQ 39
PPPPS+ A C+GCRT G + C CQ
Sbjct: 3 PPPPSSIYAPPMLVNCSGCRTPLQLPSGARSIRCALCQ 40
>At1g14190 putative mandelonitrile lyase
Length = 501
Score = 32.0 bits (71), Expect = 0.66
Identities = 42/161 (26%), Positives = 69/161 (42%), Gaps = 29/161 (18%)
Query: 74 PQCHVR---IDVHADVEEVNEKIDSEPVISCF-HKFSQTVSQELDMLQNPRETVGKQSVL 129
P+ H++ I V +++EV K+ P IS +FSQ ++ L+ Q G + +L
Sbjct: 264 PENHLKDFDIPVIVNLKEVGRKMSDNPAISLLVDRFSQ--NRTLEPPQVAAIAEGYKFIL 321
Query: 130 VNEVEQEEGDGGIAGETFTDYRPSKLSIGSPHPDPIVETSSLSAVQPPEPTYDPKIK-ND 188
+EV TD +++SI + P ++ + P +P +K N
Sbjct: 322 ESEVLP------------TDITTTRISIAAKIAFP--KSKGRLKLNSTNPRENPSVKFNY 367
Query: 189 LERSKAL-SCLQIETLVYACQRHLQHVPSGPRAGFFIGDGA 228
LE L +CL++ HLQHV FF+G A
Sbjct: 368 LENKADLDACLEMVL-------HLQHVARSETVTFFLGTQA 401
>At1g05120 hypothetical protein, 3' partial
Length = 773
Score = 32.0 bits (71), Expect = 0.66
Identities = 26/120 (21%), Positives = 46/120 (37%), Gaps = 2/120 (1%)
Query: 31 AELPCPNCQMPHVFFVDSSAVKIRCSSCKAVVNAPSNLSKFPCPQCHVRIDVHADVEEVN 90
+E C C P +V +S + C +C ++ ++L K CP C + V +
Sbjct: 575 SEQECGLCHDPAEDYVVTSCAHVFCKAC--LIGFSASLGKVTCPTCSKLLTVDWTTKADT 632
Query: 91 EKIDSEPVISCFHKFSQTVSQELDMLQNPRETVGKQSVLVNEVEQEEGDGGIAGETFTDY 150
E S+ + F S +LD Q + + + VE++ I FT +
Sbjct: 633 EHKASKTTLKGFRASSILNRIKLDDFQTSTKIEALREEIRFMVERDGSAKAIVFSQFTSF 692
>At5g48810 cytochrome b5 (dbj|BAA74840.1)
Length = 140
Score = 30.4 bits (67), Expect = 1.9
Identities = 21/84 (25%), Positives = 40/84 (47%), Gaps = 4/84 (4%)
Query: 245 HHGRRKTLWISVGSDLKFDARRDLDDMGASCISVHALNKLPYTKLDTKSVGVREGVIFST 304
H G + + S G D A D +D+G S + L++ +DT +V V+ + T
Sbjct: 40 HPGGDEVILTSTGKD----ATDDFEDVGHSSTAKAMLDEYYVGDIDTATVPVKAKFVPPT 95
Query: 305 YSSLIASSDRGRTRMQQLVQWCGP 328
+ +A+ D+ + +L+Q+ P
Sbjct: 96 STKAVATQDKSSDFVIKLLQFLVP 119
>At1g45545 hypothetical protein
Length = 752
Score = 30.0 bits (66), Expect = 2.5
Identities = 27/95 (28%), Positives = 40/95 (41%), Gaps = 7/95 (7%)
Query: 49 SAVKIRCSSCKAVVNAPSNLSKFPCPQCHVRIDVHADVEEVNEKIDSEPVISCF--HKFS 106
S +K+ S V S +F P V ID A E V E + I+ + HK
Sbjct: 124 SRLKVPGSPRAFVHPRSSGSPRFVSPTSPVLIDTAAPFESVKEAVSKFGGITDWKAHKIQ 183
Query: 107 -----QTVSQELDMLQNPRETVGKQSVLVNEVEQE 136
+TV QEL+ +Q KQ+V+ E + +
Sbjct: 184 TIERRKTVDQELEKIQEDMPDYKKQAVVAEEAKHQ 218
>At1g68750 putative phosphoenolpyruvate carboxylase
Length = 981
Score = 29.6 bits (65), Expect = 3.3
Identities = 28/103 (27%), Positives = 46/103 (44%), Gaps = 14/103 (13%)
Query: 171 LSAVQPPEPTYDPKIKNDLERSKALSCLQIETLVYACQRHLQHV-PSGPRA--GFF-IG- 225
L+ ++PP+P + K +N +E +SC + VY L + + P+A GF IG
Sbjct: 742 LATLKPPQPPREEKWRNLMEEISGISCQHYRSTVYENPEFLSYFHEATPQAELGFLNIGS 801
Query: 226 ------DGAGVGKGRTVAGLIWENWHHGR-RKTLWISVGSDLK 261
+G+G R + + W R W+ VG+ LK
Sbjct: 802 RPTRRKSSSGIGHLRAIPWVF--AWTQTRFVLPAWLGVGAGLK 842
>At1g80400 putative RING zinc finger protein
Length = 407
Score = 28.9 bits (63), Expect = 5.6
Identities = 20/59 (33%), Positives = 24/59 (39%), Gaps = 9/59 (15%)
Query: 15 ARCAGCRTYFSAAQGVAELPCPNCQMPHVFFVDS----SAVKIRCSSCKAVVNAPSNLS 69
A C C T + + V ELPC HVF VD + C CK V S+ S
Sbjct: 353 ASCCICLTRYGDDEQVRELPC-----SHVFHVDCVDKWLKINATCPLCKNEVGESSSAS 406
