

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148345.17 + phase: 0
(528 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
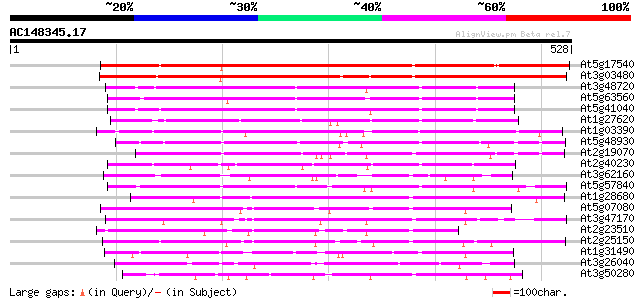
Score E
Sequences producing significant alignments: (bits) Value
At5g17540 unknown protein 412 e-115
At3g03480 putative hypersensitivity-related gene 400 e-112
At3g48720 unknown protein 229 3e-60
At5g63560 acyltransferase-like protein 215 6e-56
At5g41040 N-hydroxycinnamoyl/benzoyltransferase-like protein 214 9e-56
At1g27620 putative hypersensitivity-related protein 193 2e-49
At1g03390 hypothetical protein 176 2e-44
At5g48930 anthranilate N-benzoyltransferase 160 2e-39
At2g19070 putative anthranilate N-hydroxycinnamoyl/benzoyltransf... 156 2e-38
At2g40230 putative anthranilate N-hydroxycinnamoyl/benzoyltransf... 150 1e-36
At3g62160 unknown protein 146 3e-35
At5g57840 N-hydroxycinnamoyl/benzoyltransferase 143 3e-34
At1g28680 anthranilate N-hydroxycinnamoyl/benzoyltransferase lik... 136 3e-32
At5g07080 unknown protein 125 8e-29
At3g47170 hypersensitivity-related protein-like protein 120 2e-27
At2g23510 putative anthranilate N-hydroxycinnamoyl/benzoyltransf... 111 9e-25
At2g25150 unknown protein 110 3e-24
At1g31490 putative protein 99 8e-21
At3g26040 hypothetical protein 95 8e-20
At3g50280 anthranilate N-hydroxycinnamoyl/benzoyltransferase - l... 94 1e-19
>At5g17540 unknown protein
Length = 461
Score = 412 bits (1059), Expect = e-115
Identities = 223/447 (49%), Positives = 297/447 (65%), Gaps = 10/447 (2%)
Query: 86 SAPLLFTVRRSQPELVPPAAPTPREVKLLSDIDDQEGLRFNIPMMFIYRHEPSMKEKDPV 145
S L F + R +PELV PA PTPRE+K LSDIDDQEGLRF+IP +F YRH P+ DPV
Sbjct: 2 SGSLTFKIYRQKPELVSPAKPTPRELKPLSDIDDQEGLRFHIPTIFFYRHNPTTNS-DPV 60
Query: 146 KVLRHALSQALVYYYPFAGRIREGAGRKLMVDCTGEGVMFVEAEADVTLDEFG--DALHP 203
V+R AL++ LVYYYPFAGR+REG RKL VDCTGEGV+F+EA+ADVTL EF DAL P
Sbjct: 61 AVIRRALAETLVYYYPFAGRLREGPNRKLAVDCTGEGVLFIEADADVTLVEFEEKDALKP 120
Query: 204 PLPCFEELLYDVPGSELIIDRPIRLIQVTRLKCGSFILVVYLNHTMSDGAGLKLFMNAWA 263
P PCFEELL++V GS +++ P+ L+QVTRLKCG FI V +NH MSD GL LF+
Sbjct: 121 PFPCFEELLFNVEGSCEMLNTPLMLMQVTRLKCGGFIFAVRINHAMSDAGGLTLFLKTMC 180
Query: 264 EMARGAHKPSIQPVWNREILMARDPPHITCNHHEYEQIFSSNTIKEEDTTTLVHQSFFFR 323
E RG H P++ PVW R +L AR +T H EY+++ + T LV +S FF
Sbjct: 181 EFVRGYHAPTVAPVWERHLLSARVLLRVTHAHREYDEMPAIGTELGSRRDNLVGRSLFFG 240
Query: 324 TSDIVVLRLLVPFHL-RHCTTFDLITSCFWCCRTKALQLEADDEIRMMCIVNARSRFNAN 382
++ +R L+P +L T +++TS W RT AL+ + D E+R++ IVNARS+
Sbjct: 241 PCEMSAIRRLLPPNLVNSSTNMEMLTSFLWRYRTIALRPDQDKEMRLILIVNARSKL--K 298
Query: 383 NSPLV-GYYGNCFAYPAAVTTAGKLCGNSLGYAVELVRKLKAEVTEEYMHSVADLMVIKE 441
N PL GYYGN FA+P A+ TA +L L +A+ L+++ K+ VTEEYM S+ADLMVIK
Sbjct: 299 NPPLPRGYYGNAFAFPVAIATANELTKKPLEFALRLIKEAKSSVTEEYMRSLADLMVIKG 358
Query: 442 RCLFTTIRSCVVSDVTRIKLREVDFG-WGEAVYGGAAKSGAGPFPGAAYIIPHKNVEGEE 500
R F++ + +VSDV RI ++DFG WG+ VYGG +G PGA++ + + GE
Sbjct: 359 RPSFSSDGAYLVSDV-RI-FADIDFGIWGKPVYGGIGTAGVEDLPGASFYVSFEKRNGEI 416
Query: 501 GFILLVCLSFEAMKRFAKELDEMLGKQ 527
G ++ VCL +AM+RF +EL+ + Q
Sbjct: 417 GIVVPVCLPEKAMQRFVEELEGVFNGQ 443
>At3g03480 putative hypersensitivity-related gene
Length = 454
Score = 400 bits (1029), Expect = e-112
Identities = 214/445 (48%), Positives = 291/445 (65%), Gaps = 11/445 (2%)
Query: 85 TSAPLLFTVRRSQPELVPPAAPTPREVKLLSDIDDQEGLRFNIPMMFIYRHEPSMKEKDP 144
T+ L F V R Q ELV PA PTPRE+K LSDIDDQ+GLRF IP++F YR S + DP
Sbjct: 11 TTTGLSFKVHRQQRELVTPAKPTPRELKPLSDIDDQQGLRFQIPVIFFYRPNLS-SDLDP 69
Query: 145 VKVLRHALSQALVYYYPFAGRIREGAGRKLMVDCTGEGVMFVEAEADVTLDEF--GDALH 202
V+V++ AL+ ALVYYYPFAGR+RE + RKL VDCTGEGV+F+EAEADV L E DAL
Sbjct: 70 VQVIKKALADALVYYYPFAGRLRELSNRKLAVDCTGEGVLFIEAEADVALAELEEADALL 129
Query: 203 PPLPCFEELLYDVPGSELIIDRPIRLIQVTRLKCGSFILVVYLNHTMSDGAGLKLFMNAW 262
PP P EELL+DV GS +++ P+ L+QVTRLKC FI + NHTM+DGAGL LF+ +
Sbjct: 130 PPFPFLEELLFDVEGSSDVLNTPLLLVQVTRLKCCGFIFALRFNHTMTDGAGLSLFLKSL 189
Query: 263 AEMARGAHKPSIQPVWNREIL-MARDPPHITCNHHEYEQIFSSNTIKEEDTTTLVHQSFF 321
E+A G H PS+ PVWNR +L ++ +T H EY+ + + LV +SFF
Sbjct: 190 CELACGLHAPSVPPVWNRHLLTVSASEARVTHTHREYDDQVGIDVVATGH--PLVSRSFF 247
Query: 322 FRTSDIVVLRLLVPFHLRHCTTFDLITSCFWCCRTKALQLEADDEIRMMCIVNARSRFNA 381
FR +I +R L+P L H T+F+ ++S W CRT AL + + E+R+ CI+N+RS+
