

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148348.6 + phase: 0
(552 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
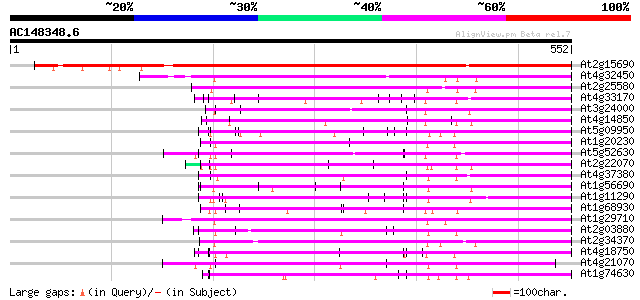
Score E
Sequences producing significant alignments: (bits) Value
At2g15690 unknown protein 540 e-154
At4g32450 putative protein 285 4e-77
At2g25580 putative selenium-binding protein 276 3e-74
At4g33170 putative protein 258 5e-69
At3g24000 hypothetical protein 254 7e-68
At4g14850 unknown protein 245 5e-65
At5g09950 selenium-binding protein-like 244 1e-64
At1g20230 hypothetical protein 244 1e-64
At5g52630 selenium-binding protein-like 242 4e-64
At2g22070 hypothetical protein 242 4e-64
At4g37380 putative protein 240 2e-63
At1g56690 hypothetical protein 239 2e-63
At1g11290 hypothetical protein 236 2e-62
At1g68930 hypothetical protein 236 3e-62
At1g29710 229 4e-60
At2g03880 putative selenium-binding protein 228 7e-60
At2g34370 putative selenium-binding protein 227 1e-59
At4g18750 putative protein 223 2e-58
At4g21070 putative protein (fragment) 221 6e-58
At1g74630 hypothetical protein 218 5e-57
>At2g15690 unknown protein
Length = 579
Score = 540 bits (1392), Expect = e-154
Identities = 280/547 (51%), Positives = 358/547 (65%), Gaps = 33/547 (6%)
Query: 25 KTLSTSAIPNEYGRLPP------HQVQQPPSDHNQHFNHHNTQNAPGRFPQQ--DWNPQN 76
K LSTSA N+Y + P HQ PP Q F+ N N R PQ W+ Q+
Sbjct: 47 KHLSTSAAANDYHQNPQSGSPSQHQRPYPP----QSFDSQNQTNTNQRVPQSPNQWSTQH 102
Query: 77 PNFRPPQPPQNPNYRQQPPP---QNPNFRPPPP------PQNPNFRPPPPPQNPNFRQPI 127
P QNP + Q PP QNP PQ+ RP Q P + P
Sbjct: 103 GGQIPQYGGQNPQHGGQRPPYGGQNPQQGGQMSQYGGHNPQHGGHRPQYGGQRPQYGGPG 162
Query: 128 R--QNPNFQPPNGPNRGINQNQWNPQNGNLNQFQNPNNQFQTPNVQEQAPPPPSIVDLTR 185
QN N Q N Q Q PQ P + Q+PN + PPPS+ ++ R
Sbjct: 163 NNYQNQNVQQSNQSQYYTPQQQQQPQP--------PRSSNQSPNQMNEVAPPPSVEEVMR 214
Query: 186 FCQEGKVKEALELMEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMH 245
CQ K+A+EL++KG D CF +LF+ C KS+E +KKVHD+FLQS FR D K++
Sbjct: 215 LCQRRLYKDAIELLDKGAMPDRECFVLLFESCANLKSLEHSKKVHDHFLQSKFRGDPKLN 274
Query: 246 NKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLE 305
N VI M+G C S+TDA+RVFDHM +++MDSWH+M+ Y+++ MGD+ L LFE+M + GL+
Sbjct: 275 NMVISMFGECSSITDAKRVFDHMVDKDMDSWHLMMCAYSDNGMGDDALHLFEEMTKHGLK 334
Query: 306 ITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEE 365
ET L V AC + +E+A+++ +SMK+++GI P EHY+G+L VLG+ G+L EAE+
Sbjct: 335 PNEETFLTVFLACATVGGIEEAFLHFDSMKNEHGISPKTEHYLGVLGVLGKCGHLVEAEQ 394
Query: 366 FIEQLPFEPTVTVFETLKNYARIHGDVDLEDHVEELIVSLDPSKAVANKIPTPPPKKYTA 425
+I LPFEPT +E ++NYAR+HGD+DLED++EEL+V +DPSKAV NKIPTPPPK +
Sbjct: 395 YIRDLPFEPTADFWEAMRNYARLHGDIDLEDYMEELMVDVDPSKAVINKIPTPPPKSFKE 454
Query: 426 ISMLDGKNRIIEYKNPTLYKDDEKLIAMNSMKDAGYVPDTRYVLHDIDQEAKEQALLYHS 485
+M+ K+RI+E++N T YKD+ K M + K YVPDTR+VLHDIDQEAKEQALLYHS
Sbjct: 455 TNMVTSKSRILEFRNLTFYKDEAK--EMAAKKGVVYVPDTRFVLHDIDQEAKEQALLYHS 512
Query: 486 ERLAIAYGLISTPPRTPLRIIKNLRVCGDCHNAIKIMSRIVGRELIVRDNKRFHHFKDGK 545
ERLAIAYG+I TPPR L IIKNLRVCGDCHN IKIMS+I+GR LIVRDNKRFHHFKDGK
Sbjct: 513 ERLAIAYGIICTPPRKTLTIIKNLRVCGDCHNFIKIMSKIIGRVLIVRDNKRFHHFKDGK 572
Query: 546 CSCGDYW 552
CSCGDYW
Sbjct: 573 CSCGDYW 579
>At4g32450 putative protein
Length = 537
Score = 285 bits (730), Expect = 4e-77
Identities = 161/442 (36%), Positives = 250/442 (56%), Gaps = 29/442 (6%)
Query: 128 RQNPNFQPPNGPNRGINQNQWNPQNGNLNQFQNPNNQFQTPNVQEQAPPPPSIVDLTRFC 187
++N N+Q +G + PQ N N FQ + S+ +L C
Sbjct: 108 QRNQNWQSSDGCSSYGTTGNGVPQENNTG-----GNHFQQDHSGHS-----SLDELDSIC 157
Query: 188 QEGKVKEALELME----KGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFK 243
+EGKVK+A+E+++ +G D + LCG ++++++AK VH++ S SD
Sbjct: 158 REGKVKKAVEIIKSWRNEGYVVDLPRLFWIAQLCGDAQALQEAKVVHEFITSSVGISDIS 217
Query: 244 MHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELG 303
+N +IEMY C S+ DA VF+ MP RN+++W +IR +A + G++ + F + + G
Sbjct: 218 AYNSIIEMYSGCGSVEDALTVFNSMPERNLETWCGVIRCFAKNGQGEDAIDTFSRFKQEG 277
Query: 304 LEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEA 363
+ E + ACG + + ++ ESM +YGI P +EHY+ L+ +L + GYL EA
Sbjct: 278 NKPDGEMFKEIFFACGVLGDMNEGLLHFESMYKEYGIIPCMEHYVSLVKMLAEPGYLDEA 337
Query: 364 EEFIEQLPFEPTVTVFETLKNYARIHGDVDLEDHVEELIVSLDPSKA-VANKIPTPPPKK 422
F+E + EP V ++ETL N +R+HGD+ L D ++++ LD S+ +K P K
Sbjct: 338 LRFVESM--EPNVDLWETLMNLSRVHGDLILGDRCQDMVEQLDASRLNKESKAGLVPVKS 395
Query: 423 YTAIS-----MLDGKNRIIEYK---NPTLYKDDEKLIAMNSMKD----AGYVPDTRYVLH 470
+ M G N I Y + + ++ E +A+ S+K+ GYVP ++ LH
Sbjct: 396 SDLVKEKLQRMAKGPNYGIRYMAAGDISRPENRELYMALKSLKEHMIEIGYVPLSKLALH 455
Query: 471 DIDQEAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLRVCGDCHNAIKIMSRIVGREL 530
D+DQE+K++ L H+ER A + TP R+ +R++KNLRVC DCHNA+K+MS+IVGREL
Sbjct: 456 DVDQESKDENLFNHNERFAFISTFLDTPARSLIRVMKNLRVCADCHNALKLMSKIVGREL 515
Query: 531 IVRDNKRFHHFKDGKCSCGDYW 552
I RD KRFHH KDG CSC +YW
Sbjct: 516 ISRDAKRFHHMKDGVCSCREYW 537
>At2g25580 putative selenium-binding protein
Length = 567
Score = 276 bits (705), Expect = 3e-74
Identities = 149/395 (37%), Positives = 231/395 (57%), Gaps = 24/395 (6%)
Query: 180 IVDLTRFCQEGKVKEALE----LMEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQ 235
I + FC+ GKVK+AL L D + L +CG+++ +++AK VH
Sbjct: 175 IEEYDAFCKHGKVKKALYTIDILASMNYVVDLSRLLRLAKICGEAEGLQEAKTVHGKISA 234
Query: 236 STFRSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQL 295
S D ++ ++EMY NC +A VF+ M +N+++W ++IR +A + G++ + +
Sbjct: 235 SVSHLDLSSNHVLLEMYSNCGLANEAASVFEKMSEKNLETWCIIIRCFAKNGFGEDAIDM 294
Query: 296 FEQMNELGLEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLG 355
F + E G + + ACG V++ ++ ESM YGI P +E Y+ L+++
Sbjct: 295 FSRFKEEGNIPDGQLFRGIFYACGMLGDVDEGLLHFESMSRDYGIAPSIEDYVSLVEMYA 354
Query: 356 QSGYLKEAEEFIEQLPFEPTVTVFETLKNYARIHGDVDLEDHVEELIVSLDPSK------ 409
G+L EA EF+E++P EP V V+ETL N +R+HG+++L D+ E++ LDP++
Sbjct: 355 LPGFLDEALEFVERMPMEPNVDVWETLMNLSRVHGNLELGDYCAEVVEFLDPTRLNKQSR 414
Query: 410 -----AVANKIPTPPPKKYTAISMLDG-KNRIIEYK--NPTLYKDDEKLIAMNSMK---- 457
A+ + KK + I L G K+ + E++ + L ++DE + ++K
Sbjct: 415 EGFIPVKASDVEKESLKKRSGI--LHGVKSSMQEFRAGDTNLPENDELFQLLRNLKMHMV 472
Query: 458 DAGYVPDTRYVLHDIDQEAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLRVCGDCHN 517
+ GYV +TR LHDIDQE+KE LL HSER+A A ++++ PR P +IKNLRVC DCHN
Sbjct: 473 EVGYVAETRMALHDIDQESKETLLLGHSERIAFARAVLNSAPRKPFTVIKNLRVCVDCHN 532
Query: 518 AIKIMSRIVGRELIVRDNKRFHHFKDGKCSCGDYW 552
A+KIMS IVGRE+I RD KRFH K+G C+C DYW
Sbjct: 533 ALKIMSDIVGREVITRDIKRFHQMKNGACTCKDYW 567
>At4g33170 putative protein
Length = 990
Score = 258 bits (660), Expect = 5e-69
Identities = 133/360 (36%), Positives = 206/360 (56%), Gaps = 31/360 (8%)
Query: 222 SVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIR 281
++E +++H L+ +D + +++MY C S+ DA +F + N+ +W+ M+
Sbjct: 633 ALEQGRQIHANALKLNCTNDPFVGTSLVDMYAKCGSIDDAYCLFKRIEMMNITAWNAMLV 692
Query: 282 GYANSTMGDEGLQLFEQMNELGLEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIE 341
G A G E LQLF+QM LG++ T + VLSAC + V +AY ++ SM YGI+
Sbjct: 693 GLAQHGEGKETLQLFKQMKSLGIKPDKVTFIGVLSACSHSGLVSEAYKHMRSMHGDYGIK 752
Query: 342 PGVEHYMGLLDVLGQSGYLKEAEEFIEQLPFEPTVTVFETLKNYARIHGDVDLEDHVEEL 401
P +EHY L D LG++G +K+AE IE + E + +++ TL R+ GD + V
Sbjct: 753 PEIEHYSCLADALGRAGLVKQAENLIESMSMEASASMYRTLLAACRVQGDTETGKRVATK 812
Query: 402 IVSLDP---------------------SKAVANKIPTPPPKKYTAISMLDGKNRIIEY-- 438
++ L+P K + KK S ++ KN+I +
Sbjct: 813 LLELEPLDSSAYVLLSNMYAAASKWDEMKLARTMMKGHKVKKDPGFSWIEVKNKIHIFVV 872
Query: 439 ------KNPTLYKDDEKLIAMNSMKDAGYVPDTRYVLHDIDQEAKEQALLYHSERLAIAY 492
+ +Y+ + +I +K GYVP+T + L D+++E KE+AL YHSE+LA+A+
Sbjct: 873 DDRSNRQTELIYRKVKDMI--RDIKQEGYVPETDFTLVDVEEEEKERALYYHSEKLAVAF 930
Query: 493 GLISTPPRTPLRIIKNLRVCGDCHNAIKIMSRIVGRELIVRDNKRFHHFKDGKCSCGDYW 552
GL+STPP TP+R+IKNLRVCGDCHNA+K ++++ RE+++RD RFH FKDG CSCGDYW
Sbjct: 931 GLLSTPPSTPIRVIKNLRVCGDCHNAMKYIAKVYNREIVLRDANRFHRFKDGICSCGDYW 990
Score = 62.0 bits (149), Expect = 8e-10
Identities = 51/197 (25%), Positives = 91/197 (45%), Gaps = 8/197 (4%)
Query: 183 LTRFCQEGKVKEALELMEKGIKADANCFEILFDL----CGKSKSVEDAKKVHDYFLQSTF 238
L+ + G+ L+ +++D C ++ F L K S+ ++VH L+
