

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148347.8 - phase: 0 /pseudo
(97 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
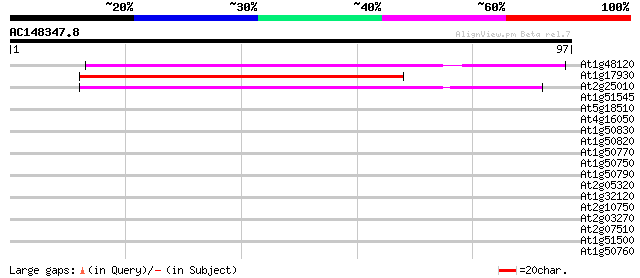
Score E
Sequences producing significant alignments: (bits) Value
At1g48120 serine/threonine phosphatase PP7, putative 52 7e-08
At1g17930 unknown protein 52 7e-08
At2g25010 unknown protein 50 2e-07
At1g51545 unknown protein 34 0.011
At5g18510 putative protein 33 0.032
At4g16050 hypothetical protein 31 0.092
At1g50830 hypothetical protein 30 0.16
At1g50820 hypothetical protein 30 0.16
At1g50770 hypothetical protein 30 0.27
At1g50750 hypothetical protein 29 0.46
At1g50790 hypothetical protein 27 1.3
At2g05320 putative N-acetylglucosaminyltransferase 27 2.3
At1g32120 hypothetical protein 27 2.3
At2g10750 pseudogene 25 5.0
At2g03270 helicase like protein 25 5.0
At2g07510 hypothetical protein 25 6.6
At1g51500 ATP-dependent transmembrane transporter, putative 25 8.6
At1g50760 hypothetical protein 25 8.6
>At1g48120 serine/threonine phosphatase PP7, putative
Length = 1338
Score = 51.6 bits (122), Expect = 7e-08
Identities = 28/83 (33%), Positives = 46/83 (54%), Gaps = 3/83 (3%)
Query: 14 GLQWLQHVSYLVIDQGLLSVIAERWHEVTSSFHLPIWEMIVTLDDLSCLLHLSIESHMLY 73
GL + V+++ +D L++ + ERW T +FHLP E+ VTL D++ LL L ++ +
Sbjct: 68 GLYGVYKVAFIQLDYALITALVERWRPETHTFHLPAGEITVTLQDVNILLGLRVDGPAVT 127
Query: 74 HNIYLTRSEAVDLMVELLGSDSG 96
+ T+ DL +LLG G
Sbjct: 128 GS---TKYNWADLCEDLLGHRPG 147
>At1g17930 unknown protein
Length = 478
Score = 51.6 bits (122), Expect = 7e-08
Identities = 20/56 (35%), Positives = 36/56 (63%)
Query: 13 SGLQWLQHVSYLVIDQGLLSVIAERWHEVTSSFHLPIWEMIVTLDDLSCLLHLSIE 68
+G W + V + ++ L+S + ERW T++FH P EM +TLD++S +L L+++
Sbjct: 49 AGFGWFRLVGSISLNNSLISALVERWRRETNTFHFPCGEMTITLDEVSLILGLAVD 104
>At2g25010 unknown protein
Length = 509
Score = 49.7 bits (117), Expect = 2e-07
Identities = 24/80 (30%), Positives = 47/80 (58%), Gaps = 1/80 (1%)
Query: 13 SGLQWLQHVSYLVIDQGLLSVIAERWHEVTSSFHLPIWEMIVTLDDLSCLLHLSIESHML 72
+G + + + + ++ L+S + ERW T++FHLP+ EM +TLD+++ +L L I+ +
Sbjct: 58 AGFGYFRKIGPMSLNNSLISALVERWRRETNTFHLPLGEMTITLDEVALVLGLEIDGDPI 117
Query: 73 YHNIYLTRSEAVDLMVELLG 92
+ + A+D+ LLG
Sbjct: 118 VGS-KVGDEVAMDMCGRLLG 136
>At1g51545 unknown protein
Length = 629
Score = 34.3 bits (77), Expect = 0.011
Identities = 16/47 (34%), Positives = 26/47 (55%)
Query: 27 DQGLLSVIAERWHEVTSSFHLPIWEMIVTLDDLSCLLHLSIESHMLY 73