Sbjct: 248 FRAEEISAIRKLLPPDL-HNTSFEALSSFLWRCRTIALNPDPNTEMRLTCIINSRSKL-- 304
Query: 382 NNSPL-VGYYGNCFAYPAAVTTAGKLCGNSLGYAVELVRKLKAEVTEEYMHSVADLMVIK 440
N PL GYYGN F PAA+ TA L L +A+ L+++ K+ VTE+Y+ SV LM +
Sbjct: 305 RNPPLEPGYYGNVFVIPAAIATARDLIEKPLEFALRLIQETKSSVTEDYVRSVTALMATR 364
Query: 441 ERCLFTTIRSCVVSDVTRIKLREVDFG-WGEAVYGGAAKSGAGPFPGAAYIIPHKNVEGE 499
R +F + ++SD+ L ++DFG WG+ VYGG AK+G FPG ++ +P KN +GE
Sbjct: 365 GRPMFVASGNYIISDLRHFDLGKIDFGPWGKPVYGGTAKAGIALFPGVSFYVPFKNKKGE 424
Query: 500 EGFILLVCLSFEAMKRFAKELDEML 524
G ++ + L AM+ F EL+ +L
Sbjct: 425 TGTVVAISLPVRAMETFVAELNGVL 449
>At3g48720 unknown protein
Length = 430
Score = 229 bits (584), Expect = 3e-60
Identities = 133/389 (34%), Positives = 221/389 (56%), Gaps = 11/389 (2%)
Query: 91 FTVRRSQPELVPPAAPTPREVKLLSDIDDQEGLRFNIPMMFIYRHEPSMKEKDPVKVLRH 150
F V R PEL+PP + TP LS++D + + + ++ Y+ E S ++ V++
Sbjct: 7 FIVTRKNPELIPPVSETPNGHYYLSNLD--QNIAIIVKTLYYYKSE-SRTNQESYNVIKK 63
Query: 151 ALSQALVYYYPFAGRIREGAGRKLMVDCTGEGVMFVEAEADVTLDEFGDALHPP-LPCFE 209
+LS+ LV+YYP AGR+ K+ V+CTGEGV+ VEAEA+ +D +A+ + E
Sbjct: 64 SLSEVLVHYYPVAGRLTISPEGKIAVNCTGEGVVVVEAEANCGIDTIKEAISENRMETLE 123
Query: 210 ELLYDVPGSELIIDRPIRLIQVTRLKCGSFILVVYLNHTMSDGAGLKLFMNAWAEMARGA 269
+L+YDVPG+ I++ P ++QVT KCG F+L + ++H M DG F+N+W EMA+G
Sbjct: 124 KLVYDVPGARNILEIPPVVVQVTNFKCGGFVLGLGMSHNMFDGVAAAEFLNSWCEMAKGL 183
Query: 270 HKPSIQPVWNREILMARDPPHITCNHHEYEQIFS-SNTIKEEDTTTLVHQSFFFRTSDIV 328
S+ P +R IL +R+PP I H+E+++I S+T K D L+++SF F +
Sbjct: 184 -PLSVPPFLDRTILRSRNPPKIEFPHNEFDEIEDISDTGKIYDEEKLIYKSFLFEPEKLE 242
Query: 329 VLRLLV--PFHLRHCTTFDLITSCFWCCRTKALQLEADDEIRMMCIVNARSRFNANNSPL 386
L+++ + +TF +T W R +AL+ + D ++++ + RSRF
Sbjct: 243 KLKIMAIEENNNNKVSTFQALTGFLWKSRCEALRFKPDQRVKLLFAADGRSRFIPRLPQ- 301
Query: 387 VGYYGNCFAYPAAVTTAGKLCGNSLGYAVELVRKLKAEVTEEYMHSVADLMVIKERCLFT 446
GY GN VT++G+L GN L ++V LV++L VT+ +M S D + R +
Sbjct: 302 -GYCGNGIVLTGLVTSSGELVGNPLSHSVGLVKRLVELVTDGFMRSAMDYFEV-NRTRPS 359
Query: 447 TIRSCVVSDVTRIKLREVDFGWGEAVYGG 475
+ +++ +++ L ++DFGWGE V+ G
Sbjct: 360 MNATLLITSWSKLTLHKLDFGWGEPVFSG 388
>At5g63560 acyltransferase-like protein
Length = 426
Score = 215 bits (547), Expect = 6e-56
Identities = 134/386 (34%), Positives = 205/386 (52%), Gaps = 13/386 (3%)
Query: 93 VRRSQPELVPPAAPTPREVKLLSDIDDQEGLRFNIPMMFIYRHEPSMKEKDPVKVLRHAL 152
V R +P LV PA+ TP+ + LS++D + I F Y S ++ +V++ +L
Sbjct: 9 VTRKEPVLVSPASETPKGLHYLSNLDQNIAI---IVKTFYYFKSNSRSNEESYEVIKKSL 65
Query: 153 SQALVYYYPFAGRIREGAGRKLMVDCTGEGVMFVEAEADVTLDEFGDALHP--PLPCFEE 210
S+ LV+YYP AGR+ K+ VDCTGEGV+ VEAEA+ +++ A+ E+
Sbjct: 66 SEVLVHYYPAAGRLTISPEGKIAVDCTGEGVVVVEAEANCGIEKIKKAISEIDQPETLEK 125
Query: 211 LLYDVPGSELIIDRPIRLIQVTRLKCGSFILVVYLNHTMSDGAGLKLFMNAWAEMARGAH 270
L+YDVPG+ I++ P ++QVT KCG F+L + +NH M DG F+N+WAE ARG
Sbjct: 126 LVYDVPGARNILEIPPVVVQVTNFKCGGFVLGLGMNHNMFDGIAAMEFLNSWAETARGL- 184
Query: 271 KPSIQPVWNREILMARDPPHITCNHHEYEQIFS-SNTIKEEDTTTLVHQSFFFRTSDIVV 329
S+ P +R +L R PP I H+E+E + S T K LV++SF F +
Sbjct: 185 PLSVPPFLDRTLLRPRTPPKIEFPHNEFEDLEDISGTGKLYSDEKLVYKSFLFGPEKLER 244
Query: 330 LRLLVPFHLRHCTTFDLITSCFWCCRTKALQLEADDEIRMMCIVNARSRFNANNSPLVGY 389
L+++ TTF +T W R +AL L+ D I+++ + RSRF GY
Sbjct: 245 LKIMAE---TRSTTFQTLTGFLWRARCQALGLKPDQRIKLLFAADGRSRFVPELPK--GY 299
Query: 390 YGNCFAYPAAVTTAGKLCGNSLGYAVELVRKLKAEVTEEYMHSVADLMVIKERCLFTTIR 449
GN + VTTAG++ N L ++V LV++ V + +M S D + R +
Sbjct: 300 SGNGIVFTYCVTTAGEVTLNPLSHSVCLVKRAVEMVNDGFMRSAIDYFEV-TRARPSLTA 358
Query: 450 SCVVSDVTRIKLREVDFGWGEAVYGG 475
+ +++ ++ DFGWGE V G
Sbjct: 359 TLLITSWAKLSFHTKDFGWGEPVVSG 384
>At5g41040 N-hydroxycinnamoyl/benzoyltransferase-like protein
Length = 457
Score = 214 bits (545), Expect = 9e-56
Identities = 132/389 (33%), Positives = 212/389 (53%), Gaps = 13/389 (3%)
Query: 93 VRRSQPELVPPAAPTPREVKLLSDIDDQEGLRFNIPMMFIYRHEPSMKEKDPVKVLRHAL 152
V + +P LV P + T + + LS++D + + + ++ ++ E E + V+V++ AL
Sbjct: 33 VHQKEPALVKPESETRKGLYFLSNLD--QNIAVIVRTIYCFKSEERGNE-EAVQVIKKAL 89
Query: 153 SQALVYYYPFAGRIREGAGRKLMVDCTGEGVMFVEAEADVTLDEFGDALHPPLPCFEELL 212
SQ LV+YYP AGR+ KL VDCT EGV+FVEAEA+ +DE GD P +L+
Sbjct: 90 SQVLVHYYPLAGRLTISPEGKLTVDCTEEGVVFVEAEANCKMDEIGDITKPDPETLGKLV 149