Sbjct: 287 LSEYLHSGQYSALLKCFADMVESDVECDQVTFILMLATAVKVDSLALGQQVHCMALKLGL 346
Query: 239 RSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQ 298
+ N +I MY + AR VFD+M R++ SW+ +I G A + + E + LF Q
Sbjct: 347 DLMLTVSNSLINMYCKLRKFGFARTVFDNMSERDLISWNSVIAGIAQNGLEVEAVCLFMQ 406
Query: 299 MNELGLEITSETMLAVLSACGS-AEAVE-DAYIYLESMKSKYGIEPGVEHYMGLLDVLGQ 356
+ GL+ TM +VL A S E + +++ ++K + V L+D +
Sbjct: 407 LLRCGLKPDQYTMTSVLKAASSLPEGLSLSKQVHVHAIKINNVSDSFVS--TALIDAYSR 464
Query: 357 SGYLKEAEEFIEQLPFE 373
+ +KEAE E+ F+
Sbjct: 465 NRCMKEAEILFERHNFD 481
Score = 58.9 bits (141), Expect = 7e-09
Identities = 43/175 (24%), Positives = 80/175 (45%), Gaps = 4/175 (2%)
Query: 191 KVKEALELMEK-GIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVI 249
K + LM K G ++D +F CG ++ K+VH Y ++S + D + + ++
Sbjct: 500 KTLKLFALMHKQGERSDDFTLATVFKTCGFLFAINQGKQVHAYAIKSGYDLDLWVSSGIL 559
Query: 250 EMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSE 309
+MY C M+ A+ FD +P + +W MI G + + +F QM +G+
Sbjct: 560 DMYVKCGDMSAAQFAFDSIPVPDDVAWTTMISGCIENGEEERAFHVFSQMRLMGVLPDEF 619
Query: 310 TMLAVLSACGSAEAVEDA-YIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEA 363
T+ + A A+E I+ ++K +P V L+D+ + G + +A
Sbjct: 620 TIATLAKASSCLTALEQGRQIHANALKLNCTNDPFVG--TSLVDMYAKCGSIDDA 672
Score = 57.8 bits (138), Expect = 2e-08
Identities = 41/149 (27%), Positives = 71/149 (47%), Gaps = 14/149 (9%)
Query: 246 NKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMG-----DEGLQLFEQMN 300
N +I MY C S+T ARRVFD MP+R++ SW+ ++ YA S+ + LF +
Sbjct: 78 NNLISMYSKCGSLTYARRVFDKMPDRDLVSWNSILAAYAQSSECVVENIQQAFLLFRILR 137
Query: 301 ELGLEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGV---EHYMG-LLDVLGQ 356
+ + + T+ +L C + Y++ Y + G+ E G L+++ +
Sbjct: 138 QDVVYTSRMTLSPMLKLC-----LHSGYVWASESFHGYACKIGLDGDEFVAGALVNIYLK 192
Query: 357 SGYLKEAEEFIEQLPFEPTVTVFETLKNY 385
G +KE + E++P+ V LK Y
Sbjct: 193 FGKVKEGKVLFEEMPYRDVVLWNLMLKAY 221
Score = 55.8 bits (133), Expect = 6e-08
Identities = 47/205 (22%), Positives = 97/205 (46%), Gaps = 6/205 (2%)
Query: 196 LELMEKGIKADANCF-EILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGN 254
++L+ G+K D +L + + +K+VH + ++ SD + +I+ Y
Sbjct: 405 MQLLRCGLKPDQYTMTSVLKAASSLPEGLSLSKQVHVHAIKINNVSDSFVSTALIDAYSR 464
Query: 255 CKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSETMLAV 314
+ M +A +F+ N ++ +W+ M+ GY S G + L+LF M++ G T+ V
Sbjct: 465 NRCMKEAEILFERH-NFDLVAWNAMMAGYTQSHDGHKTLKLFALMHKQGERSDDFTLATV 523
Query: 315 LSACGSAEAV-EDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLPFE 373
CG A+ + ++ ++KS Y ++ V G+LD+ + G + A+ + +P
Sbjct: 524 FKTCGFLFAINQGKQVHAYAIKSGYDLDLWVS--SGILDMYVKCGDMSAAQFAFDSIPV- 580
Query: 374 PTVTVFETLKNYARIHGDVDLEDHV 398
P + T+ + +G+ + HV
Sbjct: 581 PDDVAWTTMISGCIENGEEERAFHV 605
Score = 36.6 bits (83), Expect = 0.037
Identities = 20/99 (20%), Positives = 43/99 (43%)
Query: 213 LFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRN 272
+ LC S V ++ H Y + D + ++ +Y + + + +F+ MP R+
Sbjct: 151 MLKLCLHSGYVWASESFHGYACKIGLDGDEFVAGALVNIYLKFGKVKEGKVLFEEMPYRD 210
Query: 273 MDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSETM 311
+ W++M++ Y +E + L + GL T+
Sbjct: 211 VVLWNLMLKAYLEMGFKEEAIDLSSAFHSSGLNPNEITL 249
>At3g24000 hypothetical protein
Length = 633
Score = 254 bits (650), Expect = 7e-68
Identities = 139/391 (35%), Positives = 219/391 (55%), Gaps = 32/391 (8%)
Query: 193 KEALELME----KGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKV 248
++ALEL + G + + LF C + +E K VH Y ++S + N +
Sbjct: 244 EKALELFQGMLRDGFRPSHFSYASLFGACSSTGFLEQGKWVHAYMIKSGEKLVAFAGNTL 303
Query: 249 IEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITS 308
++MY S+ DAR++FD + R++ SW+ ++ YA G E + FE+M +G+
Sbjct: 304 LDMYAKSGSIHDARKIFDRLAKRDVVSWNSLLTAYAQHGFGKEAVWWFEEMRRVGIRPNE 363
Query: 309 ETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIE 368
+ L+VL+AC + +++ + Y E MK K GI P HY+ ++D+LG++G L A FIE
Sbjct: 364 ISFLSVLTACSHSGLLDEGWHYYELMK-KDGIVPEAWHYVTVVDLLGRAGDLNRALRFIE 422
Query: 369 QLPFEPTVTVFETLKNYARIHGDVDLEDHVEELIVSLDP--------------------- 407
++P EPT +++ L N R+H + +L + E + LDP
Sbjct: 423 EMPIEPTAAIWKALLNACRMHKNTELGAYAAEHVFELDPDDPGPHVILYNIYASGGRWND 482
Query: 408 SKAVANKIPTPPPKKYTAISMLDGKNRIIEY-KNPTLYKDDEKLI-----AMNSMKDAGY 461
+ V K+ KK A S ++ +N I + N + E++ + +K+ GY
Sbjct: 483 AARVRKKMKESGVKKEPACSWVEIENAIHMFVANDERHPQREEIARKWEEVLAKIKELGY 542
Query: 462 VPDTRYVLHDIDQEAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLRVCGDCHNAIKI 521
VPDT +V+ +DQ+ +E L YHSE++A+A+ L++TPP + + I KN+RVCGDCH AIK+
Sbjct: 543 VPDTSHVIVHVDQQEREVNLQYHSEKIALAFALLNTPPGSTIHIKKNIRVCGDCHTAIKL 602
Query: 522 MSRIVGRELIVRDNKRFHHFKDGKCSCGDYW 552
S++VGRE+IVRD RFHHFKDG CSC DYW
Sbjct: 603 ASKVVGREIIVRDTNRFHHFKDGNCSCKDYW 633
Score = 81.3 bits (199), Expect = 1e-15
Identities = 55/188 (29%), Positives = 84/188 (44%), Gaps = 1/188 (0%)
Query: 203 IKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNCKSMTDAR 262
I AD + L C K + + VH + LQS FR D M N ++ MY C S+ +AR
Sbjct: 56 IPADRRFYNTLLKKCTVFKLLIQGRIVHAHILQSIFRHDIVMGNTLLNMYAKCGSLEEAR 115
Query: 263 RVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSETMLAVLSACGSAE 322
+VF+ MP R+ +W +I GY+ + L F QM G T+ +V+ A +AE
Sbjct: 116 KVFEKMPQRDFVTWTTLISGYSQHDRPCDALLFFNQMLRFGYSPNEFTLSSVIKA-AAAE 174
Query: 323 AVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLPFEPTVTVFETL 382
L K G + V LLD+ + G + +A+ + L V+ +
Sbjct: 175 RRGCCGHQLHGFCVKCGFDSNVHVGSALLDLYTRYGLMDDAQLVFDALESRNDVSWNALI 234
Query: 383 KNYARIHG 390
+AR G
Sbjct: 235 AGHARRSG 242
Score = 58.5 bits (140), Expect = 9e-09
Identities = 41/163 (25%), Positives = 82/163 (50%), Gaps = 2/163 (1%)
Query: 228 KVHDYFLQSTFRSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANST 287
++H + ++ F S+ + + ++++Y M DA+ VFD + +RN SW+ +I G+A +
Sbjct: 182 QLHGFCVKCGFDSNVHVGSALLDLYTRYGLMDDAQLVFDALESRNDVSWNALIAGHARRS 241
Query: 288 MGDEGLQLFEQMNELGLEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHY 347
++ L+LF+ M G + + ++ AC S +E ++ + K G +
Sbjct: 242 GTEKALELFQGMLRDGFRPSHFSYASLFGACSSTGFLEQGK-WVHAYMIKSGEKLVAFAG 300
Query: 348 MGLLDVLGQSGYLKEAEEFIEQLPFEPTVTVFETLKNYARIHG 390
LLD+ +SG + +A + ++L V+ L YA+ HG
Sbjct: 301 NTLLDMYAKSGSIHDARKIFDRLAKRDVVSWNSLLTAYAQ-HG 342
>At4g14850 unknown protein
Length = 684
Score = 245 bits (625), Expect = 5e-65
Identities = 133/365 (36%), Positives = 207/365 (56%), Gaps = 29/365 (7%)
Query: 217 CGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSW 276
C +E + +H + +++ + + +++MYG C + D+ + FD MP +N+ +
Sbjct: 320 CAGMAGLELGRSIHAHAVKACVERTIFVGSALVDMYGKCGCIEDSEQAFDEMPEKNLVTR 379
Query: 277 HMMIRGYANSTMGDEGLQLFEQMNELGLEITSE--TMLAVLSACGSAEAVEDAYIYLESM 334
+ +I GYA+ D L LFE+M G T T +++LSAC A AVE+ +SM
Sbjct: 380 NSLIGGYAHQGQVDMALALFEEMAPRGCGPTPNYMTFVSLLSACSRAGAVENGMKIFDSM 439
Query: 335 KSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLPFEPTVTVFETLKNYARIHGDVDL 394
+S YGIEPG EHY ++D+LG++G ++ A EFI+++P +PT++V+ L+N R+HG L
Sbjct: 440 RSTYGIEPGAEHYSCIVDMLGRAGMVERAYEFIKKMPIQPTISVWGALQNACRMHGKPQL 499
Query: 395 EDHVEELIVSLDP---------------------SKAVANKIPTPPPKKYTAISMLDGKN 433
E + LDP + V ++ KK S + KN
Sbjct: 500 GLLAAENLFKLDPKDSGNHVLLSNTFAAAGRWAEANTVREELKGVGIKKGAGYSWITVKN 559
Query: 434 RIIEY----KNPTLYKDDEKLIAM--NSMKDAGYVPDTRYVLHDIDQEAKEQALLYHSER 487
++ + ++ L K+ + +A N M+ AGY PD + L+D+++E K + +HSE+
Sbjct: 560 QVHAFQAKDRSHILNKEIQTTLAKLRNEMEAAGYKPDLKLSLYDLEEEEKAAEVSHHSEK 619
Query: 488 LAIAYGLISTPPRTPLRIIKNLRVCGDCHNAIKIMSRIVGRELIVRDNKRFHHFKDGKCS 547
LA+A+GL+S P P+RI KNLR+CGDCH+ K +S V RE+IVRDN RFH FKDG CS
Sbjct: 620 LALAFGLLSLPLSVPIRITKNLRICGDCHSFFKFVSGSVKREIIVRDNNRFHRFKDGICS 679
Query: 548 CGDYW 552
C DYW
Sbjct: 680 CKDYW 684
Score = 69.7 bits (169), Expect = 4e-12
Identities = 57/251 (22%), Positives = 117/251 (45%), Gaps = 17/251 (6%)
Query: 189 EGKVKEALELMEKGIKADANCFEILF----DLCGKSKSVEDAKKVHDYFLQSTFRSDFKM 244
+G+ +EA+E + + D + I F + C + ++H L+S F +D +
Sbjct: 187 DGRPREAIEAFIEFRRIDGHPNSITFCAFLNACSDWLHLNLGMQLHGLVLRSGFDTDVSV 246
Query: 245 HNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGL 304