>At4g38770 extensin - like protein
Length = 448
Score = 28.5 bits (62), Expect = 7.3
Identities = 11/30 (36%), Positives = 15/30 (49%)
Query: 160 PHPDPIVETSSLSAVQPPEPTYDPKIKNDL 189
P P P+ E + PP P YDP K ++
Sbjct: 193 PPPVPVYEPPPKKEIPPPVPVYDPPPKKEV 222
>At5g45520 putative protein
Length = 1167
Score = 28.1 bits (61), Expect = 9.5
Identities = 17/56 (30%), Positives = 30/56 (53%), Gaps = 1/56 (1%)
Query: 87 EEVNEKIDSEPVISCFHKFSQTVSQELDMLQNPRETVGKQSVLVNEV-EQEEGDGG 141
E NEK +++ ++ K + +E D +QN +T ++ V E +QE+GD G
Sbjct: 561 ETTNEKKENQGGLATDKKLKDLMEREDDQVQNYGQTSKEEKGNVEETGKQEDGDQG 616
>At3g52130 5B protein like protein
Length = 125
Score = 28.1 bits (61), Expect = 9.5
Identities = 20/55 (36%), Positives = 24/55 (43%), Gaps = 6/55 (10%)
Query: 4 QPQPPPPSTARARCAGC-RTYFSAAQGVAELPCPNCQMPHVFFVDSSAVKIRCSS 57
Q PPPP GC RT+FSA V +PC P F +I CS+
Sbjct: 32 QQPPPPPMLPEEEVGGCSRTFFSAL--VQLIPCRAAVAP---FSPIPPTEICCSA 81
>At3g29380 transcription initiation factor IIB, putative
Length = 336
Score = 28.1 bits (61), Expect = 9.5
Identities = 39/186 (20%), Positives = 69/186 (36%), Gaps = 20/186 (10%)
Query: 32 ELPCPNCQMPHVFFVDSSAVKIRCSSCKAVVNA-----PSNLSKFPCPQCHVRIDVHADV 86
E C +C+ P + VD S+ CS C V+ A F H D +
Sbjct: 3 EETCLDCKRPTIMVVDHSSGDTICSECGLVLEAHIIEYSQEWRTFASDDNHSDRDPNRVG 62
Query: 87 EEVNEKIDSEPVISCFHKFSQTVSQELDMLQNPRETVGKQSVLVNEVEQEEGDGGIAGET 146
N + S +++ K +T S L ++ + N+V+ E D + +
Sbjct: 63 AATNPFLKSGDLVTIIEKPKETASSVLS-----KDDISTLFRAHNQVKNHEED--LIKQA 115
Query: 147 FTDYRPSKLSIGSPHPDPIVETSSLSAVQPPEPTYDPKIKNDLERSKALSCLQIETLVYA 206
F + + ++ D ++ + + V YD L R K L+ + ++ A
Sbjct: 116 FEEIQRMTDALDL---DIVINSRACEIVS----KYDGHANTKLRRGKKLNAICAASVSTA 168
Query: 207 CQRHLQ 212
C R LQ
Sbjct: 169 C-RELQ 173
>At2g33250 unknown protein
Length = 529
Score = 28.1 bits (61), Expect = 9.5
Identities = 19/58 (32%), Positives = 31/58 (52%), Gaps = 9/58 (15%)
Query: 149 DYRPSKLSIGSPHPDPIVETSSLSAVQPPE-PTYDPKIKNDL---ERSKALSCLQIET 202
++RP K P+PDP++ S +QP E +K+D+ +R+ A +CL ET
Sbjct: 440 EFRPYK-----PNPDPLLHICSTWDIQPNEVMMVGDSLKDDIACGKRAGAFTCLLDET 492
>At1g28460 hypothetical protein
Length = 182
Score = 28.1 bits (61), Expect = 9.5
Identities = 9/21 (42%), Positives = 17/21 (80%)
Query: 118 NPRETVGKQSVLVNEVEQEEG 138
NP++T GKQ + + ++E++EG
Sbjct: 2 NPKKTKGKQKINIKKIEKDEG 22
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.136 0.419
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,297,215
Number of Sequences: 26719
Number of extensions: 472855
Number of successful extensions: 1546
Number of sequences better than 10.0: 19
Number of HSP's better than 10.0 without gapping: 9
Number of HSP's successfully gapped in prelim test: 10
Number of HSP's that attempted gapping in prelim test: 1517
Number of HSP's gapped (non-prelim): 31
length of query: 422
length of database: 11,318,596
effective HSP length: 102
effective length of query: 320
effective length of database: 8,593,258
effective search space: 2749842560
effective search space used: 2749842560
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC148360.15


