

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148289.16 + phase: 0
(200 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
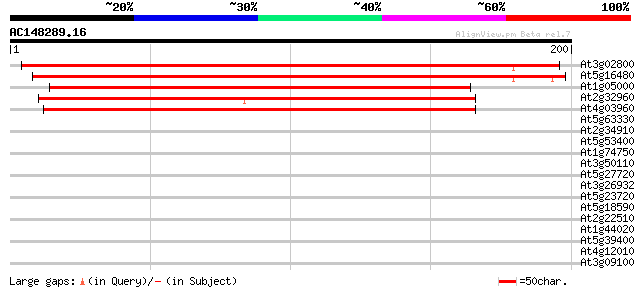
Score E
Sequences producing significant alignments: (bits) Value
At3g02800 unknown protein 278 1e-75
At5g16480 unknown protein 273 4e-74
At1g05000 unkown protein 204 3e-53
At2g32960 unknown protein 194 3e-50
At4g03960 unknown protein 192 1e-49
At5g63330 unknown protein 31 0.50
At2g34910 hypothetical protein 29 1.9
At5g53400 unknown protein 28 2.5
At1g74750 28 3.2
At3g50110 tyrosine phosphatase like protein 28 4.2
At5g27720 glycine rich protein - like 27 5.5
At3g26932 putative protein 27 5.5
At5g23720 unknown protein 27 7.2
At5g18590 RanGAP1 interacting protein 27 7.2
At2g22510 unknown protein 27 7.2
At1g44020 hypothetical protein 27 7.2
At5g39400 PTEN -like protein 27 9.4
At4g12010 like disease resistance protein (TMV N-like) 27 9.4
At3g09100 putative mRNA capping enzyme, RNA guanylyltransferase 27 9.4
>At3g02800 unknown protein
Length = 199
Score = 278 bits (711), Expect = 1e-75
Identities = 130/193 (67%), Positives = 158/193 (81%), Gaps = 1/193 (0%)
Query: 5 VENIDEDDDVLIPPPNFSMVEDCIYRSSLPKPSSFPFLQTLNLRSIIYLCPEPYPEENLD 64
+E D + DVL PP NFSMVED IYRS P+P +F FL+TLNLRSIIYLCPEPYPEENL
Sbjct: 1 METDDHNGDVLAPPSNFSMVEDGIYRSGFPRPENFSFLKTLNLRSIIYLCPEPYPEENLK 60
Query: 65 FLKEQNIRLFQFGIEGKTEVSLPALRDSIMEALKVLVDVRNHPILVHCKQGKHRTGCLVG 124
FL+ NI+L+QFGIEGKT+ P +D++++ALKVLVDVRNHPIL+HCK+GKHRTGCLVG
Sbjct: 61 FLEANNIKLYQFGIEGKTDPPTPMPKDTVLDALKVLVDVRNHPILIHCKRGKHRTGCLVG 120
Query: 125 CFRKLQNWCLSSAFEEYQRFAGVKSRAADLTFIERFDLVSLRQCLYSIIYQYQG-ASKKR 183
C RK+Q+WCLSS EEYQ+ AG+K R DL FIE FD+VSLRQCL SI+YQY G K+R
Sbjct: 121 CLRKVQSWCLSSVLEEYQKNAGLKWRQRDLNFIEAFDIVSLRQCLLSIMYQYHGYGFKRR 180
Query: 184 RLMYQDENIQKPR 196
RL Y++EN++ P+
Sbjct: 181 RLAYEEENVKTPK 193
>At5g16480 unknown protein
Length = 204
Score = 273 bits (698), Expect = 4e-74
Identities = 131/195 (67%), Positives = 156/195 (79%), Gaps = 5/195 (2%)
Query: 9 DEDDDVLIPPPNFSMVEDCIYRSSLPKPSSFPFLQTLNLRSIIYLCPEPYPEENLDFLKE 68
D D +VLIPPPNFSMVED IYRS P+ +F FL TLNLRSIIYLCPEPYPE+NL L
Sbjct: 8 DNDGEVLIPPPNFSMVEDEIYRSGFPELENFGFLSTLNLRSIIYLCPEPYPEDNLKSLAS 67
Query: 69 QNIRLFQFGIEGKTEVSLPALRDSIMEALKVLVDVRNHPILVHCKQGKHRTGCLVGCFRK 128
NI+LFQFGIEGKT+ P +D+++ AL+VLVDVRNHPIL+HCK+GKHRTGCLVGC RK
Sbjct: 68 NNIKLFQFGIEGKTDPPTPMPKDTVLSALRVLVDVRNHPILIHCKRGKHRTGCLVGCLRK 127
Query: 129 LQNWCLSSAFEEYQRFAGVKSRAADLTFIERFDLVSLRQCLYSIIYQYQG-ASKKRRLMY 187
+QNWCLSS EEYQ+ AG+K R DL FIE FD++ L+QCLYSIIYQY G K+R+L+Y
Sbjct: 128 VQNWCLSSVLEEYQKCAGLKWRQRDLRFIEDFDVLRLKQCLYSIIYQYNGYGLKRRKLLY 187
Query: 188 QDENI----QKPRLT 198