N +I+ YG CK + + +F M +N SW ++ Y + ++ L+ + + +
Sbjct: 247 CNGLIDFYGKCKQIRSSEIIFTEMGTKNAVSWCSLVAAYVQNHEDEKASVLYLRSRKDIV 306
Query: 305 EITSETMLAVLSACGSAEAVE-DAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEA 363
E + + +VLSAC +E I+ ++K+ +E + L+D+ G+ G ++++
Sbjct: 307 ETSDFMISSVLSACAGMAGLELGRSIHAHAVKA--CVERTIFVGSALVDMYGKCGCIEDS 364
Query: 364 EEFIEQLPFEPTVTVFETLKNYARIHGDVDLEDHVEELIVSLDPSKAVANKIPTPPPKKY 423
E+ +++P + VT + YA G VD+ +L + +A + P P
Sbjct: 365 EQAFDEMPEKNLVTRNSLIGGYAH-QGQVDM---------ALALFEEMAPRGCGPTPNYM 414
Query: 424 TAISMLDGKNR 434
T +S+L +R
Sbjct: 415 TFVSLLSACSR 425
Score = 55.5 bits (132), Expect = 8e-08
Identities = 40/198 (20%), Positives = 81/198 (40%), Gaps = 1/198 (0%)
Query: 194 EALELMEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYG 253
E E+ +G+ + F F + K++H ++ D + +MY
Sbjct: 95 EFFEMRREGVVPNDFTFPCAFKAVASLRLPVTGKQIHALAVKCGRILDVFVGCSAFDMYC 154
Query: 254 NCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSETMLA 313
+ DAR++FD +P RN+++W+ I E ++ F + + S T A
Sbjct: 155 KTRLRDDARKLFDEIPERNLETWNAFISNSVTDGRPREAIEAFIEFRRIDGHPNSITFCA 214
Query: 314 VLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLPFE 373
L+AC + + + L + + G + V GL+D G+ ++ +E ++ +
Sbjct: 215 FLNACSDWLHL-NLGMQLHGLVLRSGFDTDVSVCNGLIDFYGKCKQIRSSEIIFTEMGTK 273
Query: 374 PTVTVFETLKNYARIHGD 391
V+ + Y + H D
Sbjct: 274 NAVSWCSLVAAYVQNHED 291
Score = 30.4 bits (67), Expect = 2.7
Identities = 22/75 (29%), Positives = 28/75 (37%)
Query: 246 NKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLE 305
N +I MY AR V P RN+ SW +I G A + L F +M G+
Sbjct: 46 NYLINMYSKLDHPESARLVLRLTPARNVVSWTSLISGLAQNGHFSTALVEFFEMRREGVV 105
Query: 306 ITSETMLAVLSACGS 320
T A S
Sbjct: 106 PNDFTFPCAFKAVAS 120
>At5g09950 selenium-binding protein-like
Length = 995
Score = 244 bits (622), Expect = 1e-64
Identities = 140/386 (36%), Positives = 217/386 (55%), Gaps = 31/386 (8%)
Query: 198 LMEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNCKS 257
+++ G + D+ + + ++E +VH +++ SD + + +++MY C
Sbjct: 610 MLQTGQRLDSFMYATVLSAFASVATLERGMEVHACSVRACLESDVVVGSALVDMYSKCGR 669
Query: 258 MTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSE-TMLAVLS 316
+ A R F+ MP RN SW+ MI GYA G+E L+LFE M G T + VLS
Sbjct: 670 LDYALRFFNTMPVRNSYSWNSMISGYARHGQGEEALKLFETMKLDGQTPPDHVTFVGVLS 729
Query: 317 ACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLPFEPTV 376
AC A +E+ + + ESM YG+ P +EH+ + DVLG++G L + E+FIE++P +P V
Sbjct: 730 ACSHAGLLEEGFKHFESMSDSYGLAPRIEHFSCMADVLGRAGELDKLEDFIEKMPMKPNV 789
Query: 377 TVFET-LKNYARIHG-DVDLEDHVEELIVSLDPSKAV---------------------AN 413
++ T L R +G +L E++ L+P AV
Sbjct: 790 LIWRTVLGACCRANGRKAELGKKAAEMLFQLEPENAVNYVLLGNMYAAGGRWEDLVKARK 849
Query: 414 KIPTPPPKK---YTAISMLDGKNRII--EYKNPTLYKDDEKLIAMN-SMKDAGYVPDTRY 467
K+ KK Y+ ++M DG + + + +P +KL +N M+DAGYVP T +
Sbjct: 850 KMKDADVKKEAGYSWVTMKDGVHMFVAGDKSHPDADVIYKKLKELNRKMRDAGYVPQTGF 909
Query: 468 VLHDIDQEAKEQALLYHSERLAIAYGLISTPPRT-PLRIIKNLRVCGDCHNAIKIMSRIV 526
L+D++QE KE+ L YHSE+LA+A+ L + T P+RI+KNLRVCGDCH+A K +S+I
Sbjct: 910 ALYDLEQENKEEILSYHSEKLAVAFVLAAQRSSTLPIRIMKNLRVCGDCHSAFKYISKIE 969
Query: 527 GRELIVRDNKRFHHFKDGKCSCGDYW 552
GR++I+RD+ RFHHF+DG CSC D+W
Sbjct: 970 GRQIILRDSNRFHHFQDGACSCSDFW 995
Score = 63.2 bits (152), Expect = 4e-10
Identities = 44/152 (28%), Positives = 79/152 (51%), Gaps = 6/152 (3%)
Query: 223 VEDAKKVHDYFLQSTFRSDFKMH--NKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMI 280
++ ++VH + + +T DF + N ++ MY C S+ DARRVF M +++ SW+ MI
Sbjct: 329 LKKGREVHGHVI-TTGLVDFMVGIGNGLVNMYAKCGSIADARRVFYFMTDKDSVSWNSMI 387
Query: 281 RGYANSTMGDEGLQLFEQMNELGLEITSETMLAVLSACGSAE-AVEDAYIYLESMKSKYG 339
G + E ++ ++ M + S T+++ LS+C S + A I+ ES+ K G
Sbjct: 388 TGLDQNGCFIEAVERYKSMRRHDILPGSFTLISSLSSCASLKWAKLGQQIHGESL--KLG 445
Query: 340 IEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLP 371
I+ V L+ + ++GYL E + +P
Sbjct: 446 IDLNVSVSNALMTLYAETGYLNECRKIFSSMP 477
Score = 57.8 bits (138), Expect = 2e-08
Identities = 47/196 (23%), Positives = 79/196 (39%), Gaps = 3/196 (1%)
Query: 196 LELMEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNC 255
L G K + F + E K++H L++ + N +I YG C
Sbjct: 506 LNAQRAGQKLNRITFSSVLSAVSSLSFGELGKQIHGLALKNNIADEATTENALIACYGKC 565
Query: 256 KSMTDARRVFDHMPNRNMD-SWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSETMLAV 314
M ++F M R + +W+ MI GY ++ + + L L M + G + S V
Sbjct: 566 GEMDGCEKIFSRMAERRDNVTWNSMISGYIHNELLAKALDLVWFMLQTGQRLDSFMYATV 625
Query: 315 LSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLPFEP 374
LSA S +E + + + + +E V L+D+ + G L A F +P
Sbjct: 626 LSAFASVATLERG-MEVHACSVRACLESDVVVGSALVDMYSKCGRLDYALRFFNTMPVRN 684
Query: 375 TVTVFETLKNYARIHG 390
+ + + YAR HG
Sbjct: 685 SYSWNSMISGYAR-HG 699
Score = 49.3 bits (116), Expect = 6e-06
Identities = 33/126 (26%), Positives = 53/126 (41%), Gaps = 3/126 (2%)
Query: 226 AKKVHDYFLQSTFRSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYAN 285
A+ H ++ D + N +I Y AR+VFD MP RN SW ++ GY+
Sbjct: 20 ARFFHSRLYKNRLDKDVYLCNNLINAYLETGDSVSARKVFDEMPLRNCVSWACIVSGYSR 79
Query: 286 STMGDEGLQLFEQMNELGLEITSETMLAVLSAC---GSAEAVEDAYIYLESMKSKYGIEP 342
+ E L M + G+ ++VL AC GS + I+ K Y ++
Sbjct: 80 NGEHKEALVFLRDMVKEGIFSNQYAFVSVLRACQEIGSVGILFGRQIHGLMFKLSYAVDA 139
Query: 343 GVEHYM 348
V + +
Sbjct: 140 VVSNVL 145
Score = 44.3 bits (103), Expect = 2e-04
Identities = 46/201 (22%), Positives = 84/201 (40%), Gaps = 8/201 (3%)
Query: 186 FCQEGKVKEAL----ELMEKGIKADANCFEILFDLCGKSKSVED--AKKVHDYFLQSTFR 239
+ + G+ KEAL +++++GI ++ F + C + SV +++H + ++
Sbjct: 77 YSRNGEHKEALVFLRDMVKEGIFSNQYAFVSVLRACQEIGSVGILFGRQIHGLMFKLSYA 136
Query: 240 SDFKMHNKVIEMYGNC-KSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQ 298
D + N +I MY C S+ A F + +N SW+ +I Y+ + ++F
Sbjct: 137 VDAVVSNVLISMYWKCIGSVGYALCAFGDIEVKNSVSWNSIISVYSQAGDQRSAFRIFSS 196
Query: 299 MNELGLEITSETM-LAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQS 357
M G T T V +AC E + K G+ + GL+ +S
Sbjct: 197 MQYDGSRPTEYTFGSLVTTACSLTEPDVRLLEQIMCTIQKSGLLTDLFVGSGLVSAFAKS 256
Query: 358 GYLKEAEEFIEQLPFEPTVTV 378
G L A + Q+ VT+
Sbjct: 257 GSLSYARKVFNQMETRNAVTL 277
Score = 41.2 bits (95), Expect = 0.002
Identities = 42/194 (21%), Positives = 80/194 (40%), Gaps = 8/194 (4%)
Query: 183 LTRFCQEGKVKEALELMEKGIKADA--NCFEILFDL--CGKSKSVEDAKKVHDYFLQSTF 238
+T Q G EA+E + + D F ++ L C K + +++H L+
Sbjct: 387 ITGLDQNGCFIEAVERYKSMRRHDILPGSFTLISSLSSCASLKWAKLGQQIHGESLKLGI 446
Query: 239 RSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMG-DEGLQLFE 297
+ + N ++ +Y + + R++F MP + SW+ +I A S E + F
Sbjct: 447 DLNVSVSNALMTLYAETGYLNECRKIFSSMPEHDQVSWNSIIGALARSERSLPEAVVCFL 506
Query: 298 QMNELGLEITSETMLAVLSACGSAEAVE-DAYIYLESMKSKYGIEPGVEHYMGLLDVLGQ 356
G ++ T +VLSA S E I+ ++K+ E E+ L+ G+
Sbjct: 507 NAQRAGQKLNRITFSSVLSAVSSLSFGELGKQIHGLALKNNIADEATTEN--ALIACYGK 564
Query: 357 SGYLKEAEEFIEQL 370
G + E+ ++
Sbjct: 565 CGEMDGCEKIFSRM 578
Score = 40.0 bits (92), Expect = 0.003
Identities = 28/134 (20%), Positives = 65/134 (47%), Gaps = 6/134 (4%)
Query: 235 QSTFRSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQ 294
+S +D + + ++ + S++ AR+VF+ M RN + + ++ G G+E +
Sbjct: 236 KSGLLTDLFVGSGLVSAFAKSGSLSYARKVFNQMETRNAVTLNGLMVGLVRQKWGEEATK 295
Query: 295 LFEQMNELGLEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYM-----G 349
LF MN + ++++ E+ + +LS+ E+ + + I G+ +M G
Sbjct: 296 LFMDMNSM-IDVSPESYVILLSSFPEYSLAEEVGLKKGREVHGHVITTGLVDFMVGIGNG 354
Query: 350 LLDVLGQSGYLKEA 363
L+++ + G + +A
Sbjct: 355 LVNMYAKCGSIADA 368
>At1g20230 hypothetical protein
Length = 760
Score = 244 bits (622), Expect = 1e-64
Identities = 127/396 (32%), Positives = 219/396 (55%), Gaps = 31/396 (7%)
Query: 188 QEGKVKEALELMEK----GIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFK 243
Q GK EALEL + G+K + + CG ++ + H + ++ +
Sbjct: 365 QNGKDIEALELFREMQVAGVKPNHVTIPSMLPACGNIAALGHGRSTHGFAVRVHLLDNVH 424
Query: 244 MHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELG 303
+ + +I+MY C + ++ VF+ MP +N+ W+ ++ G++ E + +FE +
Sbjct: 425 VGSALIDMYAKCGRINLSQIVFNMMPTKNLVCWNSLMNGFSMHGKAKEVMSIFESLMRTR 484