Query: 213 YDVPGSELIIDRPIRLIQVTRLKCGSFILVVYLNHTMSDGAGLKLFMNAWAEMARGAHKP 272
YDV ++ I++ P QVT+ KCG F+L + +NH M DG G F+N+W ++ARG
Sbjct: 150 YDVVDAKNILEIPPVTAQVTKFKCGGFVLGLCMNHCMFDGIGAMEFVNSWGQVARGL-PL 208
Query: 273 SIQPVWNREILMARDPPHITCNHHEYEQIFSSNTIKEEDT-TTLVHQSFFFRTSDIVVLR 331
+ P +R IL AR+PP I H E+E+I + I T +++SF F I L+
Sbjct: 209 TTPPFSDRTILNARNPPKIENLHQEFEEIEDKSNINSLYTKEPTLYRSFCFDPEKIKKLK 268
Query: 332 LLVPFHL-----RHCTTFDLITSCFWCCRTKALQLEADDEIRMMCIVNARSRFNANNSPL 386
L + CT+F+ +++ W RTK+L++ +D + +++ V+ R++F
Sbjct: 269 LQATENSESLLGNSCTSFEALSAFVWRARTKSLKMLSDQKTKLLFAVDGRAKFEPQLPK- 327
Query: 387 VGYYGNCFAYPAAVTTAGKLCGNSLGYAVELVRKLKAEVTEEYMHSVADLMVIKERCLFT 446
GY+GN ++ AG+L L +AV LVR+ VT+ YM S D + R +
Sbjct: 328 -GYFGNGIVLTNSICEAGELIEKPLSFAVGLVREAIKMVTDGYMRSAIDYFEV-TRARPS 385
Query: 447 TIRSCVVSDVTRIKLREVDFGWGEAVYGG 475
+ +++ +R+ DFGWGE + G
Sbjct: 386 LSSTLLITTWSRLGFHTTDFGWGEPILSG 414
>At1g27620 putative hypersensitivity-related protein
Length = 442
Score = 193 bits (491), Expect = 2e-49
Identities = 129/393 (32%), Positives = 209/393 (52%), Gaps = 22/393 (5%)
Query: 96 SQPELVPPAAPTPREVKLLSDIDDQEGLRFNIPMMFIYRHEPSMKEKDPVKVLRHALSQA 155
+QP L+ P +PTP LS++DD LRF+I +++++ K P+ L+ +LS+
Sbjct: 12 NQPTLITPLSPTPNHSLYLSNLDDHHFLRFSIKYLYLFQ-----KSISPL-TLKDSLSRV 65
Query: 156 LVYYYPFAGRIR-EGAGRKLMVDCTGEGVMFVEAEADVTLDEFGDALHPPLPCFEELLYD 214
LV YYPFAGRIR G KL VDC GEG +F EA D+T +F P + +LL+
Sbjct: 66 LVDYYPFAGRIRVSDEGSKLEVDCNGEGAVFAEAFMDITCQDFVQLSPKPNKSWRKLLFK 125
Query: 215 VPGSELIIDRPIRLIQVTRLKCGSFILVVYLNHTMSDGAGLKLFMNAWAEMARG-AHKPS 273
V ++ +D P +IQVT L+CG IL +NH + DG G F++AWA AH P+
Sbjct: 126 VQ-AQSFLDIPPLVIQVTYLRCGGMILCTAINHCLCDGIGTSQFLHAWAHATTSQAHLPT 184
Query: 274 IQPVWNREILMARDPPHITCNHHEYEQ---IFSSNTI---KEEDTTTLVHQSFFFRTSDI 327
+P +R +L R+PP +T +H + + + S+T K + L + F S +
Sbjct: 185 -RPFHSRHVLDPRNPPRVTHSHPGFTRTTTVDKSSTFDISKYLQSQPLAPATLTFNQSHL 243
Query: 328 VVLRLLVPFHLRHCTTFDLITSCFWCCRTKALQLEADDEIRMMCIVNARSRFNANNSPLV 387
+ L+ L+ CTTF+ + + W ++L L ++++ VN R R
Sbjct: 244 LRLKKTCAPSLK-CTTFEALAANTWRSWAQSLDLPMTMLVKLLFSVNMRKRLTPELPQ-- 300
Query: 388 GYYGNCFAYPAAVTTAGKLCGNSLGYAVELVRKLKAEVTEEYMHSVADLMVIKERCLFTT 447
GYYGN F A + L ++ +AV+ +++ K+ +T+EY+ S DL ++++ + T
Sbjct: 301 GYYGNGFVLACAESKVQDLVNGNIYHAVKSIQEAKSRITDEYVRSTIDL--LEDKTVKTD 358
Query: 448 IR-SCVVSDVTRIKLREVDFGWGEAVYGGAAKS 479
+ S V+S ++ L E+D G G+ +Y G S
Sbjct: 359 VSCSLVISQWAKLGLEELDLGGGKPMYMGPLTS 391
>At1g03390 hypothetical protein
Length = 461
Score = 176 bits (447), Expect = 2e-44
Identities = 141/456 (30%), Positives = 213/456 (45%), Gaps = 31/456 (6%)
Query: 82 HIMTSAPLLFTVRRSQPELVPPAAPTPREVKLLSDIDDQEGLRFNIPMMFIYRHEPSMKE 141
HI + F V S P PA +P LS++DD G R P ++ Y + +E
Sbjct: 14 HIPVTINQQFLVHPSSPT---PANQSPHHSLYLSNLDDIIGARVFTPSVYFYP-STNNRE 69
Query: 142 KDPVKVLRHALSQALVYYYPFAGRIREGAGRKLMVDCTGE-GVMFVEAEADVTLDEFGDA 200
+K L+ ALS+ LV YYP +GR+RE KL V E GV+ V A + + L + GD
Sbjct: 70 SFVLKRLQDALSEVLVPYYPLSGRLREVENGKLEVFFGEEQGVLMVSANSSMDLADLGD- 128
Query: 201 LHPPLPCFEELLYDVPGSEL--IIDRPIRLIQVTRLKCGSFILVVYLNHTMSDGAGLKLF 258
L P P + L++ PG E I++ P+ + QVT CG F L + L H + DG G F
Sbjct: 129 LTVPNPAWLPLIFRNPGEEAYKILEMPLLIAQVTFFTCGGFSLGIRLCHCICDGFGAMQF 188
Query: 259 MNAWAEMAR-GAHKPSIQPVWNREILMARDPPHITCNHHEYEQIFSSNTIKEE--DTTTL 315
+ +WA A+ G +PVW+RE R+PP + HHEY I + + DT L
Sbjct: 189 LGSWAATAKTGKLIADPEPVWDRETFKPRNPPMVKYPHHEYLPIEERSNLTNSLWDTKPL 248
Query: 316 -----VHQSFFFRTSDIVVLR--LLVPFHLRHCTTFDLITSCFWCCRTKALQLE-ADDEI 367
+ + F R I LV C+TFD + + W KAL ++ D +
Sbjct: 249 QKCYRISKEFQCRVKSIAQGEDPTLV------CSTFDAMAAHIWRSWVKALDVKPLDYNL 302
Query: 368 RMMCIVNARSRFNANNSPLVGYYGNCFAYPAAVTTAGKLCGNSLGYAVELVRKLKAEVTE 427
R+ VN R+R G+YGN A+++ L +SL LV+ + V+E
Sbjct: 303 RLTFSVNVRTRLETLKL-RKGFYGNVVCLACAMSSVESLINDSLSKTTRLVQDARLRVSE 361
Query: 428 EYMHSVADLMVIKERCLFTTIRSCVVSDVTRIKLRE-VDFGWGEAVYGGAAKSGAGPFPG 486
+Y+ S+ D + +K ++ TR ++ E DFGWG+ VY G P P
Sbjct: 362 DYLRSMVDYVDVKRPKRLEFGGKLTITQWTRFEMYETADFGWGKPVYAGPI--DLRPTPQ 419
Query: 487 AAYIIPHKNVE--GEEGFILLVCLSFEAMKRFAKEL 520
++P VE ++ ++ +CL A+ F + L
Sbjct: 420 VCVLLPQGGVESGNDQSMVVCLCLPPTAVHTFTRLL 455
>At5g48930 anthranilate N-benzoyltransferase
Length = 433
Score = 160 bits (405), Expect = 2e-39