Q+EN+ QKP+ T
Sbjct: 188 QEENVVQEQQKPQAT 202
>At1g05000 unkown protein
Length = 215
Score = 204 bits (519), Expect = 3e-53
Identities = 97/150 (64%), Positives = 114/150 (75%)
Query: 15 LIPPPNFSMVEDCIYRSSLPKPSSFPFLQTLNLRSIIYLCPEPYPEENLDFLKEQNIRLF 74
LIPP NFSMV++ I+RS P ++F FLQTL LRSIIYLCPEPYPE NL FLK IRLF
Sbjct: 53 LIPPLNFSMVDNGIFRSGFPDSANFSFLQTLGLRSIIYLCPEPYPESNLQFLKSNGIRLF 112
Query: 75 QFGIEGKTEVSLPALRDSIMEALKVLVDVRNHPILVHCKQGKHRTGCLVGCFRKLQNWCL 134
QFGIEG E + I ALKVL+D +NHP+L+HCK+GKHRTGCLVGC RKLQ WCL
Sbjct: 113 QFGIEGNKEPFVNIPDHKIRMALKVLLDEKNHPVLIHCKRGKHRTGCLVGCLRKLQKWCL 172
Query: 135 SSAFEEYQRFAGVKSRAADLTFIERFDLVS 164
+S F+EYQRFA K+R +D F+E FD+ S
Sbjct: 173 TSIFDEYQRFAAAKARVSDQRFMEIFDVSS 202
>At2g32960 unknown protein
Length = 218
Score = 194 bits (492), Expect = 3e-50
Identities = 92/158 (58%), Positives = 116/158 (73%), Gaps = 2/158 (1%)
Query: 11 DDDVLIPPPNFSMVEDCIYRSSLPKPSSFPFLQTLNLRSIIYLCPEPYPEENLDFLKEQN 70
D+ LIPP NFSMV++ I+RS P ++F F++TL LRSII LCPEPYPE N+ FLK
Sbjct: 50 DELNLIPPLNFSMVDNGIFRSGFPDSANFSFIKTLGLRSIISLCPEPYPENNMQFLKSNG 109
Query: 71 IRLFQFGIEGKT--EVSLPALRDSIMEALKVLVDVRNHPILVHCKQGKHRTGCLVGCFRK 128
I LFQFGIEG E + L I EALKVL+D +NHP+L+HCK+GKHRTGCLVGC RK
Sbjct: 110 ISLFQFGIEGSKSKEPFVDILDQKIREALKVLLDEKNHPLLIHCKRGKHRTGCLVGCMRK 169
Query: 129 LQNWCLSSAFEEYQRFAGVKSRAADLTFIERFDLVSLR 166
LQ WC++S +EY+RFA K+R +D F+E FD+ L+
Sbjct: 170 LQKWCITSILDEYKRFAAAKARVSDQRFLESFDVSGLK 207
>At4g03960 unknown protein
Length = 198
Score = 192 bits (487), Expect = 1e-49
Identities = 88/154 (57%), Positives = 115/154 (74%)
Query: 13 DVLIPPPNFSMVEDCIYRSSLPKPSSFPFLQTLNLRSIIYLCPEPYPEENLDFLKEQNIR 72
++ +PP NF+MV++ I+RS P+P SF FLQ+L L+SIIYLCPE YPE N +F K I+
Sbjct: 27 ELFVPPLNFAMVDNGIFRSGFPEPVSFSFLQSLRLKSIIYLCPEAYPEVNREFAKSNGIQ 86
Query: 73 LFQFGIEGKTEVSLPALRDSIMEALKVLVDVRNHPILVHCKQGKHRTGCLVGCFRKLQNW 132
+FQFGIE E + + I EAL+VL+D NHP+L+HCK GKHRTGCLVGC RK+Q W
Sbjct: 87 VFQFGIERCKEPFVNIPDEVIREALQVLLDTENHPVLIHCKSGKHRTGCLVGCVRKIQRW 146
Query: 133 CLSSAFEEYQRFAGVKSRAADLTFIERFDLVSLR 166
CLSS F+EYQRFA K+R +D F+E FD+ +L+
Sbjct: 147 CLSSIFDEYQRFAAAKARISDQRFMELFDISNLK 180
>At5g63330 unknown protein
Length = 477
Score = 30.8 bits (68), Expect = 0.50
Identities = 17/47 (36%), Positives = 29/47 (61%), Gaps = 1/47 (2%)
Query: 56 EPYPEENLDFLKEQNIRLFQFGIEGKTEVSLPALRDSIMEALKVLVD 102
E +P++ D L+EQ+ Q G EG+ E+ + AL D I+ ++ L+D
Sbjct: 331 EDFPQKIADLLREQSGSDGQSG-EGEIEIDIEALSDEILFMVRKLLD 376
>At2g34910 hypothetical protein
Length = 288
Score = 28.9 bits (63), Expect = 1.9
Identities = 26/102 (25%), Positives = 46/102 (44%), Gaps = 6/102 (5%)
Query: 1 MIVEVENIDEDDDVLIPPPNFSMVED-CIYRSSLPKPSSFPFLQTLNLRSIIYLCPEPYP 59