Query: 304 LEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEA 363
L+ + ++LSACG ++ + Y + M +YGI+P +EHY ++++LG++G L+EA
Sbjct: 485 LKPDFISFTSLLSACGQVGLTDEGWKYFKMMSEEYGIKPRLEHYSCMVNLLGRAGKLQEA 544
Query: 364 EEFIEQLPFEPTVTVFETLKNYARIHGDVDLEDHVEELIVSLDPSK-------------- 409
+ I+++PFEP V+ L N R+ +VDL + E + L+P
Sbjct: 545 YDLIKEMPFEPDSCVWGALLNSCRLQNNVDLAEIAAEKLFHLEPENPGTYVLLSNIYAAK 604
Query: 410 -------AVANKIPTPPPKKYTAISMLDGKNRII-----EYKNPTLYKDDEKLIAMNS-M 456
++ NK+ + KK S + KNR+ + +P + + EK+ ++ M
Sbjct: 605 GMWTEVDSIRNKMESLGLKKNPGCSWIQVKNRVYTLLAGDKSHPQIDQITEKMDEISKEM 664
Query: 457 KDAGYVPDTRYVLHDIDQEAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLRVCGDCH 516
+ +G+ P+ + LHD++++ +EQ L HSE+LA+ +GL++TP TPL++IKNLR+CGDCH
Sbjct: 665 RKSGHRPNLDFALHDVEEQEQEQMLWGHSEKLAVVFGLLNTPDGTPLQVIKNLRICGDCH 724
Query: 517 NAIKIMSRIVGRELIVRDNKRFHHFKDGKCSCGDYW 552
IK +S GRE+ +RD RFHHFKDG CSCGD+W
Sbjct: 725 AVIKFISSYAGREIFIRDTNRFHHFKDGICSCGDFW 760
Score = 52.4 bits (124), Expect = 7e-07
Identities = 29/137 (21%), Positives = 63/137 (45%)
Query: 198 LMEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNCKS 257
+ G+ D++ LF +C + + + K++H S D + + MY C
Sbjct: 107 MFSHGLIPDSHVLPNLFKVCAELSAFKVGKQIHCVSCVSGLDMDAFVQGSMFHMYMRCGR 166
Query: 258 MTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSETMLAVLSA 317
M DAR+VFD M ++++ + ++ YA +E +++ +M G+E + +LS
Sbjct: 167 MGDARKVFDRMSDKDVVTCSALLCAYARKGCLEEVVRILSEMESSGIEANIVSWNGILSG 226
Query: 318 CGSAEAVEDAYIYLESM 334
+ ++A + + +
Sbjct: 227 FNRSGYHKEAVVMFQKI 243
Score = 41.6 bits (96), Expect = 0.001
Identities = 32/165 (19%), Positives = 73/165 (43%), Gaps = 7/165 (4%)
Query: 183 LTRFCQEGKVKEALELMEK----GIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTF 238
L+ F + G KEA+ + +K G D + G S+ + + +H Y ++
Sbjct: 224 LSGFNRSGYHKEAVVMFQKIHHLGFCPDQVTVSSVLPSVGDSEMLNMGRLIHGYVIKQGL 283
Query: 239 RSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQ 298
D + + +I+MYG + +F+ + I G + + + D+ L++FE
Sbjct: 284 LKDKCVISAMIDMYGKSGHVYGIISLFNQFEMMEAGVCNAYITGLSRNGLVDKALEMFEL 343
Query: 299 MNELGLEITSETMLAVLSACG-SAEAVEDAYIYLESMKSKYGIEP 342
E +E+ + ++++ C + + +E ++ E + G++P
Sbjct: 344 FKEQTMELNVVSWTSIIAGCAQNGKDIEALELFREMQVA--GVKP 386
>At5g52630 selenium-binding protein-like
Length = 588
Score = 242 bits (617), Expect = 4e-64
Identities = 139/401 (34%), Positives = 219/401 (53%), Gaps = 38/401 (9%)
Query: 186 FCQEGKVKEAL----ELMEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSD 241
+ Q G+ +EAL E + + + + F + +C S +E +++H ++S+F S
Sbjct: 192 YAQMGENEEALWLFKEALFENLAVNDYSFSSVISVCANSTLLELGRQIHGLSIKSSFDSS 251
Query: 242 FKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNE 301
+ + ++ +Y C A +VF+ +P +N+ W+ M++ YA + + ++LF++M
Sbjct: 252 SFVGSSLVSLYSKCGVPEGAYQVFNEVPVKNLGIWNAMLKAYAQHSHTQKVIELFKRMKL 311
Query: 302 LGLEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLK 361
G++ T L VL+AC A V++ Y + MK IEP +HY L+D+LG++G L+
Sbjct: 312 SGMKPNFITFLNVLNACSHAGLVDEGRYYFDQMKESR-IEPTDKHYASLVDMLGRAGRLQ 370
Query: 362 EAEEFIEQLPFEPTVTVFETLKNYARIHGDVDLEDHVEELIVSLDP-------------- 407
EA E I +P +PT +V+ L +H + +L + + L P
Sbjct: 371 EALEVITNMPIDPTESVWGALLTSCTVHKNTELAAFAADKVFELGPVSSGMHISLSNAYA 430
Query: 408 ------SKAVANKIPTPP-PKKYTAISMLDGKNRIIEY--------KNPTLYKDDEKLIA 452
A A K+ KK T +S ++ +N++ + K+ +Y EKL
Sbjct: 431 ADGRFEDAAKARKLLRDRGEKKETGLSWVEERNKVHTFAAGERRHEKSKEIY---EKLAE 487
Query: 453 MNS-MKDAGYVPDTRYVLHDIDQEAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLRV 511
+ M+ AGY+ DT YVL ++D + K Q + YHSERLAIA+GLI+ P P+R++KNLRV
Sbjct: 488 LGEEMEKAGYIADTSYVLREVDGDEKNQTIRYHSERLAIAFGLITFPADRPIRVMKNLRV 547
Query: 512 CGDCHNAIKIMSRIVGRELIVRDNKRFHHFKDGKCSCGDYW 552
CGDCHNAIK MS R +IVRDN RFH F+DGKCSC DYW
Sbjct: 548 CGDCHNAIKFMSVCTRRVIIVRDNNRFHRFEDGKCSCNDYW 588
Score = 65.5 bits (158), Expect = 7e-11
Identities = 54/244 (22%), Positives = 106/244 (43%), Gaps = 10/244 (4%)
Query: 152 NGNLNQFQNPNNQFQTPNVQEQAPPPPSIV---DLTRFCQEGKVKEALELMEK----GIK 204
N +N + F + E +P S ++ F Q +LE ++K ++
Sbjct: 54 NNLINFYSKSQLPFDSRRAFEDSPQKSSTTWSSIISCFAQNELPWMSLEFLKKMMAGNLR 113
Query: 205 ADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNCKSMTDARRV 264
D + C + + VH +++ + +D + + +++MY C + AR++
Sbjct: 114 PDDHVLPSATKSCAILSRCDIGRSVHCLSMKTGYDADVFVGSSLVDMYAKCGEIVYARKM 173
Query: 265 FDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSETMLAVLSACGSAEAV 324
FD MP RN+ +W M+ GYA +E L LF++ L + + +V+S C ++ +
Sbjct: 174 FDEMPQRNVVTWSGMMYGYAQMGENEEALWLFKEALFENLAVNDYSFSSVISVCANSTLL 233
Query: 325 E-DAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLPFEPTVTVFETLK 383
E I+ S+KS + V L+ + + G + A + ++P + LK
Sbjct: 234 ELGRQIHGLSIKSSFDSSSFVG--SSLVSLYSKCGVPEGAYQVFNEVPVKNLGIWNAMLK 291
Query: 384 NYAR 387
YA+
Sbjct: 292 AYAQ 295
Score = 44.7 bits (104), Expect = 1e-04
Identities = 33/171 (19%), Positives = 74/171 (42%), Gaps = 3/171 (1%)
Query: 219 KSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHM 278
+++S ++H Y ++S + N +I Y + D+RR F+ P ++ +W
Sbjct: 27 RTRSTIKGLQLHGYVVKSGLSLIPLVANNLINFYSKSQLPFDSRRAFEDSPQKSSTTWSS 86
Query: 279 MIRGYANSTMGDEGLQLFEQMNELGLEITSETMLAVLSACGSAEAVE-DAYIYLESMKSK 337
+I +A + + L+ ++M L + + +C + ++ SMK+
Sbjct: 87 IISCFAQNELPWMSLEFLKKMMAGNLRPDDHVLPSATKSCAILSRCDIGRSVHCLSMKTG 146
Query: 338 YGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLPFEPTVTVFETLKNYARI 388
Y + V L+D+ + G + A + +++P VT + YA++
Sbjct: 147 YDADVFVG--SSLVDMYAKCGEIVYARKMFDEMPQRNVVTWSGMMYGYAQM 195
>At2g22070 hypothetical protein
Length = 786
Score = 242 bits (617), Expect = 4e-64
Identities = 147/397 (37%), Positives = 214/397 (53%), Gaps = 32/397 (8%)
Query: 188 QEGKVKEALELMEK----GIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFK 243
Q G EA+ L G + ++ + + S+ K++H ++S
Sbjct: 390 QHGSYGEAINLFRSMVGGGQRPNSYTLAAMLSVASSLASLSHGKQIHGSAVKSGEIYSVS 449
Query: 244 MHNKVIEMYGNCKSMTDARRVFDHMP-NRNMDSWHMMIRGYANSTMGDEGLQLFEQMNEL 302
+ N +I MY ++T A R FD + R+ SW MI A +E L+LFE M
Sbjct: 450 VSNALITMYAKAGNITSASRAFDLIRCERDTVSWTSMIIALAQHGHAEEALELFETMLME 509
Query: 303 GLEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKE 362
GL T + V SAC A V Y + MK I P + HY ++D+ G++G L+E
Sbjct: 510 GLRPDHITYVGVFSACTHAGLVNQGRQYFDMMKDVDKIIPTLSHYACMVDLFGRAGLLQE 569
Query: 363 AEEFIEQLPFEPTVTVFETLKNYARIHGDVDLEDHVEELIVSLDPSK-----AVAN---- 413
A+EFIE++P EP V + +L + R+H ++DL E ++ L+P A+AN
Sbjct: 570 AQEFIEKMPIEPDVVTWGSLLSACRVHKNIDLGKVAAERLLLLEPENSGAYSALANLYSA 629
Query: 414 ------------KIPTPPPKKYTAISMLDGKNRIIEY--KNPTLYKDDEKLIAM----NS 455
+ KK S ++ K+++ + ++ T + +E + M +
Sbjct: 630 CGKWEEAAKIRKSMKDGRVKKEQGFSWIEVKHKVHVFGVEDGTHPEKNEIYMTMKKIWDE 689
Query: 456 MKDAGYVPDTRYVLHDIDQEAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLRVCGDC 515
+K GYVPDT VLHD+++E KEQ L +HSE+LAIA+GLISTP +T LRI+KNLRVC DC
Sbjct: 690 IKKMGYVPDTASVLHDLEEEVKEQILRHHSEKLAIAFGLISTPDKTTLRIMKNLRVCNDC 749
Query: 516 HNAIKIMSRIVGRELIVRDNKRFHHFKDGKCSCGDYW 552
H AIK +S++VGRE+IVRD RFHHFKDG CSC DYW
Sbjct: 750 HTAIKFISKLVGREIIVRDTTRFHHFKDGFCSCRDYW 786
Score = 52.8 bits (125), Expect = 5e-07
Identities = 30/117 (25%), Positives = 58/117 (48%), Gaps = 1/117 (0%)
Query: 197 ELMEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNCK 256
+++++GI+ + ++ +E KKVH + ++ R + + N ++ MY C
Sbjct: 136 DMVKEGIEPTQFTLTNVLASVAATRCMETGKKVHSFIVKLGLRGNVSVSNSLLNMYAKCG 195
Query: 257 SMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSETMLA 313
A+ VFD M R++ SW+ MI + D + FEQM E + +T +M++
Sbjct: 196 DPMMAKFVFDRMVVRDISSWNAMIALHMQVGQMDLAMAQFEQMAERDI-VTWNSMIS 251
Score = 46.2 bits (108), Expect = 5e-05
Identities = 46/198 (23%), Positives = 78/198 (39%), Gaps = 18/198 (9%)
Query: 174 APPPPSIVDLTRFC----------QEGKVKEAL---ELMEKGIKADANCFEILFDLCGKS 220
AP P S+ L C G+ L +++ G+ L ++ K+
Sbjct: 3 APVPLSLSTLLELCTNLLQKSVNKSNGRFTAQLVHCRVIKSGLMFSVYLMNNLMNVYSKT 62
Query: 221 KSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMI 280
A+K+ D + R+ F N V+ Y M FD +P R+ SW MI
Sbjct: 63 GYALHARKLFD---EMPLRTAFSW-NTVLSAYSKRGDMDSTCEFFDQLPQRDSVSWTTMI 118
Query: 281 RGYANSTMGDEGLQLFEQMNELGLEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGI 340