+Q LL + E+W T SF P E +TL+D+ LL S++ ++
Sbjct: 28 NQSLLLALVEKWCPETKSFLFPWGEATITLEDVLVLLGFSVQGSPVF 74
>At5g18510 putative protein
Length = 702
Score = 32.7 bits (73), Expect = 0.032
Identities = 16/44 (36%), Positives = 24/44 (54%)
Query: 24 LVIDQGLLSVIAERWHEVTSSFHLPIWEMIVTLDDLSCLLHLSI 67
++ D + IAE+W T SF P E +TL+D+ LL S+
Sbjct: 91 IIKDTSSILSIAEKWCSETKSFIFPWGEATITLEDVMVLLGFSV 134
>At4g16050 hypothetical protein
Length = 900
Score = 31.2 bits (69), Expect = 0.092
Identities = 17/57 (29%), Positives = 32/57 (55%), Gaps = 1/57 (1%)
Query: 30 LLSVIAERWHEVTSSFHLPIWEMIVTLDDLSCLLHLSIESHMLYHNIYLTR-SEAVD 85
L+ +A+ W T++F P E +TL+D++ LL SI ++ ++ + EAV+
Sbjct: 363 LILSLAQNWCPETNTFVFPWGEATITLEDVNVLLGFSISGSSVFASLQSSEMKEAVE 419
>At1g50830 hypothetical protein
Length = 768
Score = 30.4 bits (67), Expect = 0.16
Identities = 14/38 (36%), Positives = 22/38 (57%)
Query: 30 LLSVIAERWHEVTSSFHLPIWEMIVTLDDLSCLLHLSI 67
L+ ++E+W T SF P E +TL+D+ LL S+
Sbjct: 119 LILSVSEKWCPETKSFVFPWGEATITLEDVMVLLGFSV 156
>At1g50820 hypothetical protein
Length = 528
Score = 30.4 bits (67), Expect = 0.16
Identities = 15/41 (36%), Positives = 23/41 (55%)
Query: 27 DQGLLSVIAERWHEVTSSFHLPIWEMIVTLDDLSCLLHLSI 67
D L+ +AE+W T +F P E +TL+D+ LL S+
Sbjct: 94 DTDLVLGLAEKWCPDTKTFIFPWGEATITLEDVMVLLGFSV 134
>At1g50770 hypothetical protein
Length = 632
Score = 29.6 bits (65), Expect = 0.27
Identities = 14/34 (41%), Positives = 20/34 (58%)
Query: 34 IAERWHEVTSSFHLPIWEMIVTLDDLSCLLHLSI 67
IAE+W T +F P E +TL+D+ LL S+
Sbjct: 104 IAEKWCPDTKTFVFPWGETTITLEDVMLLLGFSV 137
>At1g50750 hypothetical protein
Length = 816
Score = 28.9 bits (63), Expect = 0.46
Identities = 15/38 (39%), Positives = 21/38 (54%)
Query: 30 LLSVIAERWHEVTSSFHLPIWEMIVTLDDLSCLLHLSI 67
L+ IAE+W T +F P E VTL+D+ L S+
Sbjct: 76 LILGIAEKWCPYTKTFVFPWGETAVTLEDVMVLSGFSV 113
>At1g50790 hypothetical protein
Length = 812
Score = 27.3 bits (59), Expect = 1.3
Identities = 14/38 (36%), Positives = 22/38 (57%)
Query: 30 LLSVIAERWHEVTSSFHLPIWEMIVTLDDLSCLLHLSI 67
L+ IAE+W T++F E +TL+D+ LL S+
Sbjct: 101 LVMGIAEKWCPDTNTFVFSWGEATITLEDVMVLLGFSV 138
>At2g05320 putative N-acetylglucosaminyltransferase
Length = 419
Score = 26.6 bits (57), Expect = 2.3
Identities = 12/26 (46%), Positives = 17/26 (65%)
Query: 28 QGLLSVIAERWHEVTSSFHLPIWEMI 53
+GL S++AER V SF+ +WE I
Sbjct: 266 EGLESLVAERMGNVGYSFNRSVWENI 291
>At1g32120 hypothetical protein