Identities = 127/441 (28%), Positives = 205/441 (45%), Gaps = 34/441 (7%)
Query: 100 LVPPAAPTPREVKLLSDIDDQEGLRFNIPMMFIYRHEPSMKEKDPVKVLRHALSQALVYY 159
+V PA TP S++D RF+ P ++ YR + DP +V++ ALS+ALV +
Sbjct: 10 MVRPATETPITNLWNSNVDLVIP-RFHTPSVYFYRPTGASNFFDP-QVMKEALSKALVPF 67
Query: 160 YPFAGRIREGAGRKLMVDCTGEGVMFVEAEADVTLDEFGDALHPPLPCFEELLYDVPGSE 219
YP AGR++ ++ +DC G GV+FV A+ +D+FGD P +L+ +V S
Sbjct: 68 YPMAGRLKRDDDGRIEIDCNGAGVLFVVADTPSVIDDFGD--FAPTLNLRQLIPEVDHSA 125
Query: 220 LIIDRPIRLIQVTRLKCGSFILVVYLNHTMSDGAGLKLFMNAWAEMARGAHKPSIQPVWN 279
I P+ ++QVT KCG L V + H +DG F+N W++MARG +I P +
Sbjct: 126 GIHSFPLLVLQVTFFKCGGASLGVGMQHHAADGFSGLHFINTWSDMARGLDL-TIPPFID 184
Query: 280 REILMARDPPHITCNHHEYEQIFSSNTIKE------EDTTTLVHQSFFFRTSDIVV---L 330
R +L ARDPP +H EY+ S + E+TT S F T D +V
Sbjct: 185 RTLLRARDPPQPAFHHVEYQPAPSMKIPLDPSKSGPENTTV----SIFKLTRDQLVALKA 240
Query: 331 RLLVPFHLRHCTTFDLITSCFWCCRTKALQLEADDEIRMMCIVNARSRFNANNSPLVGYY 390
+ + ++++++ W KA L D E ++ + RSR P GY+
Sbjct: 241 KSKEDGNTVSYSSYEMLAGHVWRSVGKARGLPNDQETKLYIATDGRSRLRPQLPP--GYF 298
Query: 391 GNCFAYPAAVTTAGKLCGNSLGYAVELVRKLKAEVTEEYMHSVADLMVIKERCLFTTIR- 449
GN + AG L YA + + + Y+ S D + ++ L +R
Sbjct: 299 GNVIFTATPLAVAGDLLSKPTWYAAGQIHDFLVRMDDNYLRSALDYLEMQPD-LSALVRG 357
Query: 450 -------SCVVSDVTRIKLREVDFGWGEAVYGGAAKSGAGPFPGAAYIIPHKNVEGEEGF 502
+ ++ R+ + + DFGWG ++ G G P+ G ++++P +G
Sbjct: 358 AHTYKCPNLGITSWVRLPIYDADFGWGRPIFMG---PGGIPYEGLSFVLPSPTNDG--SL 412
Query: 503 ILLVCLSFEAMKRFAKELDEM 523
+ + L E MK F K L E+
Sbjct: 413 SVAIALQSEHMKLFEKFLFEI 433
>At2g19070 putative anthranilate
N-hydroxycinnamoyl/benzoyltransferase
Length = 451
Score = 156 bits (395), Expect = 2e-38
Identities = 130/434 (29%), Positives = 203/434 (45%), Gaps = 43/434 (9%)
Query: 119 DQEGLRFNIPMMFIYRHEPSMKEKDPVKVLRHALSQALVYYYPFAGRIREGAGRKLMVDC 178
DQ G +IP ++ Y + + V++L+ +LS+ LV++YP AGR+R + ++C
Sbjct: 29 DQVGTITHIPTLYFYDKPSESFQGNVVEILKTSLSRVLVHFYPMAGRLRWLPRGRFELNC 88
Query: 179 TGEGVMFVEAEADVTLDEFGDALHPPLPCFEELLYDVPGSELIIDRPIRLIQVTRLKCGS 238
EGV F+EAE++ L +F D P P FE L+ V I P+ L QVT+ KCG
Sbjct: 89 NAEGVEFIEAESEGKLSDFKD--FSPTPEFENLMPQVNYKNPIETIPLFLAQVTKFKCGG 146
Query: 239 FILVVYLNHTMSDGAGLKLFMNAWAEMARGAHKPSIQPVWNREILMARD--PPHIT---C 293
L V ++H + DG ++ W +ARG ++ P +R+IL A + PP ++
Sbjct: 147 ISLSVNVSHAIVDGQSALHLISEWGRLARGEPLETV-PFLDRKILWAGEPLPPFVSPPKF 205
Query: 294 NHHEYEQ----IFSSNTIKEEDTTTLVHQSFFFRTSDIVVLRLLV-----PFHLRHCTTF 344
+H E++Q I ++ ++E T+V TS + LR + T +
Sbjct: 206 DHKEFDQPPFLIGETDNVEERKKKTIV-VMLPLSTSQLQKLRSKANGSKHSDPAKGFTRY 264
Query: 345 DLITSCFWCCRTKALQLEADDEIRMMCIVNARSRFNANNSPLV-GYYGNCFAYPAAVTTA 403
+ +T W C KA + + ++ RSR PL GY+GN A +T+
Sbjct: 265 ETVTGHVWRCACKARGHSPEQPTALGICIDTRSRM---EPPLPRGYFGNATLDVVAASTS 321
Query: 404 GKLCGNSLGYAVELVRKLKAEVTEEY-MHSVADLMVIKERCLFTTIRSC----------- 451
G+L N LG+A L+ K VT EY M + L K+ F + +
Sbjct: 322 GELISNELGFAASLISKAIKNVTNEYVMIGIEYLKNQKDLKKFQDLHALGSTEGPFYGNP 381
Query: 452 ---VVSDVTRIKLREVDFGWGEAVYGGAAKSGAGPFPGAAYIIPHKNVEGEEGFILLVCL 508
VVS +T + + +DFGWG+ Y G G F G + I+P +N +G IL CL
Sbjct: 382 NLGVVSWLT-LPMYGLDFGWGKEFYTG---PGTHDFDGDSLILPDQNEDG--SVILATCL 435
Query: 509 SFEAMKRFAKELDE 522
M+ F K E
Sbjct: 436 QVAHMEAFKKHFYE 449
>At2g40230 putative anthranilate
N-hydroxycinnamoyl/benzoyltransferase
Length = 433
Score = 150 bits (380), Expect = 1e-36
Identities = 118/396 (29%), Positives = 204/396 (50%), Gaps = 21/396 (5%)
Query: 93 VRRSQPELVPPAAPTPREVKLLSDIDDQEGLRFNIPMMFIYRHEPSMKEKDPVKVLRHAL 152
V + ++ P+ TP V LS +D Q LRF I + +Y S E L+ AL
Sbjct: 5 VHVKEATVITPSDQTPSSVLSLSALDSQLFLRFTIEYLLVYP-PVSDPEYSLSGRLKSAL 63
Query: 153 SQALVYYYPFAGRIRE--GAGRKLMVDCTGEGVMFVEAEADVTLDEFGDALHPP--LPCF 208
S+ALV Y+PF+GR+RE G L V+C G+G +F+EA +D+ D PP + +
Sbjct: 64 SRALVPYFPFSGRVREKPDGGGGLEVNCRGQGALFLEAVSDILTCL--DFQKPPRHVTSW 121
Query: 209 EELLYDVPGSELIIDRPIRLIQVTRLKCGSFILVVYLNHTMSDGAGLKLFMNAWAEMARG 268
+LL + +++ P ++Q+T L+ G L V +NH +SDG G F+ +AE+++
Sbjct: 122 RKLL-SLHVIDVLAGAPPLVVQLTWLRDGGAALAVGVNHCVSDGIGSAEFLTLFAELSKD 180
Query: 269 AHKPSI---QPVWNREILMARDPPHITCNHHEYEQIFS-SNTIKEEDTTTLVHQSFFFRT 324
+ + + +W+R++LM P + +H E+ ++ + + LV S F
Sbjct: 181 SLSQTELKRKHLWDRQLLMP-SPIRDSLSHPEFNRVPDLCGFVNRFNAERLVPTSVVFER 239
Query: 325 SDIVVLRLLVP----FHLRHCTTFDLITSCFWCCRTKALQLEADDEIRMMCIVNARSRFN 380