GY N + +++ M + G+E T T+ VL++ + +E + S K G+
Sbjct: 119 VGYKNIGQYHKAIRVMGDMVKEGIEPTQFTLTNVLASVAATRCMETGK-KVHSFIVKLGL 177
Query: 341 EPGVEHYMGLLDVLGQSG 358
V LL++ + G
Sbjct: 178 RGNVSVSNSLLNMYAKCG 195
Score = 40.0 bits (92), Expect = 0.003
Identities = 54/260 (20%), Positives = 89/260 (33%), Gaps = 69/260 (26%)
Query: 183 LTRFCQEGKVKEALELMEKGIK-----ADANCFEILFDLCGKSKSVEDAKKVHDYFLQST 237
++ F Q G AL++ K ++ D + C + + K++H + + +
Sbjct: 250 ISGFNQRGYDLRALDIFSKMLRDSLLSPDRFTLASVLSACANLEKLCIGKQIHSHIVTTG 309
Query: 238 FRSDFKMHNKVIEMYGNCKSMTDARR---------------------------------V 264
F + N +I MY C + ARR +
Sbjct: 310 FDISGIVLNALISMYSRCGGVETARRLIEQRGTKDLKIEGFTALLDGYIKLGDMNQAKNI 369
Query: 265 FDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSETMLAVLSACGS---- 320
F + +R++ +W MI GY E + LF M G S T+ A+LS S
Sbjct: 370 FVSLKDRDVVAWTAMIVGYEQHGSYGEAINLFRSMVGGGQRPNSYTLAAMLSVASSLASL 429
Query: 321 -------AEAVEDAYIY-------LESMKSKYG-------------IEPGVEHYMGLLDV 353
AV+ IY L +M +K G E + ++
Sbjct: 430 SHGKQIHGSAVKSGEIYSVSVSNALITMYAKAGNITSASRAFDLIRCERDTVSWTSMIIA 489
Query: 354 LGQSGYLKEAEEFIEQLPFE 373
L Q G+ +EA E E + E
Sbjct: 490 LAQHGHAEEALELFETMLME 509
>At4g37380 putative protein
Length = 632
Score = 240 bits (612), Expect = 2e-63
Identities = 137/402 (34%), Positives = 213/402 (52%), Gaps = 37/402 (9%)
Query: 186 FCQEGKVKEALELMEKGI-----KADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRS 240
+ Q G +AL L +K + K D C + ++E + +H + S R
Sbjct: 233 YAQHGFPNDALMLFQKLLAEGKPKPDEITVVAALSACSQIGALETGRWIHVFVKSSRIRL 292
Query: 241 DFKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMN 300
+ K+ +I+MY C S+ +A VF+ P +++ +W+ MI GYA + L+LF +M
Sbjct: 293 NVKVCTGLIDMYSKCGSLEEAVLVFNDTPRKDIVAWNAMIAGYAMHGYSQDALRLFNEMQ 352
Query: 301 EL-GLEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGY 359
+ GL+ T T + L AC A V + ESM +YGI+P +EHY L+ +LG++G
Sbjct: 353 GITGLQPTDITFIGTLQACAHAGLVNEGIRIFESMGQEYGIKPKIEHYGCLVSLLGRAGQ 412
Query: 360 LKEAEEFIEQLPFEPTVTVFETLKNYARIHGDVDLEDHVEELIVSLDPSKA--------- 410
LK A E I+ + + ++ ++ ++HGD L + E ++ L+ +
Sbjct: 413 LKRAYETIKNMNMDADSVLWSSVLGSCKLHGDFVLGKEIAEYLIGLNIKNSGIYVLLSNI 472
Query: 411 ------------VANKIPTPPPKKYTAISMLDGKNRIIEY--------KNPTLYKDDEKL 450
V N + K IS ++ +N++ E+ K+ +Y K+
Sbjct: 473 YASVGDYEGVAKVRNLMKEKGIVKEPGISTIEIENKVHEFRAGDREHSKSKEIYTMLRKI 532
Query: 451 IAMNSMKDAGYVPDTRYVLHDIDQEAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLR 510
+K GYVP+T VL D+++ KEQ+L HSERLAIAYGLIST P +PL+I KNLR
Sbjct: 533 --SERIKSHGYVPNTNTVLQDLEETEKEQSLQVHSERLAIAYGLISTKPGSPLKIFKNLR 590
Query: 511 VCGDCHNAIKIMSRIVGRELIVRDNKRFHHFKDGKCSCGDYW 552
VC DCH K++S+I GR++++RD RFHHF DG CSCGD+W
Sbjct: 591 VCSDCHTVTKLISKITGRKIVMRDRNRFHHFTDGSCSCGDFW 632
Score = 65.1 bits (157), Expect = 1e-10
Identities = 56/197 (28%), Positives = 93/197 (46%), Gaps = 13/197 (6%)
Query: 198 LMEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNCKS 257
+++ G+ D L D+ K V A+KV D + + S M I Y +
Sbjct: 152 VLKFGLGIDPYVATGLVDVYAKGGDVVSAQKVFDRMPERSLVSSTAM----ITCYAKQGN 207
Query: 258 MTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSE-TMLAVLS 316
+ AR +FD M R++ SW++MI GYA ++ L LF+++ G E T++A LS
Sbjct: 208 VEAARALFDSMCERDIVSWNVMIDGYAQHGFPNDALMLFQKLLAEGKPKPDEITVVAALS 267
Query: 317 ACGSAEAVEDA---YIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLPFE 373
AC A+E +++++S + I V+ GL+D+ + G L+EA P +
Sbjct: 268 ACSQIGALETGRWIHVFVKSSR----IRLNVKVCTGLIDMYSKCGSLEEAVLVFNDTPRK 323
Query: 374 PTVTVFETLKNYARIHG 390
V + YA +HG
Sbjct: 324 DIVAWNAMIAGYA-MHG 339
>At1g56690 hypothetical protein
Length = 704
Score = 239 bits (611), Expect = 2e-63
Identities = 126/397 (31%), Positives = 215/397 (53%), Gaps = 32/397 (8%)
Query: 188 QEGKVKEALELM----EKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFK 243
++G EAL+L ++G++ + +C S++ ++VH + ++ F D
Sbjct: 308 RKGFELEALDLFAQMQKQGVRPSFPSLISILSVCATLASLQYGRQVHAHLVRCQFDDDVY 367
Query: 244 MHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELG 303
+ + ++ MY C + A+ VFD ++++ W+ +I GYA+ +G+E L++F +M G
Sbjct: 368 VASVLMTMYVKCGELVKAKLVFDRFSSKDIIMWNSIISGYASHGLGEEALKIFHEMPSSG 427
Query: 304 LEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEA 363
T++A+L+AC A +E+ ESM+SK+ + P VEHY +D+LG++G + +A
Sbjct: 428 TMPNKVTLIAILTACSYAGKLEEGLEIFESMESKFCVTPTVEHYSCTVDMLGRAGQVDKA 487
Query: 364 EEFIEQLPFEPTVTVFETLKNYARIHGDVDLEDHVEELIVSLDPSKA------------- 410
E IE + +P TV+ L + H +DL + + + +P A
Sbjct: 488 MELIESMTIKPDATVWGALLGACKTHSRLDLAEVAAKKLFENEPDNAGTYVLLSSINASR 547
Query: 411 --------VANKIPTPPPKKYTAISMLDGKNRIIEYKNPTLYKDDEKLIAM-------NS 455
V + T K+ S ++ ++ + + E+ + +
Sbjct: 548 SKWGDVAVVRKNMRTNNVSKFPGCSWIEVGKKVHMFTRGGIKNHPEQAMILMMLEKTDGL 607
Query: 456 MKDAGYVPDTRYVLHDIDQEAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLRVCGDC 515
+++AGY PD +VLHD+D+E K +L HSERLA+AYGL+ P P+R++KNLRVCGDC
Sbjct: 608 LREAGYSPDCSHVLHDVDEEEKVDSLSRHSERLAVAYGLLKLPEGVPIRVMKNLRVCGDC 667
Query: 516 HNAIKIMSRIVGRELIVRDNKRFHHFKDGKCSCGDYW 552
H AIK++S++ RE+I+RD RFHHF +G+CSC DYW
Sbjct: 668 HAAIKLISKVTEREIILRDANRFHHFNNGECSCRDYW 704
Score = 51.6 bits (122), Expect = 1e-06
Identities = 25/80 (31%), Positives = 45/80 (56%)
Query: 246 NKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLE 305
N +I +G ++ ARRVFD M +R+ +W MI+ Y E L LF QM + G+
Sbjct: 269 NAMIVGFGEVGEISKARRVFDLMEDRDNATWRGMIKAYERKGFELEALDLFAQMQKQGVR 328
Query: 306 ITSETMLAVLSACGSAEAVE 325
+ +++++LS C + +++
Sbjct: 329 PSFPSLISILSVCATLASLQ 348
Score = 48.1 bits (113), Expect = 1e-05
Identities = 50/204 (24%), Positives = 86/204 (41%), Gaps = 17/204 (8%)
Query: 186 FCQEGKVKEALELMEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMH 245
+ QEG V EA L + + + + ++F ++ A+K++D + M
Sbjct: 120 YMQEGMVGEAESLFWRMPERNEVSWTVMFGGLIDDGRIDKARKLYDMMPVKDVVASTNM- 178
Query: 246 NKVIEMYGNCKS--MTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELG 303
+ G C+ + +AR +FD M RN+ +W MI GY + D +LFE M E
Sbjct: 179 -----IGGLCREGRVDEARLIFDEMRERNVVTWTTMITGYRQNNRVDVARKLFEVMPE-K 232
Query: 304 LEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEA 363
E++ +ML + G +EDA + E M K I ++ G+ G + +A
Sbjct: 233 TEVSWTSMLLGYTLSG---RIEDAEEFFEVMPMKPVIACN-----AMIVGFGEVGEISKA 284
Query: 364 EEFIEQLPFEPTVTVFETLKNYAR 387
+ + T +K Y R
Sbjct: 285 RRVFDLMEDRDNATWRGMIKAYER 308
Score = 42.7 bits (99), Expect = 5e-04
Identities = 21/56 (37%), Positives = 31/56 (54%)
Query: 246 NKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNE 301
N ++ Y + + +AR VF+ MP RN+ SW M++GY M E LF +M E
Sbjct: 83 NGLVSGYIKNRMIVEARNVFELMPERNVVSWTAMVKGYMQEGMVGEAESLFWRMPE 138
Score = 39.3 bits (90), Expect = 0.006
Identities = 19/56 (33%), Positives = 31/56 (54%)
Query: 246 NKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNE 301
N ++ Y + +AR++FD M RN+ SW+ ++ GY + M E +FE M E
Sbjct: 52 NSIVSGYFSNGLPKEARQLFDEMSERNVVSWNGLVSGYIKNRMIVEARNVFELMPE 107
Score = 36.6 bits (83), Expect = 0.037
Identities = 14/44 (31%), Positives = 29/44 (65%)
Query: 258 MTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNE 301
+ +AR+ FD + + + SW+ ++ GY ++ + E QLF++M+E
Sbjct: 33 INEARKFFDSLQFKAIGSWNSIVSGYFSNGLPKEARQLFDEMSE 76
>At1g11290 hypothetical protein
Length = 809
Score = 236 bits (602), Expect = 2e-62
Identities = 133/398 (33%), Positives = 211/398 (52%), Gaps = 32/398 (8%)
Query: 186 FCQEGKVKEAL----ELMEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSD 241
F Q G+ +AL ++ + +K D + + + AK +H ++S +
Sbjct: 413 FAQNGRPIDALNYFSQMRSRTVKPDTFTYVSVITAIAELSITHHAKWIHGVVMRSCLDKN 472
Query: 242 FKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNE 301
+ +++MY C ++ AR +FD M R++ +W+ MI GY G L+LFE+M +
Sbjct: 473 VFVTTALVDMYAKCGAIMIARLIFDMMSERHVTTWNAMIDGYGTHGFGKAALELFEEMQK 532
Query: 302 LGLEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLK 361
++ T L+V+SAC + VE MK Y IE ++HY ++D+LG++G L
Sbjct: 533 GTIKPNGVTFLSVISACSHSGLVEAGLKCFYMMKENYSIELSMDHYGAMVDLLGRAGRLN 592
Query: 362 EAEEFIEQLPFEPTVTVFETLKNYARIHGDVDLEDHVEELIVSLDPSKA----------- 410
EA +FI Q+P +P V V+ + +IH +V+ + E + L+P
Sbjct: 593 EAWDFIMQMPVKPAVNVYGAMLGACQIHKNVNFAEKAAERLFELNPDDGGYHVLLANIYR 652