M V +++E+ ++ P S D + LP+P P + + S++ L YP
Sbjct: 50 MTVSRPSLNEESFRMVLPLAMSPPRDNAVPLPVLPEPMMKPRKKLSHQESMLSLRKSRYP 109
Query: 60 EENLDFLKEQNIRLFQFGIE----GKTEVSLPALRDSIMEAL 97
E+N + +E+N + F + GK V P DSI + +
Sbjct: 110 EKNF-YQEEENFKCNAFCLSLPGFGKRPVRSPKSEDSIKKKM 150
>At5g53400 unknown protein
Length = 304
Score = 28.5 bits (62), Expect = 2.5
Identities = 21/77 (27%), Positives = 39/77 (50%), Gaps = 5/77 (6%)
Query: 22 SMVEDCIYRSSLPKPSSFPFLQTLNLRSIIYLCPEPYPEENLDFLKEQNIRLFQFGIEGK 81
+++ + SS +P FPF TL+ + P + E+ DFL EQ+ L + E +
Sbjct: 2 AIISEVEEESSSSRPMIFPFRATLSSAN-----PLGFLEKVFDFLGEQSDFLKKPSAEDE 56
Query: 82 TEVSLPALRDSIMEALK 98
V++ A ++ + +A K
Sbjct: 57 IVVAVRAAKEKLKKAEK 73
>At1g74750
Length = 855
Score = 28.1 bits (61), Expect = 3.2
Identities = 14/47 (29%), Positives = 23/47 (48%), Gaps = 4/47 (8%)
Query: 93 IMEALKVLVDVRNHPILVHCKQGKHRTGCLVGCFRKLQNWCLSSAFE 139
+ +A++ L+++ N P +GC VG L+NW L S E
Sbjct: 808 VRQAVEELLNIFNFPFFTE----NGNSGCFVGSGEPLKNWLLESYVE 850
>At3g50110 tyrosine phosphatase like protein
Length = 632
Score = 27.7 bits (60), Expect = 4.2
Identities = 11/24 (45%), Positives = 17/24 (70%), Gaps = 1/24 (4%)
Query: 102 DVRNHPILVHCKQGKHRTGCLVGC 125
D++N ++VHCK G RTG ++ C
Sbjct: 298 DIQN-VVVVHCKAGMARTGLMICC 320
>At5g27720 glycine rich protein - like
Length = 129
Score = 27.3 bits (59), Expect = 5.5
Identities = 13/32 (40%), Positives = 17/32 (52%)
Query: 94 MEALKVLVDVRNHPILVHCKQGKHRTGCLVGC 125
M L +L + HP+LV K G+ G LV C
Sbjct: 1 MLPLSLLKTAQGHPMLVELKNGETYNGHLVNC 32
>At3g26932 putative protein
Length = 357
Score = 27.3 bits (59), Expect = 5.5
Identities = 19/50 (38%), Positives = 24/50 (48%), Gaps = 4/50 (8%)
Query: 17 PPPNFSMVEDCIYRSSLPKPSSFPFLQTL----NLRSIIYLCPEPYPEEN 62
P P S VE Y LP PS Q L ++RS+I +C P P+ N
Sbjct: 283 PSPLSSSVERNCYSKLLPFPSFVLNHQKLAPAVHIRSVIPVCSAPPPKPN 332
>At5g23720 unknown protein
Length = 929
Score = 26.9 bits (58), Expect = 7.2
Identities = 24/112 (21%), Positives = 49/112 (43%), Gaps = 9/112 (8%)
Query: 22 SMVEDCIYRSSLPKPSSFPFLQTLNLRSIIYLCPEPYPEENL---DFLKEQNIRLFQFGI 78
SM+++ ++ S LQ L + ++ LC + + D + QN F I
Sbjct: 705 SMIQENLFIGGGLAARSIYTLQHLGITHVLCLCANEIGQSDTQYPDLFEYQN-----FSI 759
Query: 79 EGKTEVSLPALRDSIMEALKVLVDVRNHPILVHCKQGKHRTGCLVGCFRKLQ 130
+ ++ ++ ++ +K + ILVHC +G+ R+ +V + LQ
Sbjct: 760 TDDEDSNIESIFQEALDFIKHGEETGGK-ILVHCFEGRSRSATVVLAYLMLQ 810
>At5g18590 RanGAP1 interacting protein
Length = 708
Score = 26.9 bits (58), Expect = 7.2
Identities = 13/54 (24%), Positives = 25/54 (46%), Gaps = 3/54 (5%)
Query: 11 DDDVLIPPPNFSMVEDCIYRSSLPKPSSFPFLQTLNLRSIIYLCPEPYPEENLD 64
++D+ +PPP +D + + + + + S+ C E YP EN+D
Sbjct: 486 EEDISVPPPT---TDDNTGGAKITAEKTLSMVSDREILSLQKQCSETYPLENVD 536