+ L+ L F+ +H T+F+++++ W ++L L ++ ++++ VN R R
Sbjct: 240 QKLNELKKLASRLGEFNSKH-TSFEVLSAHVWRSWARSLNLPSNQVLKLLFSVNIRDRVK 298
Query: 381 ANNSPLVGYYGNCFAYPAAVTTAGKLCGNSLGYAVELVRKLKAEVTEEYMHSVADLMVIK 440
+ G+YGN F A TT L L YA LV++ K V +EY+ SV + V K
Sbjct: 299 PSLPS--GFYGNAFVVGCAQTTVKDLTEKGLSYATMLVKQAKERVGDEYVRSVVE-AVSK 355
Query: 441 ERCLFTTIRSCVVSDVTRIKLREVDFGWGEAVYGGA 476
ER ++ ++S +R+ L ++DFG G+ V+ G+
Sbjct: 356 ERASPDSVGVLILSQWSRLGLEKLDFGLGKPVHVGS 391
>At3g62160 unknown protein
Length = 428
Score = 146 bits (368), Expect = 3e-35
Identities = 115/413 (27%), Positives = 193/413 (45%), Gaps = 67/413 (16%)
Query: 89 LLFTVRRSQPELVPPAAPTPREVKLLSDIDDQEGLRFNIPMMFIYRHEPSMKEKDPVKVL 148
+ F++ R L+ TP V LS ID+ LR + ++ H P D + +
Sbjct: 1 MAFSLTRVNRVLIQACETTPSVVLDLSLIDNIPVLRCFARTIHVFTHGP-----DAARAI 55
Query: 149 RHALSQALVYYYPFAGRIREGAGRKLMVDCTGE-GVMFVEAEADVTLDEFGDALHPPLPC 207
+ AL++ALV YYP +GR++E KL +DCTG+ G+ FV+A DV L+ G
Sbjct: 56 QEALAKALVSYYPLSGRLKELNQGKLQIDCTGKTGIWFVDAVTDVKLESVG--------- 106
Query: 208 FEELLYDVPGSELIIDR--------PIRLIQVTRLKCGSFILVVYLNHTMSDGAGLKLFM 259
+ + L ++P +L+ D+ P+ +QVT+ CG F++ + H + DG G F+
Sbjct: 107 YFDNLMEIPYDDLLPDQIPKDDDAEPLVQMQVTQFSCGGFVMGLRFCHAICDGLGAAQFL 166
Query: 260 NAWAEMARGAHKPSIQPVWNREIL----MARD------PPHITCNHHEYEQIFSSNTIKE 309
A E+A G + PVW+R+ + A D PP EY+ S I
Sbjct: 167 TAVGEIACGQTNLGVSPVWHRDFIPQQHRANDVISPAIPPPFPPAFPEYKLEHLSFDI-P 225
Query: 310 EDTTTLVHQSFFFRTSDIVVLRLLVPFHLRHCTTFDLITSCFWCCRTKALQLEADDEIRM 369
D + F T +I C+ F++I +CFW RT+A+ L D E+++
Sbjct: 226 SDLIERFKREFRASTGEI-------------CSAFEVIAACFWKLRTEAINLREDTEVKL 272
Query: 370 MCIVNARSRFNANNSPL-VGYYGNCFAYPAAVTTAGKLCG--NSLGYAVELVRKLKAEVT 426
+ N R + N PL +G+YGNCF +P ++ S+ V+L+++ K ++
Sbjct: 273 VFFANCR---HLVNPPLPIGFYGNCF-FPVKISAESHEISKEKSVFDVVKLIKQAKTKLP 328
Query: 427 EEYMHSVAD------LMVIKERCLFTTIRSCVVSDVTRIKLREVDFGWGEAVY 473
EE+ V+ + LF +S+ ++ +VD+GWG ++
Sbjct: 329 EEFEKFVSGDGDDPFAPAVGYNTLF-------LSEWGKLGFNQVDYGWGPPLH 374
>At5g57840 N-hydroxycinnamoyl/benzoyltransferase
Length = 443
Score = 143 bits (360), Expect = 3e-34
Identities = 126/456 (27%), Positives = 203/456 (43%), Gaps = 44/456 (9%)
Query: 93 VRRSQPELVPPAAPTPREVKLLSDIDDQEGLRFNIPMMFIYRHEP-SMKEKDPVKVLRHA 151
+R Q +V PA TP LS++D ++ + M +Y ++P S ++ + L A
Sbjct: 3 IRVKQATIVKPAEETPTHNLWLSNLDL---IQVRLHMGTLYFYKPCSSSDRPNTQSLIEA 59
Query: 152 LSQALVYYYPFAGRIREGAGRKLMVDCTGEGVMFVEAEADVTLDEFGDALHPPLPCFEEL 211
LS+ LV++YP AGR+++ +L V C GEGV+FVEAE+D T+ + G L +L
Sbjct: 60 LSKVLVFFYPAAGRLQKNTNGRLEVQCNGEGVLFVEAESDSTVQDIG--LLTQSLDLSQL 117
Query: 212 LYDVPGSELIIDRPIRLIQVTRLKCGSFILVVYLNHTMSDGAGLKLFMNAWAEMARGAHK 271
+ V + I P+ L QVT KCG+ + ++HT + L M AW+ ARG
Sbjct: 118 VPTVDYAGDISSYPLLLFQVTYFKCGTICVGSSIHHTFGEATSLGYIMEAWSLTARGL-L 176
Query: 272 PSIQPVWNREILMARDPPHITCNHHEYEQIFSSNTIKEEDTTTLVHQSFFFRTSDIVVLR 331
+ P +R +L AR+PP H EY+ S +L ++S S I L+
Sbjct: 177 VKLTPFLDRTVLHARNPPSSVFPHTEYQ----SPPFHNHPMKSLAYRSNPESDSAIATLK 232
Query: 332 L----LVPFHLR------HCTTFDLITSCFWCCRTKALQ-LEADDEIRMMCIVNARSRFN 380
L L R +T++++ + W C + A + L + R+ I++ R R
Sbjct: 233 LTRLQLKALKARAEIADSKFSTYEVLVAHTWRCASFANEDLSEEHSTRLHIIIDGRPRLQ 292
Query: 381 ANNSPLVGYYGNCFAYPAAVTTAGKLCGNSLGYAVELVRKLKAEVTEEYMHSVAD----- 435
GY GN + V+ G S VE V + ++ EY+ S D
Sbjct: 293 PKLPQ--GYIGNTLFHARPVSPLGAFHRESFSETVERVHREIRKMDNEYLRSAIDYLERH 350
Query: 436 -----LMVIKERCLFTTIRSCVVSDVTRIKLREVDFGWGEAVYGGAA--KSGAGPFPGAA 488
L+ + +F+ + + + + E+DFGWG+A Y A+ G G G
Sbjct: 351 PDLDQLVPGEGNTIFSCAANFCIVSLIKQTAYEMDFGWGKAYYKRASHLNEGKGFVTG-- 408
Query: 489 YIIPHKNVEGEEGFILLVCLSFEAMKRFAKELDEML 524
N + E +L +CL + +F K E L
Sbjct: 409 ------NPDEEGSMMLTMCLKKTQLSKFRKLFYEFL 438
>At1g28680 anthranilate N-hydroxycinnamoyl/benzoyltransferase like
protein
Length = 451
Score = 136 bits (342), Expect = 3e-32
Identities = 114/421 (27%), Positives = 191/421 (45%), Gaps = 17/421 (4%)
Query: 114 LSDIDDQEGLRFNIPMMFIYRHEPS-MKEKDPVKVLRHALSQALVYYYPFAGRIREGAG- 171
LS +D+ L + + +Y S + + P V+ +L+ ALV+YYP AG +R A
Sbjct: 27 LSHLDNDNNLHVSFRYLRVYSSSSSTVAGESPSAVVSASLATALVHYYPLAGSLRRSASD 86
Query: 172 -RKLMVDCTGEGVMFVEAEADVTLDEFGDALHPPLPCFEELLYDVPGSELIIDRPIRLIQ 230
R ++ G+ V V A + TL+ G L P P F E L P E + P ++Q