Query: 411 ----------VANKIPTPPPKKYTAISMLDGKNRIIE-YKNPTLYKDDEKLIA-----MN 454
V + +K SM++ KN + + T + D +K+ A +
Sbjct: 653 AASMWEKVGQVRVSMLRQGLRKTPGCSMVEIKNEVHSFFSGSTAHPDSKKIYAFLEKLIC 712
Query: 455 SMKDAGYVPDTRYVLHDIDQEAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLRVCGD 514
+K+AGYVPDT VL ++ + KEQ L HSE+LAI++GL++T T + + KNLRVC D
Sbjct: 713 HIKEAGYVPDTNLVL-GVENDVKEQLLSTHSEKLAISFGLLNTTAGTTIHVRKNLRVCAD 771
Query: 515 CHNAIKIMSRIVGRELIVRDNKRFHHFKDGKCSCGDYW 552
CHNA K +S + GRE++VRD +RFHHFK+G CSCGDYW
Sbjct: 772 CHNATKYISLVTGREIVVRDMQRFHHFKNGACSCGDYW 809
Score = 73.9 bits (180), Expect = 2e-13
Identities = 43/160 (26%), Positives = 81/160 (49%), Gaps = 3/160 (1%)
Query: 210 FEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNCKSMTDARRVFDHMP 269
F L +CG + K++H ++S F D + MY C+ + +AR+VFD MP
Sbjct: 138 FTYLLKVCGDEAELRVGKEIHGLLVKSGFSLDLFAMTGLENMYAKCRQVNEARKVFDRMP 197
Query: 270 NRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSETMLAVLSACGSAEAVE-DAY 328
R++ SW+ ++ GY+ + M L++ + M E L+ + T+++VL A + +
Sbjct: 198 ERDLVSWNTIVAGYSQNGMARMALEMVKSMCEENLKPSFITIVSVLPAVSALRLISVGKE 257
Query: 329 IYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIE 368
I+ +M+S G + V L+D+ + G L+ A + +
Sbjct: 258 IHGYAMRS--GFDSLVNISTALVDMYAKCGSLETARQLFD 295
Score = 70.5 bits (171), Expect = 2e-12
Identities = 41/172 (23%), Positives = 83/172 (47%), Gaps = 5/172 (2%)
Query: 186 FCQEGKVKEALELM----EKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSD 241
+ Q G + ALE++ E+ +K + + + K++H Y ++S F S
Sbjct: 211 YSQNGMARMALEMVKSMCEENLKPSFITIVSVLPAVSALRLISVGKEIHGYAMRSGFDSL 270
Query: 242 FKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNE 301
+ +++MY C S+ AR++FD M RN+ SW+ MI Y + E + +F++M +
Sbjct: 271 VNISTALVDMYAKCGSLETARQLFDGMLERNVVSWNSMIDAYVQNENPKEAMLIFQKMLD 330
Query: 302 LGLEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDV 353
G++ T +++ L AC +E ++ + + G++ V L+ +
Sbjct: 331 EGVKPTDVSVMGALHACADLGDLERGR-FIHKLSVELGLDRNVSVVNSLISM 381
Score = 52.8 bits (125), Expect = 5e-07
Identities = 42/209 (20%), Positives = 89/209 (42%), Gaps = 6/209 (2%)
Query: 186 FCQEGKVKEAL----ELMEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSD 241
+ Q KEA+ +++++G+K C +E + +H ++ +
Sbjct: 312 YVQNENPKEAMLIFQKMLDEGVKPTDVSVMGALHACADLGDLERGRFIHKLSVELGLDRN 371
Query: 242 FKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNE 301
+ N +I MY CK + A +F + +R + SW+ MI G+A + + L F QM
Sbjct: 372 VSVVNSLISMYCKCKEVDTAASMFGKLQSRTLVSWNAMILGFAQNGRPIDALNYFSQMRS 431
Query: 302 LGLEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLK 361
++ + T ++V++A A ++ + + ++ V L+D+ + G +
Sbjct: 432 RTVKPDTFTYVSVITAIAELSITHHAK-WIHGVVMRSCLDKNVFVTTALVDMYAKCGAIM 490
Query: 362 EAEEFIEQLPFEPTVTVFETLKNYARIHG 390
A + + E VT + + + HG
Sbjct: 491 IARLIFDMMS-ERHVTTWNAMIDGYGTHG 518
Score = 42.0 bits (97), Expect = 9e-04
Identities = 36/185 (19%), Positives = 82/185 (43%), Gaps = 8/185 (4%)
Query: 207 ANCFE----ILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNCKSMTDAR 262
AN +E +L + C S+++ +++ ++ + K++ ++ S+ +A
Sbjct: 33 ANVYEHPAALLLERCS---SLKELRQILPLVFKNGLYQEHFFQTKLVSLFCRYGSVDEAA 89
Query: 263 RVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSETMLAVLSACGSAE 322
RVF+ + ++ +H M++G+A + D+ LQ F +M +E +L CG E
Sbjct: 90 RVFEPIDSKLNVLYHTMLKGFAKVSDLDKALQFFVRMRYDDVEPVVYNFTYLLKVCGD-E 148
Query: 323 AVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLPFEPTVTVFETL 382
A + + K G + GL ++ + + EA + +++P V+ +
Sbjct: 149 AELRVGKEIHGLLVKSGFSLDLFAMTGLENMYAKCRQVNEARKVFDRMPERDLVSWNTIV 208
Query: 383 KNYAR 387
Y++
Sbjct: 209 AGYSQ 213
>At1g68930 hypothetical protein
Length = 743
Score = 236 bits (601), Expect = 3e-62
Identities = 140/396 (35%), Positives = 206/396 (51%), Gaps = 31/396 (7%)
Query: 188 QEGKVKEA----LELMEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFK 243
Q G+ +EA L++ GI D C S+E+ + H + S
Sbjct: 348 QTGRAEEAVKIFLDMQRSGIDPDHYTLGQAISACANVSSLEEGSQFHGKAITSGLIHYVT 407
Query: 244 MHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELG 303
+ N ++ +YG C + D+ R+F+ M R+ SW M+ YA E +QLF++M + G
Sbjct: 408 VSNSLVTLYGKCGDIDDSTRLFNEMNVRDAVSWTAMVSAYAQFGRAVETIQLFDKMVQHG 467
Query: 304 LEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEA 363
L+ T+ V+SAC A VE Y + M S+YGI P + HY ++D+ +SG L+EA
Sbjct: 468 LKPDGVTLTGVISACSRAGLVEKGQRYFKLMTSEYGIVPSIGHYSCMIDLFSRSGRLEEA 527
Query: 364 EEFIEQLPFEPTVTVFETLKNYARIHGDVDLEDHVEELIVSLDP---------SKAVANK 414
FI +PF P + TL + R G++++ E ++ LDP S A+K
Sbjct: 528 MRFINGMPFPPDAIGWTTLLSACRNKGNLEIGKWAAESLIELDPHHPAGYTLLSSIYASK 587
Query: 415 ------------IPTPPPKKYTAISMLDGKNRIIEYK-----NPTLYKDDEKLIAMNS-M 456
+ KK S + K ++ + +P L + KL +N+ +
Sbjct: 588 GKWDSVAQLRRGMREKNVKKEPGQSWIKWKGKLHSFSADDESSPYLDQIYAKLEELNNKI 647
Query: 457 KDAGYVPDTRYVLHDIDQEAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLRVCGDCH 516
D GY PDT +V HD+++ K + L YHSERLAIA+GLI P P+R+ KNLRVC DCH
Sbjct: 648 IDNGYKPDTSFVHHDVEEAVKVKMLNYHSERLAIAFGLIFVPSGQPIRVGKNLRVCVDCH 707
Query: 517 NAIKIMSRIVGRELIVRDNKRFHHFKDGKCSCGDYW 552
NA K +S + GRE++VRD RFH FKDG CSCGD+W
Sbjct: 708 NATKHISSVTGREILVRDAVRFHRFKDGTCSCGDFW 743
Score = 85.9 bits (211), Expect = 5e-17
Identities = 51/208 (24%), Positives = 99/208 (47%), Gaps = 13/208 (6%)
Query: 188 QEGKVKEALELMEK----GIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFK 243
Q G KEA+E + G+K D F + CG ++ + K++H +++ F+
Sbjct: 247 QNGLAKEAIECFREMKVQGLKMDQYPFGSVLPACGGLGAINEGKQIHACIIRTNFQDHIY 306
Query: 244 MHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELG 303
+ + +I+MY CK + A+ VFD M +N+ SW M+ GY + +E +++F M G
Sbjct: 307 VGSALIDMYCKCKCLHYAKTVFDRMKQKNVVSWTAMVVGYGQTGRAEEAVKIFLDMQRSG 366
Query: 304 LEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYM----GLLDVLGQSGY 359
++ T+ +SAC + ++E+ S I G+ HY+ L+ + G+ G
Sbjct: 367 IDPDHYTLGQAISACANVSSLEEG-----SQFHGKAITSGLIHYVTVSNSLVTLYGKCGD 421
Query: 360 LKEAEEFIEQLPFEPTVTVFETLKNYAR 387
+ ++ ++ V+ + YA+
Sbjct: 422 IDDSTRLFNEMNVRDAVSWTAMVSAYAQ 449
Score = 47.0 bits (110), Expect = 3e-05
Identities = 27/102 (26%), Positives = 50/102 (48%), Gaps = 1/102 (0%)
Query: 227 KKVHDYFLQSTFRSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANS 286
K +H +++ + ++N ++ Y KS T ARRVFD +P N+ SW+ ++ Y+ +
Sbjct: 26 KMIHGNIIRALPYPETFLYNNIVHAYALMKSSTYARRVFDRIPQPNLFSWNNLLLAYSKA 85
Query: 287 TMGDEGLQLFEQMNELGLEITSETMLAVLSACGSAEAVEDAY 328
+ E FE++ + +T ++ S G A AY
Sbjct: 86 GLISEMESTFEKLPDRD-GVTWNVLIEGYSLSGLVGAAVKAY 126
Score = 46.6 bits (109), Expect = 4e-05
Identities = 32/144 (22%), Positives = 59/144 (40%), Gaps = 30/144 (20%)
Query: 213 LFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRN 272
+ L + V K++H ++ F S + + ++ MY N ++DA++VF + +RN
Sbjct: 145 MLKLSSSNGHVSLGKQIHGQVIKLGFESYLLVGSPLLYMYANVGCISDAKKVFYGLDDRN 204
Query: 273 ------------------------------MDSWHMMIRGYANSTMGDEGLQLFEQMNEL 302
SW MI+G A + + E ++ F +M
Sbjct: 205 TVMYNSLMGGLLACGMIEDALQLFRGMEKDSVSWAAMIKGLAQNGLAKEAIECFREMKVQ 264
Query: 303 GLEITSETMLAVLSACGSAEAVED 326
GL++ +VL ACG A+ +
Sbjct: 265 GLKMDQYPFGSVLPACGGLGAINE 288
>At1g29710
Length = 475
Score = 229 bits (583), Expect = 4e-60
Identities = 136/417 (32%), Positives = 220/417 (52%), Gaps = 23/417 (5%)
Query: 151 QNGNLNQFQNPNNQFQTPNVQEQAPPPPSIVDLTRFCQEGKVKEALELME----KGIKAD 206
QN + Q++ + NV +I C +G +EA+E+++ KG D
Sbjct: 67 QNSMVGQYKTTVSPSVAQNV--------TIETFDSLCIQGNWREAVEVLDYLENKGYAMD 118
Query: 207 ANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNCKSMTDARRVFD 266
L LCGK +++E A+ VH+ + D N +IEMY C S+ DA +VF+
Sbjct: 119 LIRLLGLAKLCGKPEALEAARVVHECIIALVSPCDVGARNAIIEMYSGCCSVDDALKVFE 178
Query: 267 HMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSETMLAVLSACGSAEAVED 326
MP N + +M+R + N+ G+E + LF + E G + E V S C V++
Sbjct: 179 EMPEWNSGTLCVMMRCFVNNGYGEEAIDLFTRFKEEGNKPNGEIFNQVFSTCTLTGDVKE 238
Query: 327 AYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLPFEPTVTVFETLKNYA 386
+ ++M +YGI P +EHY + +L SG+L EA F+E++P EP+V V+ETL N +
Sbjct: 239 GSLQFQAMYREYGIVPSMEHYHSVTKMLATSGHLDEALNFVERMPMEPSVDVWETLMNLS 298
Query: 387 RIHGDVDLEDHVEELIVSLDPSK--------AVANKIPTPPPKKYTAIS--MLDGKNRII 436
