

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148360.12 + phase: 0 /pseudo
(102 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
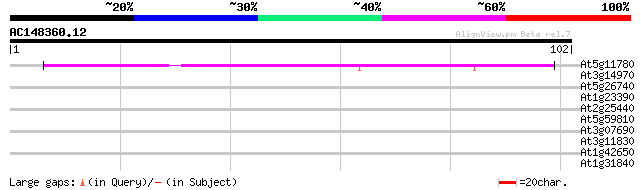
Score E
Sequences producing significant alignments: (bits) Value
At5g11780 putative protein 47 2e-06
At3g14970 hypothetical protein 27 1.3
At5g26740 unknown protein 27 1.7
At1g23390 unknown protein 27 2.2
At2g25440 putative disease resistance protein 26 2.9
At5g59810 subtilisin-like protease - like protein 26 3.8
At3g07690 putative glycerol-3-phosphate dehydrogenase 26 3.8
At3g11830 putative T-complex protein 1, ETA subunit 25 5.0
At1g42650 unknown protein 25 6.5
At1g31840 hypothetical protein 25 8.5
>At5g11780 putative protein
Length = 494
Score = 46.6 bits (109), Expect = 2e-06
Identities = 31/105 (29%), Positives = 53/105 (49%), Gaps = 14/105 (13%)
Query: 7 PPFDVIWVYSAIKFGCRKSLKGDILEQISAAKALFQLISACSASVGGSKSIALLAP---- 62
PP +++W YSAI+F K D + + FQLI + S S G K ++LL+P
Sbjct: 57 PPLELVWFYSAIRFYSSKLAFRD--DSVRLTSCFFQLIVSFSDSFSGVKKVSLLSPVVYQ 114
Query: 63 ------KAMKEVKSLVDMILGFMSICC--SKISEEKDLDLVLSFS 99
++ SL++ I+ ++S+ C +E+ D+ +V FS
Sbjct: 115 LSRLVISRRRDALSLLEGIVSYISMYCVDEPGNEDDDVLMVSGFS 159
>At3g14970 hypothetical protein
Length = 220
Score = 27.3 bits (59), Expect = 1.3
Identities = 14/37 (37%), Positives = 22/37 (58%), Gaps = 2/37 (5%)
Query: 61 APKAMKEVKSLVDMILGFMSICCSKISEEKDLDLVLS 97
AP+ + V L+D IL C KIS K++D+V++
Sbjct: 95 APRYIDVVTELIDAILDLKK--CDKISNVKEVDVVIN 129
>At5g26740 unknown protein
Length = 422
Score = 26.9 bits (58), Expect = 1.7
Identities = 15/51 (29%), Positives = 29/51 (56%), Gaps = 1/51 (1%)
Query: 23 RKSLKGDILEQISAAKALFQLISACSASVGGSKSIAL-LAPKAMKEVKSLV 72
+ S+ D + ++ A ++ +S C A VGG S+ L L+ +++K SL+
Sbjct: 66 KSSIYFDSIREVYEAWVIYNFLSLCLAWVGGPGSVVLSLSGRSLKPSWSLM 116
>At1g23390 unknown protein
Length = 394
Score = 26.6 bits (57), Expect = 2.2
Identities = 13/33 (39%), Positives = 19/33 (57%)
Query: 25 SLKGDILEQISAAKALFQLISACSASVGGSKSI 57
S+ GDILE I + L L SAC S ++++
Sbjct: 19 SIDGDILESILSYLPLLDLDSACQVSKSWNRAV 51
>At2g25440 putative disease resistance protein
Length = 671
Score = 26.2 bits (56), Expect = 2.9
Identities = 25/62 (40%), Positives = 34/62 (54%), Gaps = 7/62 (11%)
Query: 11 VIWVYSAIKFGCRKSLKGDILEQISAAKALFQLISACSASVGG-SKSIALLAPKAMKEVK 69
V+ YSAI F R L+G+I E I KAL L + +A G +S+A L KE++
Sbjct: 487 VLTSYSAIDFS-RNLLEGNIPESIGLLKALIALNLSNNAFTGHIPQSLANL-----KELQ 540
Query: 70 SL 71
SL
Sbjct: 541 SL 542
>At5g59810 subtilisin-like protease - like protein
Length = 778
Score = 25.8 bits (55), Expect = 3.8
Identities = 18/54 (33%), Positives = 28/54 (51%), Gaps = 1/54 (1%)
Query: 24 KSLKGDILEQISAAKALFQLISACSASV-GGSKSIALLAPKAMKEVKSLVDMIL 76
+S KG L + + ++ LISA A+V G+ + ALL K + K + IL
Sbjct: 373 QSFKGTSLSKPLPEEKMYSLISAADANVANGNVTDALLCKKGSLDPKKVKGKIL 426
>At3g07690 putative glycerol-3-phosphate dehydrogenase
Length = 455
Score = 25.8 bits (55), Expect = 3.8
Identities = 17/65 (26%), Positives = 32/65 (49%), Gaps = 1/65 (1%)
Query: 25 SLKGD-ILEQISAAKALFQLISACSASVGGSKSIALLAPKAMKEVKSLVDMILGFMSICC 83
S+KG +++ +SA KA F+L++ S S+ + + P + + ++ IL C
Sbjct: 359 SIKGKGMIQGVSAVKAFFELLNQSSLSLQHPEEGKPVTPAELCPILKMLYRILITREFSC 418
Query: 84 SKISE 88
I E
Sbjct: 419 EAILE 423
>At3g11830 putative T-complex protein 1, ETA subunit
Length = 557
Score = 25.4 bits (54), Expect = 5.0
Identities = 13/39 (33%), Positives = 23/39 (58%), Gaps = 1/39 (2%)
Query: 36 AAKALFQLISACSASVG-GSKSIALLAPKAMKEVKSLVD 73
AAK L + + + VG G+ ++ LLA + +KE K ++
Sbjct: 78 AAKILVDIAKSQDSEVGDGTTTVVLLAAEFLKEAKPFIE 116
>At1g42650 unknown protein
Length = 593
Score = 25.0 bits (53), Expect = 6.5
Identities = 12/36 (33%), Positives = 19/36 (52%)
Query: 67 EVKSLVDMILGFMSICCSKISEEKDLDLVLSFSSLG 102
E +VDM + F +C K + K D+V++ LG
Sbjct: 28 ERDDVVDMAVVFYEVCILKGAHAKAKDVVITVKKLG 63
>At1g31840 hypothetical protein
Length = 845
Score = 24.6 bits (52), Expect = 8.5
Identities = 11/26 (42%), Positives = 14/26 (53%)
Query: 22 CRKSLKGDILEQISAAKALFQLISAC 47
C K LKG ++QI A L L+ C
Sbjct: 255 CNKVLKGLSVDQIEVASRLLSLVLDC 280
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.321 0.135 0.381
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,119,547
Number of Sequences: 26719
Number of extensions: 62732
Number of successful extensions: 164
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 158
Number of HSP's gapped (non-prelim): 10
length of query: 102
length of database: 11,318,596
effective HSP length: 78
effective length of query: 24
effective length of database: 9,234,514
effective search space: 221628336
effective search space used: 221628336
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 52 (24.6 bits)
Medicago: description of AC148360.12