Sbjct: 87 NRFELLCSAGQSVPLVNATVNCTLESVG-YLDGPDPGFVERLVPDPTREEGMVNPC-ILQ 144
Query: 231 VTRLKCGSFILVVYLNHTMSDGAGLKLFMNAWAEMARGAHKPSIQPVWNREILMA-RDPP 289
VT +CG ++L ++H + DG G LF NA AE+ARGA K SI+PVW+RE L+ R+ P
Sbjct: 145 VTMFQCGGWVLGASIHHAICDGLGASLFFNAMAELARGATKISIEPVWDRERLLGPREKP 204
Query: 290 HITCNHHEYEQIFSSNTIKEEDTTTLVHQSFFFRTSDIVVLRL-LVPFHLRHCTTFDLIT 348
+ ++ + + + FF + L+ L+ + TTF+ +
Sbjct: 205 WVGAPVRDFLSLDKDFDPYGQAIGDVKRDCFFVTDDSLDQLKAQLLEKSGLNFTTFEALG 264
Query: 349 SCFWCCRTKALQLEADDEIRMMCIVNARSRFNANNSPL-VGYYGNCFAYPAAVTTAGKLC 407
+ W + +A + E + ++ + +N R N PL GY+GN A AG+L
Sbjct: 265 AYIWRAKVRAAKTEEKENVKFVYSINIR---RLMNPPLPKGYWGNGCVPMYAQIKAGELI 321
Query: 408 GNSLGYAVELVRKLKAEVTEEYMHSVADLMVIKERCLFTTIRSCV-VSDVTRIKLREVDF 466
+ EL+++ K+ ++EY+ S D + + +D + +DF
Sbjct: 322 EQPIWKTAELIKQSKSNTSDEYVRSFIDFQELHHKDGINAGTGVTGFTDWRYLGHSTIDF 381
Query: 467 GWGEAVYGGAAKSGAGPFPGAAYIIPHK-----NVEGEEGFILLVCLSFEAMKRFAKELD 521
GWG V + + +P+ + + GF +LV L AM F + +D
Sbjct: 382 GWGGPVTVLPLSNKLLGSMEPCFFLPYSTDAAAGSKKDSGFKVLVNLRESAMPEFKEAMD 441
Query: 522 E 522
+
Sbjct: 442 K 442
>At5g07080 unknown protein
Length = 450
Score = 125 bits (313), Expect = 8e-29
Identities = 114/410 (27%), Positives = 179/410 (42%), Gaps = 36/410 (8%)
Query: 86 SAPLLFTVRRSQPELVPPAAPTPREVKLLSDIDDQEGLRFNIPMMFIYRH-EPSMKEKDP 144
S+P V++SQ +V P+ TP LS +D+ L +++Y P+ DP
Sbjct: 7 SSPSPLVVKKSQVVIVKPSKATPDVSLSLSTLDNDPYLETLAKTIYVYAPPSPNDHVHDP 66
Query: 145 VKVLRHALSQALVYYYPFAGRIREGA-GRKLMVDCT-GEGVMFVEAEADVTLDEFGDALH 202
+ + ALS ALVYYYP AG++ G+ +L + C+ EGV FV A AD TL
Sbjct: 67 ASLFQQALSDALVYYYPLAGKLHRGSHDHRLELRCSPAEGVPFVRATADCTLSSLNYLKD 126
Query: 203 PPLPCFEELLYDV--PGSELIIDRPIRLIQVTRLKCGSFILVVYLNHTMSDGAGLKLFMN 260
++ + DV P + P+ L Q+T CG L L+H++ DG G F
Sbjct: 127 MDTDLYQLVPCDVAAPSGDY---NPLAL-QITLFACGGLTLATALSHSLCDGFGASQFFK 182
Query: 261 AWAEMARGAHKPSIQPVWNREILMARDPPHITCNHHEYE-----------QIFSSNTIKE 309
E+A G +PSI PVW+R L + + + N E S+ T
Sbjct: 183 TLTELAAGKTQPSIIPVWDRHRLTSNN---FSLNDQVEEGQAPKLVDFGEACSSAATSPY 239
Query: 310 EDTTTLVHQSFFFRTSDIVVLRLLVPFHLRHCTTFDLITSCFWCCRTKALQLEADDEIRM 369
+ +V + + DI L+ V + TT +++ + W R +AL+L D
Sbjct: 240 TPSNDMVCEILNVTSEDITQLKEKVAGVV---TTLEILAAHVWRARCRALKLSPDGTSLF 296
Query: 370 MCIVNARSRFNANNSPLV-GYYGNCFAYPAAVTTAGKLCGNSLGYAVELVRKLKAEVTE- 427
V R PL GYYGN F AG+L + L + V+L+++ K E
Sbjct: 297 GMAVGIR---RIVEPPLPEGYYGNAFVKANVAMKAGELSNSPLSHVVQLIKEAKKAAQEK 353
Query: 428 ----EYMHSVADLMVIKERCLFTTIRSCVVSDVTRI-KLREVDFGWGEAV 472
E + + + C +++D ++ L E+DFG+G +V
Sbjct: 354 RYVLEQLRETEKTLKMNVACEGGKGAFMLLTDWRQLGLLDEIDFGYGGSV 403
>At3g47170 hypersensitivity-related protein-like protein
Length = 447
Score = 120 bits (301), Expect = 2e-27
Identities = 117/468 (25%), Positives = 204/468 (43%), Gaps = 66/468 (14%)
Query: 91 FTVRRSQPELVPPAAPTPREVKLLSDIDDQEGLRFNIPMMFIYRHEPSMKEKD---PVKV 147
F V +S+ LV P+ PTP LS ID+ + ++ ++ P +++ P
Sbjct: 10 FVVEKSEVVLVKPSKPTPDVTLSLSTIDNDPNNEVILDILCVFSPNPYVRDHTNYHPASF 69
Query: 148 LRHALSQALVYYYPFAGRI-REGAGRKLMVDCT-GEGVMFVEAEADVTLDEFG------- 198
++ ALS ALVYYYP AG++ R + + L +DCT G+GV ++A A TL
Sbjct: 70 IQLALSNALVYYYPLAGKLHRRTSDQTLQLDCTQGDGVPLIKATASCTLSSLNYLESGNH 129
Query: 199 -DALHPPLPCFEELLYDVPGSELIIDRPIRLIQVTRLKCGSFILVVYLNHTMSDGAGLKL 257
+A + +P + L G L RP+ L Q+T+ CG + +HT+ DG G+
Sbjct: 130 LEATYQLVPSHDTL----KGCNLGY-RPLAL-QITKFTCGGMTIGSVHSHTVCDGIGMAQ 183
Query: 258 FMNAWAEMARGAHKPSIQPVWNREILMARDPPHI-----TCNHHEYEQIFSSNTIKEEDT 312
F A E+A G +P++ PVW+R ++ + + N+ + + + + T
Sbjct: 184 FSQAILELAAGRAQPTVIPVWDRHMITSNQISTLCKLGNDKNNPKLVDVEKDCSSPDTPT 243
Query: 313 TTLVHQSFFFRTSDIVVLRLLV---------PFHLRHCTTFDLITSCFWCCRTKALQLEA 363
+V + + DI L+ ++ TT +++ + W R +A++L
Sbjct: 244 EDMVREILNITSEDITKLKNIIIEDENLTNEKEKNMKITTVEVLAAYVWRARCRAMKLNP 303
Query: 364 DDEIRMMCIVNARSRFNANNSPL-VGYYGNCFAYPAAVTTAGKLCGNSLGYAVELVRKLK 422
D ++ V+ RS PL GYYGN F + + TA +L + V L++ K
Sbjct: 304 DTITDLVISVSIRSSI---EPPLPEGYYGNAFTHASVALTAKELSKTPISRLVRLIKDAK 360
Query: 423 AEVTE-----EYMHSVADLMVIKERCLFTTIRSCVVSDVTRIKLREVDFGWGEAVYGGAA 477
+ E + + + M +K I V +T + G + V+G +
Sbjct: 361 RAALDNGYVCEQLREMENTMKLK--LASKEIHGGVFMMLTDWR----HLGLDQDVWGAS- 413
Query: 478 KSGAGPFPGAAYIIPHKNVEGEEGFI-LLVCLSFEAMKRFAKELDEML 524
K V G+ G + +L L +AM +F +E+D +L