R+HGDV+L D EL+ LD ++ VA K K+ + S R +
Sbjct: 299 RVHGDVELGDRCAELVEKLDATRLDKVSSAGLVATKASDFVKKEPSTRSEPYFYSTFRPV 358
Query: 437 EYKNPTLYKDDEKLIAMNS-MKDAGYVPDTRYVLHDIDQEAKEQALLYHSERLAIAYGLI 495
+ +P + E L+++ S +K+ GYVPDTRY I ++ + + E +A+ L+
Sbjct: 359 DSSHPQMNIIYETLMSLRSQLKEMGYVPDTRYYRSLIMAMENKEQIFGYREEIAVVESLL 418
Query: 496 STPPRTPLRIIKNLRVCGDCHNAIKIMSRIVGRELIVRDNKRFHHFKDGKCSCGDYW 552
+ PR+ + ++ N+R+ GDCH+ +K+MS I GR++I RD K +H FK+G C C + W
Sbjct: 419 KSKPRSAITLLTNIRIVGDCHDMMKLMSVITGRDMIKRDAKIYHLFKNGVCRCNNLW 475
>At2g03880 putative selenium-binding protein
Length = 630
Score = 228 bits (581), Expect = 7e-60
Identities = 128/398 (32%), Positives = 213/398 (53%), Gaps = 33/398 (8%)
Query: 186 FCQEGKVKEALELMEK----GIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSD 241
F Q + ALEL ++ G A+ + C +E + H + ++ + D
Sbjct: 235 FAQNSRSDVALELFKRMKRAGFIAEQATLTSVLRACTGLALLELGMQAHVHIVK--YDQD 292
Query: 242 FKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNE 301
++N +++MY C S+ DA RVF+ M R++ +W MI G A + E L+LFE+M
Sbjct: 293 LILNNALVDMYCKCGSLEDALRVFNQMKERDVITWSTMISGLAQNGYSQEALKLFERMKS 352
Query: 302 LGLEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLK 361
G + T++ VL AC A +ED + Y SMK YGI+P EHY ++D+LG++G L
Sbjct: 353 SGTKPNYITIVGVLFACSHAGLLEDGWYYFRSMKKLYGIDPVREHYGCMIDLLGKAGKLD 412
Query: 362 EAEEFIEQLPFEPTVTVFETLKNYARIHGDVDLEDHVEELIVSLDPSKA----------- 410
+A + + ++ EP + TL R+ ++ L ++ + +++LDP A
Sbjct: 413 DAVKLLNEMECEPDAVTWRTLLGACRVQRNMVLAEYAAKKVIALDPEDAGTYTLLSNIYA 472
Query: 411 ----------VANKIPTPPPKKYTAISMLDGKNRIIEY-----KNPTLYKDDEKLIAM-N 454
+ ++ KK S ++ +I + +P + + +KL + +
Sbjct: 473 NSQKWDSVEEIRTRMRDRGIKKEPGCSWIEVNKQIHAFIIGDNSHPQIVEVSKKLNQLIH 532
Query: 455 SMKDAGYVPDTRYVLHDIDQEAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLRVCGD 514
+ GYVP+T +VL D++ E E +L +HSE+LA+A+GL++ P +RI KNLR+CGD
Sbjct: 533 RLTGIGYVPETNFVLQDLEGEQMEDSLRHHSEKLALAFGLMTLPIEKVIRIRKNLRICGD 592
Query: 515 CHNAIKIMSRIVGRELIVRDNKRFHHFKDGKCSCGDYW 552
CH K+ S++ R +++RD R+HHF+DGKCSCGDYW
Sbjct: 593 CHVFCKLASKLEIRSIVIRDPIRYHHFQDGKCSCGDYW 630
Score = 64.3 bits (155), Expect = 2e-10
Identities = 47/193 (24%), Positives = 84/193 (43%), Gaps = 8/193 (4%)
Query: 182 DLTRFCQEGKVKEALELMEK----GIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQST 237
+ TR C + + A++ M+ G+ AD+ + L C +++V + + + +
Sbjct: 32 EFTRLCYQRDLPRAMKAMDSLQSHGLWADSATYSELIKCCISNRAVHEGNLICRHLYFNG 91
Query: 238 FRSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFE 297
R + N +I MY + DA ++FD MP RN+ SW MI Y+ + + L+L
Sbjct: 92 HRPMMFLVNVLINMYVKFNLLNDAHQLFDQMPQRNVISWTTMISAYSKCKIHQKALELLV 151
Query: 298 QMNELGLEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQS 357
M + T +VL +C V L K G+E V L+DV +
Sbjct: 152 LMLRDNVRPNVYTYSSVLRSCNGMSDVR----MLHCGIIKEGLESDVFVRSALIDVFAKL 207
Query: 358 GYLKEAEEFIEQL 370
G ++A +++
Sbjct: 208 GEPEDALSVFDEM 220
Score = 53.9 bits (128), Expect = 2e-07
Identities = 37/158 (23%), Positives = 74/158 (46%), Gaps = 9/158 (5%)
Query: 223 VEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRG 282
+ D + +H ++ SD + + +I+++ DA VFD M + W+ +I G
Sbjct: 175 MSDVRMLHCGIIKEGLESDVFVRSALIDVFAKLGEPEDALSVFDEMVTGDAIVWNSIIGG 234
Query: 283 YANSTMGDEGLQLFEQMNELGLEITSETMLAVLSACGSAEAVE---DAYIYLESMKSKYG 339
+A ++ D L+LF++M G T+ +VL AC +E A++++ KY
Sbjct: 235 FAQNSRSDVALELFKRMKRAGFIAEQATLTSVLRACTGLALLELGMQAHVHI----VKYD 290
Query: 340 IEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLPFEPTVT 377
+ + + L+D+ + G L++A Q+ +T
Sbjct: 291 QDLILNN--ALVDMYCKCGSLEDALRVFNQMKERDVIT 326
>At2g34370 putative selenium-binding protein
Length = 469
Score = 227 bits (578), Expect = 1e-59
Identities = 137/388 (35%), Positives = 214/388 (54%), Gaps = 28/388 (7%)
Query: 187 CQEGKVKEALELME----KGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDF 242
C++ K++EALE+++ KG D L LCG+ +++E+A+ VHD RS
Sbjct: 88 CKQVKIREALEVIDILEDKGYIVDFPRLLGLAKLCGEVEALEEARVVHDCITPLDARS-- 145
Query: 243 KMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNEL 302
++ VIEMY C+S DA VF+ MP RN ++W MIR A + G+ + +F + E
Sbjct: 146 --YHTVIEMYSGCRSTDDALNVFNEMPKRNSETWGTMIRCLAKNGEGERAIDMFTRFIEE 203
Query: 303 GLEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKE 362
G + E AV AC S + + ++ ESM YG+ +E Y+ ++++L G+L E
Sbjct: 204 GNKPDKEIFKAVFFACVSIGDINEGLLHFESMYRDYGMVLSMEDYVNVIEMLAACGHLDE 263
Query: 363 AEEFIEQLPFEPTVTVFETLKNYARIHGDVDLEDHVEELIVSLDPSK--------AVANK 414
A +F+E++ EP+V ++ETL N + G ++L D ELI LD S+ VA K
Sbjct: 264 ALDFVERMTVEPSVEMWETLMNLCWVQGYLELGDRFAELIKKLDASRMSKESNAGLVAAK 323
Query: 415 IPTPPPKK-----YTAISMLDGKNRIIEYK-NPTLYKDDEKLIAMNSMK----DAGYVPD 464
+K Y + D K R+ E++ T + + A S+K D G+VP
Sbjct: 324 ASDSAMEKLKELRYCQMIRDDPKKRMHEFRAGDTSHLG--TVSAFRSLKVQMLDIGFVPA 381
Query: 465 TRYVLHDIDQEAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLRVCGDCHNAIKIMSR 524
TR +++E KE+ LL+ S +LA A+ +I++ R PL +++N+R C D HN K++S
Sbjct: 382 TRVCFVTVEEEEKEEQLLFRSNKLAFAHAIINSEARRPLTVLQNMRTCIDGHNTFKMISL 441
Query: 525 IVGRELIVRDNKRFHHFKDGKCSCGDYW 552
I GR LI RD K++H +K+G CSC DYW
Sbjct: 442 ITGRALIQRDKKKYHFYKNGVCSCKDYW 469
>At4g18750 putative protein
Length = 871
Score = 223 bits (568), Expect = 2e-58
Identities = 124/384 (32%), Positives = 201/384 (52%), Gaps = 27/384 (7%)
Query: 196 LELMEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNC 255
L L EK D + C + + +++H Y +++ + SD + N +++MY C
Sbjct: 488 LLLEEKRFSPDERTVACVLPACASLSAFDKGREIHGYIMRNGYFSDRHVANSLVDMYAKC 547
Query: 256 KSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSETMLAVL 315
++ A +FD + ++++ SW +MI GY G E + LF QM + G+E + +++L
Sbjct: 548 GALLLAHMLFDDIASKDLVSWTVMIAGYGMHGFGKEAIALFNQMRQAGIEADEISFVSLL 607
Query: 316 SACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLPFEPT 375
AC + V++ + + M+ + IEP VEHY ++D+L ++G L +A FIE +P P
Sbjct: 608 YACSHSGLVDEGWRFFNIMRHECKIEPTVEHYACIVDMLARTGDLIKAYRFIENMPIPPD 667
Query: 376 VTVFETLKNYARIHGDVDLEDHVEELIVSLDPS---------------------KAVANK 414
T++ L RIH DV L + V E + L+P K + +
Sbjct: 668 ATIWGALLCGCRIHHDVKLAEKVAEKVFELEPENTGYYVLMANIYAEAEKWEQVKRLRKR 727
Query: 415 IPTPPPKKYTAISMLDGKNRII-----EYKNPTLYKDDEKLIAMNS-MKDAGYVPDTRYV 468
I +K S ++ K R+ + NP + L + + M + GY P T+Y
Sbjct: 728 IGQRGLRKNPGCSWIEIKGRVNIFVAGDSSNPETENIEAFLRKVRARMIEEGYSPLTKYA 787
Query: 469 LHDIDQEAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLRVCGDCHNAIKIMSRIVGR 528
L D ++ KE+AL HSE+LA+A G+IS+ +R+ KNLRVCGDCH K MS++ R
Sbjct: 788 LIDAEEMEKEEALCGHSEKLAMALGIISSGHGKIIRVTKNLRVCGDCHEMAKFMSKLTRR 847
Query: 529 ELIVRDNKRFHHFKDGKCSCGDYW 552
E+++RD+ RFH FKDG CSC +W
Sbjct: 848 EIVLRDSNRFHQFKDGHCSCRGFW 871
Score = 80.9 bits (198), Expect = 2e-15
Identities = 54/214 (25%), Positives = 106/214 (49%), Gaps = 6/214 (2%)
Query: 197 ELMEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNCK 256
++M G++ D+ F + +SV +++H + L+S F + N ++ Y +
Sbjct: 185 KMMSSGVEMDSYTFSCVSKSFSSLRSVHGGEQLHGFILKSGFGERNSVGNSLVAFYLKNQ 244
Query: 257 SMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSETMLAVLS 316
+ AR+VFD M R++ SW+ +I GY ++ + ++GL +F QM G+EI T+++V +
Sbjct: 245 RVDSARKVFDEMTERDVISWNSIINGYVSNGLAEKGLSVFVQMLVSGIEIDLATIVSVFA 304
Query: 317 ACGSAEAVE-DAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLPFEPT 375
C + + ++ +K+ + E LLD+ + G L A+ ++
Sbjct: 305 GCADSRLISLGRAVHSIGVKACFSRED--RFCNTLLDMYSKCGDLDSAKAVFREMSDRSV 362
Query: 376 VTVFETLKNYAR--IHGD-VDLEDHVEELIVSLD 406
V+ + YAR + G+ V L + +EE +S D
Sbjct: 363 VSYTSMIAGYAREGLAGEAVKLFEEMEEEGISPD 396
Score = 78.2 bits (191), Expect = 1e-14
Identities = 47/193 (24%), Positives = 97/193 (49%), Gaps = 3/193 (1%)
Query: 196 LELMEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNC 255
++++ GI+ D +F C S+ + + VH +++ F + + N +++MY C
Sbjct: 285 VQMLVSGIEIDLATIVSVFAGCADSRLISLGRAVHSIGVKACFSREDRFCNTLLDMYSKC 344
Query: 256 KSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSETMLAVL 315
+ A+ VF M +R++ S+ MI GYA + E ++LFE+M E G+ T+ AVL
Sbjct: 345 GDLDSAKAVFREMSDRSVVSYTSMIAGYAREGLAGEAVKLFEEMEEEGISPDVYTVTAVL 404
Query: 316 SACGSAEAVEDAYIYLESMK-SKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLPFEP 374