>At2g22510 unknown protein
Length = 124
Score = 26.9 bits (58), Expect = 7.2
Identities = 17/57 (29%), Positives = 27/57 (46%), Gaps = 4/57 (7%)
Query: 31 SSLPKPSSFPFLQTLNLRSIIYLCPEPYPEENLDFLKEQNIRLFQFGIEGKTEVSLP 87
+SLP+P++FP L + + S+ P P N+ + NI F I +LP
Sbjct: 57 TSLPQPTAFPPLPSSQIPSL----PNPAQPINIPNFPQINIPNFPISIPNNFPFNLP 109
>At1g44020 hypothetical protein
Length = 654
Score = 26.9 bits (58), Expect = 7.2
Identities = 16/53 (30%), Positives = 26/53 (48%)
Query: 22 SMVEDCIYRSSLPKPSSFPFLQTLNLRSIIYLCPEPYPEENLDFLKEQNIRLF 74
S + + R++ + +QT++L S + L +P PE L L Q I LF
Sbjct: 11 SFISQLVSRNNTDSENISCMIQTISLVSSMDLKSQPKPESKLMSLVTQTISLF 63
>At5g39400 PTEN -like protein
Length = 412
Score = 26.6 bits (57), Expect = 9.4
Identities = 12/33 (36%), Positives = 15/33 (45%)
Query: 109 LVHCKQGKHRTGCLVGCFRKLQNWCLSSAFEEY 141
+VHC GK RTG +V + A E Y
Sbjct: 149 VVHCMAGKGRTGLMVSAYLVYGGMSAEEALEMY 181
>At4g12010 like disease resistance protein (TMV N-like)
Length = 1219
Score = 26.6 bits (57), Expect = 9.4
Identities = 28/100 (28%), Positives = 42/100 (42%), Gaps = 14/100 (14%)
Query: 24 VEDCIYRSSLPKPSSFPFLQTLNLRSIIYLCPEPYPEENLDFLKEQNIRLFQFGIEGKTE 83
+ DC SLPK LQTL L L P EN++ L ++G
Sbjct: 697 LRDCTSLRSLPKGIKTQSLQTLILSGCSSLKKFPLISENVEVLL----------LDGTVI 746
Query: 84 VSLPALRDSIMEALKV-LVDVRNHPILVHCKQGKHRTGCL 122
SLP +SI ++ L++++N L H ++ CL
Sbjct: 747 KSLP---ESIQTFRRLALLNLKNCKKLKHLSSDLYKLKCL 783
>At3g09100 putative mRNA capping enzyme, RNA guanylyltransferase
Length = 672
Score = 26.6 bits (57), Expect = 9.4
Identities = 17/62 (27%), Positives = 33/62 (52%), Gaps = 5/62 (8%)
Query: 66 LKEQNIRLFQFGIEGKT----EVSLPALRDSIMEALKVLVDVRNHPILVHCKQGKHRTGC 121
LK++ I+ + +G+ VS+ A + + + + L + + ILVHC G +RTG
Sbjct: 155 LKKEGIKHVKIACKGRDAVPDNVSVNAFVNEVNQFVLNLKHSKKY-ILVHCTHGHNRTGF 213
Query: 122 LV 123
++
Sbjct: 214 MI 215
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.324 0.141 0.424
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,681,032
Number of Sequences: 26719
Number of extensions: 199685
Number of successful extensions: 512
Number of sequences better than 10.0: 19
Number of HSP's better than 10.0 without gapping: 9
Number of HSP's successfully gapped in prelim test: 10
Number of HSP's that attempted gapping in prelim test: 500
Number of HSP's gapped (non-prelim): 19
length of query: 200
length of database: 11,318,596
effective HSP length: 94
effective length of query: 106
effective length of database: 8,807,010
effective search space: 933543060
effective search space used: 933543060
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 57 (26.6 bits)
Medicago: description of AC148289.16