Sbjct: 414 ----------------KAVPGKSGGVRVLTTLPRDAMAKFKEEIDAVL 445
>At2g23510 putative anthranilate
N-hydroxycinnamoyl/benzoyltransferase
Length = 451
Score = 111 bits (278), Expect = 9e-25
Identities = 104/356 (29%), Positives = 166/356 (46%), Gaps = 32/356 (8%)
Query: 82 HIMTSAPLLFTVRRSQPELVPPAAPTPREVKLLSDIDDQEGLRFNIPMMFIYRHEPSMKE 141
HI +S PL+ V + E+V P+ P++ LS +D+ ++++ + +
Sbjct: 4 HIGSSIPLM--VEKMLTEMVKPSKHIPQQTLNLSTLDNDPYNEVIYKACYVFKAKNVADD 61
Query: 142 KD-PVKVLRHALSQALVYYYPFAGRI-REGAGRKLMVDCTGEG--VMFVEAEADVTLDEF 197
+ P +LR ALS L YYYP +G + R+ + RKL + C G+G V F A A+V L
Sbjct: 62 DNRPEALLREALSDLLGYYYPLSGSLKRQESDRKLQLSCGGDGGGVPFTVATANVELSSL 121
Query: 198 GDALHPPLPCFEELLYDVPGSELIID--RPIRLIQVTRLKCGSFILVVYLNHTMSDGAGL 255
+ + + L +P + ID RP L QVT+ +CG FIL + ++H M DG G
Sbjct: 122 KNLENIDS---DTALNFLPVLHVDIDGYRPFAL-QVTKFECGGFILGMAMSHAMCDGYGE 177
Query: 256 KLFMNAWAEMARGAHKPSIQPVWNREILMAR----DPPHITCNHHEYEQIFSSNTIKEED 311
M A ++A G KP + P+W RE L+ + PP + + +S + +D
Sbjct: 178 GHIMCALTDLAGGKKKPMVTPIWERERLVGKPEDDQPPFV-----PGDDTAASPYLPTDD 232
Query: 312 TTTLVHQSFFFRTSDIVVLRLLV----PFHLRHCTTFDLITSCFWCCRTKALQLEADDEI 367
T + R I L+ F TTF++I + W R KAL L+ D
Sbjct: 233 WVT---EKITIRADSIRRLKEATLKEYDFSNETITTFEVIGAYLWKSRVKALNLDRDGVT 289
Query: 368 RMMCIVNARSRFNANNSPLV-GYYGNCFAYPAAVTTAGKLCGNSLGYAVELVRKLK 422
+ V R N + PL GYYGN + TA ++ ++ V+L+++ K
Sbjct: 290 VLGLSVGIR---NVVDPPLPDGYYGNAYIDMYVPLTAREVEEFTISDIVKLIKEAK 342
>At2g25150 unknown protein
Length = 461
Score = 110 bits (274), Expect = 3e-24
Identities = 121/462 (26%), Positives = 205/462 (44%), Gaps = 42/462 (9%)
Query: 88 PLLFTVRRSQP-ELVPPAAPTPREVKLLSDIDDQEGLRFNIPMMFIYRHEPSMKEKDPVK 146
P+L + +P ELV P+ T E LS +D+ +++++ + DPV
Sbjct: 7 PILPLLLEKKPVELVKPSKHTHCETLSLSTLDNDPFNEVMYATIYVFKANGKNLD-DPVS 65
Query: 147 VLRHALSQALVYYYPFAGRI-REGAGRKLMVDCTGEGVMFVEAEADVTLDEFGDALH--- 202
+LR ALS+ LV+YYP +G++ R + KL + GEGV F A + + L +
Sbjct: 66 LLRKALSELLVHYYPLSGKLMRSESNGKLQLVYLGEGVPFEVATSTLDLSSLNYIENLDD 125
Query: 203 -PPLPCFEELLYDVPGSELIIDRPIRLIQVTRLKCGSFILVVYLNHTMSDGAGLKLFMNA 261
L E+ D + + P+ L QVT+ CG F + L H + DG G+ ++A
Sbjct: 126 QVALRLVPEIEIDYESN--VCYHPLAL-QVTKFACGGFTIGTALTHAVCDGYGVAQIIHA 182
Query: 262 WAEMARGAHKPSIQPVWNREILMAR---DPPHITCNHHEYEQIFSSNTIKEEDTTTLVHQ 318
E+A G +PS++ VW RE L+ + P + +H + S+ TT +V +
Sbjct: 183 LTELAAGKTEPSVKSVWQRERLVGKIDNKPGKVPGSHIDGFLATSAYL----PTTDVVTE 238
Query: 319 SFFFRTSDIVVLRLLVPFHLRHC-------TTFDLITSCFWCCRTKALQLEADDEIRMMC 371
+ R DI + L ++ C TT+++++S W R++AL+L D +
Sbjct: 239 TINIRAGDI---KRLKDSMMKECEYLKESFTTYEVLSSYIWKLRSRALKLNPDGITVLGV 295
Query: 372 IVNARSRFNANNSPL-VGYYGNCFAYPAAVTTAGKLCGNSLGYAVELVRKLKAEV----- 425
V R + + PL GYYGN + T +L +S+ V+K K
Sbjct: 296 AVGIR---HVLDPPLPKGYYGNAYIDVYVELTVRELEESSISNIANRVKKAKKTAYEKGY 352
Query: 426 TEEYMHSVADLMVIKERCLFTTIRSCV--VSDVTRIK-LREVDFGWGEAVYGGAAKSGAG 482
EE + + LM ++ +F + + ++D I +DFGW E V
Sbjct: 353 IEEELKNTERLM--RDDSMFEGVSDGLFFLTDWRNIGWFGSMDFGWNEPVNLRPLTQRES 410
Query: 483 PFPGAAYIIPHKNVEGEEGFI-LLVCLSFEAMKRFAKELDEM 523
+ P K+ EG + +++ L +AM F +E+ M
Sbjct: 411 TVHVGMILKPSKSDPSMEGGVKVIMKLPRDAMVEFKREMATM 452
>At1g31490 putative protein
Length = 444
Score = 98.6 bits (244), Expect = 8e-21
Identities = 103/406 (25%), Positives = 167/406 (40%), Gaps = 43/406 (10%)
Query: 90 LFTVRRSQPELVPPAAPTPREVKLL--SDIDDQEGLRFNIPMMFIYRHEPSMKEKDPVKV 147
+F V + +V + P +V L S+ D G RF + + Y +P + + VK
Sbjct: 3 IFDVSFTGEFIVKASGDQPEKVNSLNLSNFDLLSG-RFPVTYFYFYPKQPQLSFETIVKS 61
Query: 148 LRHALSQALVYYYPFAGRI--REGAGRKLMVDCTGEGVMFVEAEADVTLDEFGDALHPPL 205
L+ +LSQ L Y+YPFAG+I E + + M+ C G +FVEA A V L D +
Sbjct: 62 LQSSLSQTLSYFYPFAGQIVPNETSQEEPMIICNNNGALFVEARAHVDLKSL-DFYNLDA 120
Query: 206 PCFEELLYDVPGSELIIDRPIRLIQVTRLKCGSFILVVYLNHTMSDGAGLKLFMNAWAEM 265
+L+ P L IQ T +CG L +H + D + F+ W+E+
Sbjct: 121 FLQSKLVQVNPDFSL-------QIQATEFECGGLALTFTFDHALGDASSFGKFLTLWSEI 173
Query: 266 ARGAHKP-SIQPVWNREILMARDPPHITCNHHEYEQIFSSNTIK--EED------TTTLV 316
+R +KP S P R +L AR PP Y+ IK EED + TL+
Sbjct: 174 SR--NKPVSCVPDHRRNLLRARSPP-------RYDPHLDKTFIKCSEEDIKNIPMSKTLI 224
Query: 317 HQSFFFRTSDIVVLRLLVPFHLRHCTTFDLITSCFWCCRTKALQLEADDEIRMMCIVNAR 376
+ + S + L+ L + T + ++ W +++ +M +V+ R