+ C +++ E +K + G + V + L+D+ + G ++EAE ++ +
Sbjct: 405 NCCARYRLLDEGKRVHEWIKENDLGFDIFVSN--ALMDMYAKCGSMQEAELVFSEMRVKD 462
Query: 375 TVTVFETLKNYAR 387
++ + Y++
Sbjct: 463 IISWNTIIGGYSK 475
Score = 76.6 bits (187), Expect = 3e-14
Identities = 54/211 (25%), Positives = 101/211 (47%), Gaps = 9/211 (4%)
Query: 186 FCQEGKVKEALELMEK----GIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSD 241
+ +EG EA++L E+ GI D + + C + + +++ K+VH++ ++ D
Sbjct: 372 YAREGLAGEAVKLFEEMEEEGISPDVYTVTAVLNCCARYRLLDEGKRVHEWIKENDLGFD 431
Query: 242 FKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFE-QMN 300
+ N +++MY C SM +A VF M +++ SW+ +I GY+ + +E L LF +
Sbjct: 432 IFVSNALMDMYAKCGSMQEAELVFSEMRVKDIISWNTIIGGYSKNCYANEALSLFNLLLE 491
Query: 301 ELGLEITSETMLAVLSACGSAEAVEDA-YIYLESMKSKYGIEPGVEHYMGLLDVLGQSGY 359
E T+ VL AC S A + I+ M++ Y + V + L+D+ + G
Sbjct: 492 EKRFSPDERTVACVLPACASLSAFDKGREIHGYIMRNGYFSDRHVAN--SLVDMYAKCGA 549
Query: 360 LKEAEEFIEQLPFEPTVTVFETLKNYARIHG 390
L A + + + V+ + Y +HG
Sbjct: 550 LLLAHMLFDDIASKDLVSWTVMIAGYG-MHG 579
Score = 67.8 bits (164), Expect = 2e-11
Identities = 42/144 (29%), Positives = 74/144 (51%), Gaps = 2/144 (1%)
Query: 183 LTRFCQEGKVKEALELMEKGIKADANCFEI--LFDLCGKSKSVEDAKKVHDYFLQSTFRS 240
L RFC+ G ++ A++L+ K D + + + LC SKS++D K+V ++ + F
Sbjct: 68 LRRFCESGNLENAVKLLCVSGKWDIDPRTLCSVLQLCADSKSLKDGKEVDNFIRGNGFVI 127
Query: 241 DFKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMN 300
D + +K+ MY NC + +A RVFD + W++++ A S + LF++M
Sbjct: 128 DSNLGSKLSLMYTNCGDLKEASRVFDEVKIEKALFWNILMNELAKSGDFSGSIGLFKKMM 187
Query: 301 ELGLEITSETMLAVLSACGSAEAV 324
G+E+ S T V + S +V
Sbjct: 188 SSGVEMDSYTFSCVSKSFSSLRSV 211
>At4g21070 putative protein (fragment)
Length = 1495
Score = 221 bits (564), Expect = 6e-58
Identities = 131/422 (31%), Positives = 219/422 (51%), Gaps = 35/422 (8%)
Query: 151 QNGNLNQFQNPNNQFQTPNVQEQAPPPPSIV---DLTRFCQEGKVKEAL----ELMEKGI 203
QN L+ + N + V ++ P + + F + GK +EAL E+ KGI
Sbjct: 159 QNSLLHLYANCGDVASAYKVFDKMPEKDLVAWNSVINGFAENGKPEEALALYTEMNSKGI 218
Query: 204 KADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNCKSMTDARR 263
K D L C K ++ K+VH Y ++ + N ++++Y C + +A+
Sbjct: 219 KPDGFTIVSLLSACAKIGALTLGKRVHVYMIKVGLTRNLHSSNVLLDLYARCGRVEEAKT 278
Query: 264 VFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNEL-GLEITSETMLAVLSACGSAE 322
+FD M ++N SW +I G A + G E ++LF+ M GL T + +L AC
Sbjct: 279 LFDEMVDKNSVSWTSLIVGLAVNGFGKEAIELFKYMESTEGLLPCEITFVGILYACSHCG 338
Query: 323 AVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLPFEPTVTVFETL 382
V++ + Y M+ +Y IEP +EH+ ++D+L ++G +K+A E+I+ +P +P V ++ TL
Sbjct: 339 MVKEGFEYFRRMREEYKIEPRIEHFGCMVDLLARAGQVKKAYEYIKSMPMQPNVVIWRTL 398
Query: 383 KNYARIHGDVDLEDHVEELIVSLDPSKA---------------------VANKIPTPPPK 421
+HGD DL + I+ L+P+ + + ++ K
Sbjct: 399 LGACTVHGDSDLAEFARIQILQLEPNHSGDYVLLSNMYASEQRWSDVQKIRKQMLRDGVK 458
Query: 422 KYTAISMLDGKNRIIEY-----KNPTLYKDDEKLIAMNS-MKDAGYVPDTRYVLHDIDQE 475
K S+++ NR+ E+ +P KL M ++ GYVP V D+++E
Sbjct: 459 KVPGHSLVEVGNRVHEFLMGDKSHPQSDAIYAKLKEMTGRLRSEGYVPQISNVYVDVEEE 518
Query: 476 AKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLRVCGDCHNAIKIMSRIVGRELIVRDN 535
KE A++YHSE++AIA+ LISTP R+P+ ++KNLRVC DCH AIK++S++ RE++VRD
Sbjct: 519 EKENAVVYHSEKIAIAFMLISTPERSPITVVKNLRVCADCHLAIKLVSKVYNREIVVRDR 578
Query: 536 KR 537
R
Sbjct: 579 SR 580
Score = 79.7 bits (195), Expect = 4e-15
Identities = 42/171 (24%), Positives = 88/171 (50%), Gaps = 7/171 (4%)
Query: 203 IKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNCKSMTDAR 262
++ D + + L V + +H ++S F S + N ++ +Y NC + A
Sbjct: 117 VEPDTHTYPFLIKAVTTMADVRLGETIHSVVIRSGFGSLIYVQNSLLHLYANCGDVASAY 176
Query: 263 RVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSETMLAVLSACGSAE 322
+VFD MP +++ +W+ +I G+A + +E L L+ +MN G++ T++++LSAC
Sbjct: 177 KVFDKMPEKDLVAWNSVINGFAENGKPEEALALYTEMNSKGIKPDGFTIVSLLSACAKIG 236
Query: 323 AV---EDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQL 370
A+ + ++Y+ K G+ + LLD+ + G ++EA+ +++
Sbjct: 237 ALTLGKRVHVYM----IKVGLTRNLHSSNVLLDLYARCGRVEEAKTLFDEM 283
>At1g74630 hypothetical protein
Length = 643
Score = 218 bits (556), Expect = 5e-57
Identities = 126/385 (32%), Positives = 203/385 (52%), Gaps = 29/385 (7%)
Query: 197 ELMEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNCK 256
EL G+ + + C +S S E K +H + ++ + ++N +I+MY C
Sbjct: 259 ELQRAGMSPNEVSLTGVLSACSQSGSFEFGKILHGFVEKAGYSWIVSVNNALIDMYSRCG 318
Query: 257 SMTDARRVFDHMPNRN-MDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSETMLAVL 315
++ AR VF+ M + + SW MI G A G+E ++LF +M G+ + +++L
Sbjct: 319 NVPMARLVFEGMQEKRCIVSWTSMIAGLAMHGQGEEAVRLFNEMTAYGVTPDGISFISLL 378
Query: 316 SACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLPFEPT 375
AC A +E+ Y MK Y IEP +EHY ++D+ G+SG L++A +FI Q+P PT
Sbjct: 379 HACSHAGLIEEGEDYFSEMKRVYHIEPEIEHYGCMVDLYGRSGKLQKAYDFICQMPIPPT 438
Query: 376 VTVFETLKNYARIHGDVDLEDHVEELIVSLDPSKA-----VANKIPTPPP---------- 420
V+ TL HG+++L + V++ + LDP+ + ++N T
Sbjct: 439 AIVWRTLLGACSSHGNIELAEQVKQRLNELDPNNSGDLVLLSNAYATAGKWKDVASIRKS 498
Query: 421 ------KKYTAISMLDGKNRIIEY-----KNPTLYKDDEKL--IAMNSMKDAGYVPDTRY 467
KK TA S+++ + ++ K + EKL I + +AGY P+
Sbjct: 499 MIVQRIKKTTAWSLVEVGKTMYKFTAGEKKKGIDIEAHEKLKEIILRLKDEAGYTPEVAS 558
Query: 468 VLHDIDQEAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLRVCGDCHNAIKIMSRIVG 527
L+D+++E KE + HSE+LA+A+ L +RI+KNLR+C DCH +K+ S++ G
Sbjct: 559 ALYDVEEEEKEDQVSKHSEKLALAFALARLSKGANIRIVKNLRICRDCHAVMKLTSKVYG 618
Query: 528 RELIVRDNKRFHHFKDGKCSCGDYW 552
E++VRD RFH FKDG CSC DYW
Sbjct: 619 VEILVRDRNRFHSFKDGSCSCRDYW 643
Score = 53.5 bits (127), Expect = 3e-07
Identities = 44/201 (21%), Positives = 85/201 (41%), Gaps = 5/201 (2%)
Query: 190 GKVKEALELMEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVI 249
G V+ A ++ ++ + + + + C + V A+++ D L S N ++
Sbjct: 155 GCVEFARKVFDEMHQPNLVAWNAVITACFRGNDVAGAREIFDKMLVRNHTS----WNVML 210
Query: 250 EMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSE 309
Y + A+R+F MP+R+ SW MI G A++ +E F ++ G+
Sbjct: 211 AGYIKAGELESAKRIFSEMPHRDDVSWSTMIVGIAHNGSFNESFLYFRELQRAGMSPNEV 270
Query: 310 TMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQ 369
++ VLSAC + + E I L K G V L+D+ + G + A E
Sbjct: 271 SLTGVLSACSQSGSFEFGKI-LHGFVEKAGYSWIVSVNNALIDMYSRCGNVPMARLVFEG 329
Query: 370 LPFEPTVTVFETLKNYARIHG 390
+ + + + ++ +HG
Sbjct: 330 MQEKRCIVSWTSMIAGLAMHG 350
Score = 50.1 bits (118), Expect = 3e-06
Identities = 50/222 (22%), Positives = 92/222 (40%), Gaps = 40/222 (18%)
Query: 196 LELMEKG-IKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGN 254
+E+M KG + D+ F + +S+ ++H L+ S + +I MYG
Sbjct: 94 VEMMRKGFVFPDSFSFAFVIKAVENFRSLRTGFQMHCQALKHGLESHLFVGTTLIGMYGG 153
Query: 255 CKSMTDARRVFDHM--PN-----------------------------RNMDSWHMMIRGY 283
C + AR+VFD M PN RN SW++M+ GY
Sbjct: 154 CGCVEFARKVFDEMHQPNLVAWNAVITACFRGNDVAGAREIFDKMLVRNHTSWNVMLAGY 213
Query: 284 ANSTMGDEGLQLFEQMNELGLEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPG 343
+ + ++F +M +++ TM+ ++ GS ++++Y ++ + G+ P
Sbjct: 214 IKAGELESAKRIFSEMPHRD-DVSWSTMIVGIAHNGS---FNESFLYFRELQ-RAGMSPN 268
Query: 344 VEHYMGLLDVLGQSG---YLKEAEEFIEQLPFEPTVTVFETL 382
G+L QSG + K F+E+ + V+V L
Sbjct: 269 EVSLTGVLSACSQSGSFEFGKILHGFVEKAGYSWIVSVNNAL 310
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.317 0.136 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,906,792
Number of Sequences: 26719
Number of extensions: 792935
Number of successful extensions: 16914
Number of sequences better than 10.0: 874
Number of HSP's better than 10.0 without gapping: 545
Number of HSP's successfully gapped in prelim test: 351
Number of HSP's that attempted gapping in prelim test: 3947
Number of HSP's gapped (non-prelim): 3999
length of query: 552
length of database: 11,318,596
effective HSP length: 104
effective length of query: 448
effective length of database: 8,539,820
effective search space: 3825839360
effective search space used: 3825839360
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 63 (28.9 bits)
Medicago: description of AC148348.6