Length = 1206
Score = 26.6 bits (57), Expect = 2.3
Identities = 13/38 (34%), Positives = 22/38 (57%)
Query: 30 LLSVIAERWHEVTSSFHLPIWEMIVTLDDLSCLLHLSI 67
L+ + E+W T++F P E +TL+D+ L LS+
Sbjct: 121 LIVALVEKWCIETNTFVFPWGEATLTLEDMIVLGGLSV 158
>At2g10750 pseudogene
Length = 488
Score = 25.4 bits (54), Expect = 5.0
Identities = 11/32 (34%), Positives = 16/32 (49%)
Query: 58 DLSCLLHLSIESHMLYHNIYLTRSEAVDLMVE 89
DL C+ L E ++Y LT E ++VE
Sbjct: 272 DLGCVYSLGREEFVIYETTILTTQEVSKILVE 303
>At2g03270 helicase like protein
Length = 639
Score = 25.4 bits (54), Expect = 5.0
Identities = 17/45 (37%), Positives = 23/45 (50%), Gaps = 3/45 (6%)
Query: 56 LDDLSCLLHLSIESHMLYHNIYLTRS---EAVDLMVELLGSDSGE 97
L D H S+ SHML+ +T+S EA L+V+ G D E
Sbjct: 450 LYDNKITAHSSVASHMLFDLENVTKSSSTEATLLLVDTAGCDMEE 494
>At2g07510 hypothetical protein
Length = 460
Score = 25.0 bits (53), Expect = 6.6
Identities = 15/54 (27%), Positives = 27/54 (49%), Gaps = 5/54 (9%)
Query: 34 IAERWHEVTSSFHLPIWEMIVTLDDLSCLLHLSIESHMLYHNIYLTRSEAVDLM 87
I ++WH++ S + +++S LL L IE H HN + +A D++
Sbjct: 183 IDQKWHDIDLSLSDEEED-----EEVSHLLSLIIEEHPYEHNSWSGGMDACDIV 231
>At1g51500 ATP-dependent transmembrane transporter, putative
Length = 687
Score = 24.6 bits (52), Expect = 8.6
Identities = 14/45 (31%), Positives = 19/45 (42%)
Query: 18 LQHVSYLVIDQGLLSVIAERWHEVTSSFHLPIWEMIVTLDDLSCL 62
L S + Q L ++ + V SS H P E+ DDL L
Sbjct: 198 LDSASAFFVIQALRNIARDGGRTVVSSIHQPSSEVFALFDDLFLL 242
>At1g50760 hypothetical protein
Length = 649
Score = 24.6 bits (52), Expect = 8.6
Identities = 11/34 (32%), Positives = 18/34 (52%)
Query: 34 IAERWHEVTSSFHLPIWEMIVTLDDLSCLLHLSI 67
+AE+W T +F E +TL+D+ L S+
Sbjct: 94 VAEKWSPDTKTFVFSWGEATITLEDVMVLSGFSV 127
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.323 0.136 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,196,936
Number of Sequences: 26719
Number of extensions: 75799
Number of successful extensions: 176
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 16
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 159
Number of HSP's gapped (non-prelim): 19
length of query: 97
length of database: 11,318,596
effective HSP length: 73
effective length of query: 24
effective length of database: 9,368,109
effective search space: 224834616
effective search space used: 224834616
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 52 (24.6 bits)
Medicago: description of AC148347.8