Sbjct: 225 KRLYHIGASSLDALQALATVNGESRTKIEAFSAYVWKKMVDSIE-SGHKTCKMGWLVDGR 283
Query: 377 SRFNANNSPLVGYYGNCFAYPAAVTTAGKLCGNSLGYAVELVRKLKAEVTE--------E 428
R S Y GN + + L N + +V K EVT +
Sbjct: 284 GRLETVTS---SYIGNVLSIAVGEASIENLKKNHVSDIANIVHKSITEVTNDTHFTDLID 340
Query: 429 YMHSVADLMVIKERCLFTTIRSCVVSDVTRIKLREVDFGWGEAVYG 474
++ S +++ L + V+S R + E+DFG+G G
Sbjct: 341 WIESHRPGLMLARVVLGQEGPALVLSSGRRFPVAELDFGFGAPFLG 386
>At3g26040 hypothetical protein
Length = 442
Score = 95.1 bits (235), Expect = 8e-20
Identities = 95/401 (23%), Positives = 172/401 (42%), Gaps = 50/401 (12%)
Query: 99 ELVPPAAPTPREVKLLS-DIDDQEGLRFNIPMMFIYRHEPSMKEKDPVKVLRHALSQALV 157
+++ P++PTP +K + +Q G PM+F Y S+K + +++L+ +LS+ L
Sbjct: 9 DIIKPSSPTPNHLKKFKLSLLEQLGPTIFGPMVFFYSANNSIKPTEQLQMLKKSLSETLT 68
Query: 158 YYYPFAGRIREGAGRKLMVDCTGEGVMFVEAEADVTLDEFGDALHPPLPCFEELLYDVPG 217
++YP AGR++ + +DC G F+EA + L L P ++L +P
Sbjct: 69 HFYPLAGRLK----GNISIDCNDSGADFLEARVNSPLSNL--LLEPSSDSLQQL---IPT 119
Query: 218 SELIIDRPIRLI--QVTRLKCGSFILVVYLNHTMSDGAGLKLFMNAWAEM-ARGAHKPSI 274
S I+ RL+ Q + +CGS + V ++H ++D + LFM +WA + +RG+ K
Sbjct: 120 SVDSIETRTRLLLAQASFFECGSMSIGVCISHKLADATSIGLFMKSWAAISSRGSIKTIG 179
Query: 275 QPVWNREILMARDPPHITCNHHEYEQIFSSNTIKEE-DTTTLVHQSFFFRTSDIVVLRLL 333
PV++ + PP + + + ++ E + + F F +S I L+
Sbjct: 180 APVFDTVKIF---PP------GNFSETSPAPVVEPEIMMNQTLSKRFIFDSSSIQALQAK 230
Query: 334 V-PFHLRHCTTFDLITSCFWCCRTKALQLEADDEIRMMCIVNARSRFNANNSPLV-GYYG 391
F + T + +++ W KA + + + + N+ S + + P G
Sbjct: 231 ASSFEVNQPTRVEAVSALIWKSAMKATRTVSGTS-KPSILANSVSLRSRVSPPFTKNSIG 289
Query: 392 NCFAYPAAVTTAGKLCGNSLGYAVELVRKLK-----------------AEVTEEYMHSVA 434
N +Y AA G + L V +RK K E+ Y
Sbjct: 290 NLVSYFAAKAEEG-INQTKLQTLVSKIRKAKQRFRDIHIPKLVGNPNATEIICSYQKEAG 348
Query: 435 DLMVIKERCLFTTIRSCVVSDVTRIKLREVDFGWGEAVYGG 475
D++ + + + S R L E DFGWG+ V+ G
Sbjct: 349 DMIASGDFDFY------IFSSACRFGLYETDFGWGKPVWVG 383
>At3g50280 anthranilate N-hydroxycinnamoyl/benzoyltransferase - like
protein
Length = 443
Score = 94.4 bits (233), Expect = 1e-19
Identities = 103/400 (25%), Positives = 174/400 (42%), Gaps = 46/400 (11%)
Query: 107 TPREVKLLSDIDDQEGLRFNIPMMFIYRHEPSMKEKDPVKVLRHALSQALVYYYPFAGRI 166
TP ++ LL Q GL F P +P E + LR +LS AL Y+PFAGR+
Sbjct: 28 TPFDLNLLYVDYTQRGLLFPKP-------DP---ETHFISRLRTSLSSALDIYFPFAGRL 77
Query: 167 REGAGRK-----LMVDCTGEGVMFVEAEADVTLDEFGDALHPP--LPCFEELLYDVPGSE 219
+ + ++C G G F+ A +D D L P +P F + Y + G +
Sbjct: 78 NKVENHEDETVSFYINCDGSGAKFIHAVSDSV--SVSDLLRPDGSVPDFFRIFYPMNGVK 135
Query: 220 LI--IDRPIRLIQVTRLKCGSFILVVYLNHTMSDGAGLKLFMNAWAEMARGAHKPSIQPV 277
I + P+ +QVT ++ G FI Y NH ++DGA + F W+++ + ++QP+
Sbjct: 136 SIDGLSEPLLALQVTEMRDGVFIGFGY-NHMVADGASIWNFFRTWSKICSNGQRENLQPL 194
Query: 278 WNREILM--ARDPPHITCNHHEYEQIFSSNTIKEEDTTTLVHQSFFFRTSDIVVLRLLVP 335
+ + + P HI + E ++ E + T + F F +I L+ V
Sbjct: 195 ALKGLFVDGMDFPIHIPVSDTE------TSPPSRELSPTFKERVFHFTKRNISDLKAKVN 248
Query: 336 FHL----RHCTTFDLITSCFWCCRTKALQLEADDEIRMMCIVNARSRFNANNSPL-VGYY 390
+ ++ +++ W + L +++ R V+ R R N PL +
Sbjct: 249 GEIGLRDHKVSSLQAVSAHMWRSIIRHSGLNQEEKTRCFVAVDLRQRL---NPPLDKECF 305
Query: 391 GNCFAYPAAVTTAGKLCGNSLGYAVELVRKLKAEVTEE----YMHSVADLMVIKERCLFT 446
G+ TT G+L LG+A + + +T E Y + M I++ L +
Sbjct: 306 GHVIYNSVVTTTVGELHDQGLGWAFLQINNMLRSLTNEDYRIYAENWVRNMKIQKSGLGS 365
Query: 447 --TIRSCVVSDVTRIKLREVDFGWGE--AVYGGAAKSGAG 482
T S +VS R ++ + DFGWG+ AV G + S +G
Sbjct: 366 KMTRDSVIVSSSPRFEVYDNDFGWGKPIAVRAGPSNSISG 405
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.326 0.140 0.437
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,263,799
Number of Sequences: 26719
Number of extensions: 528820
Number of successful extensions: 1705
Number of sequences better than 10.0: 65
Number of HSP's better than 10.0 without gapping: 42
Number of HSP's successfully gapped in prelim test: 23
Number of HSP's that attempted gapping in prelim test: 1478
Number of HSP's gapped (non-prelim): 89
length of query: 528
length of database: 11,318,596
effective HSP length: 104
effective length of query: 424
effective length of database: 8,539,820
effective search space: 3620883680
effective search space used: 3620883680
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 62 (28.5 bits)
Medicago: description of AC148345.17


