

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
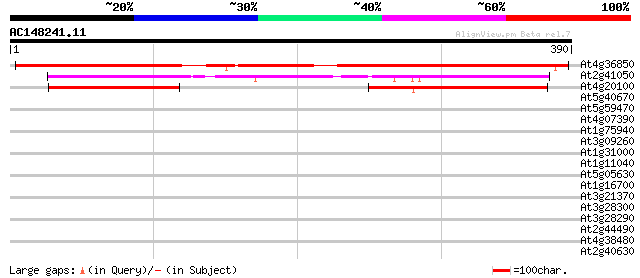
Query= AC148241.11 + phase: 0
(390 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At4g36850 unknown protein 382 e-106
At2g41050 unknown protein 215 4e-56
At4g20100 putative protein 112 3e-25
At5g40670 unknown protein 35 0.071
At5g59470 unknown protein 32 0.46
At4g07390 unknown protein 32 0.46
At1g75940 ATA27 31 1.3
At3g09260 thioglucosidase 3D precursor 30 2.3
At1g31000 F17F8.8 30 2.3
At1g11040 DnaJ isolog 30 2.3
At5g05630 unknown protein 30 3.0
At1g16700 NADH-ubiquinone oxidoreductase like protein 29 3.9
At3g21370 beta-glucosidase, putative 29 5.1
At3g28300 At14a-2 protein 28 6.6
At3g28290 At14a-1 protein 28 6.6
At2g44490 putative beta-glucosidase 28 6.6
At4g38480 unknown protein 28 8.6
At2g40630 unknown protein 28 8.6
>At4g36850 unknown protein
Length = 374
Score = 382 bits (982), Expect = e-106
Identities = 208/389 (53%), Positives = 256/389 (65%), Gaps = 37/389 (9%)
Query: 5 YCVKENKECVKWVETYFKDCLCNLRDDISFSLGLMSLVSWGVAEIPQIITIFRNKSSHGI 64
YC+KE K CV+WVE YF DCLCNL DD+SF+LG+ SL+ WGVAEIPQ+IT FR KSS+G+
Sbjct: 6 YCLKEKKTCVRWVEIYFDDCLCNLNDDVSFALGIASLLCWGVAEIPQVITNFRTKSSNGV 65
Query: 65 SLAFLLTWVAGDICNLVGCLLEPATLPTQFYTALTVINCSFMQALQSYYYCKLYTMITSS 124
SL+FLL WVAGDI NLVGCLLEPATLPTQFYTAL + + +Q+ YY +Y +
Sbjct: 66 SLSFLLAWVAGDIFNLVGCLLEPATLPTQFYTALLYTVSTVVLVIQTIYYDYIYKL---- 121
Query: 125 DGANIVRMLRVNCYFYNCIILQDNE--EEKRPLNPKPSQVYSGIAIPNGTQKEAARGEYY 182
C I Q +E EEKRPL P P + S I+IP G+ K+++R E+Y
Sbjct: 122 ------------CRHRRTKICQKDEEDEEKRPLKP-PKTMGSAISIPGGSYKDSSRREFY 168
Query: 183 YMSARSLAGSATPPSFT-HLRAAKSGPSALEFIHDSSDDDEASQVTSNISTTKPWSIPRS 241
Y SARSLAGS TPP T + R AKSG + S + + +
Sbjct: 169 YTSARSLAGSGTPPLRTSYFRVAKSGLRLWQ---------------STMMVRHRTKMRQC 213
Query: 242 VDGRYGTFLATAINLPLKGNSMRYGYIGFTGIKLLKVYVVVHSTYGQYLGWIMAAIYTCS 301
+GTFLA + +LPL+ S+ Y + +LL +V HS GQ+LGW+MAAIY
Sbjct: 214 RRAGFGTFLAASASLPLQAKSLAEKYAHASSRRLLNERIVEHSALGQWLGWLMAAIYMGG 273
Query: 302 RIPQIWLNIKRGSVEGLNPFMFVFALIANTSYVGSILVRTTEFESIKANLPWLLDATVCV 361
RIPQIWLNIKRGSVEGLNP MF+FAL+AN +YVGSILVRTTE+++IK NLPWLLDA VCV
Sbjct: 274 RIPQIWLNIKRGSVEGLNPLMFIFALVANATYVGSILVRTTEWDNIKPNLPWLLDAIVCV 333
Query: 362 ALDFFIISQYIYYRYFR--SSESSDDGEY 388
LD FII QYIYY+Y R S ES ++ Y
Sbjct: 334 VLDLFIILQYIYYKYCRIKSLESREEDAY 362
>At2g41050 unknown protein
Length = 376
Score = 215 bits (547), Expect = 4e-56
Identities = 143/367 (38%), Positives = 203/367 (54%), Gaps = 32/367 (8%)
Query: 27 NLRDDISFSLGLMSLVSWGVAEIPQIITIFRNKSSHGISLAFLLTWVAGDICNLVGCLLE 86
+ RD +S SLG++S++SWGVAEIPQI+T + KS+ G+S+ FL TW+ GDI NL+GCL+E
Sbjct: 6 SFRDGLSLSLGIISVISWGVAEIPQIMTNYSEKSTEGLSITFLTTWMIGDIFNLLGCLME 65
Query: 87 PATLPTQFYTALTVINCSFMQALQSYYYCKLYTMITSSDGANIVRMLRVNCYFYNCIILQ 146
PATLPTQFY AL + + +QS YY +Y + + +V R++ I+
Sbjct: 66 PATLPTQFYMALLYTVTTSVLYVQSIYYGHIYPRLKNRRD-QMVEAERIS------NIIS 118
Query: 147 DNEEEKRPLNPKPSQVYSGIAIP----NGTQKEAARG-EYYYMSARSLAGSATPPSFTHL 201
D + R N + G P G+Q+ + G E +Y SARSL+ S TPP+ + L
Sbjct: 119 DVKIPGRWRNSSDTTTCGGQTTPITMIPGSQRTSFTGRELFYTSARSLSSSHTPPAGSVL 178
Query: 202 RAAKSGPSALEFIHDSSDDDEASQVTSNISTTKPWSIPRSVDGRYGTFLATAINLPLKGN 261
+ + + + ++ + +TK SV GTF NL +
Sbjct: 179 AQRMARGYSEPTLEEPLLPEDVTH-----PSTKSLLCVVSVFLFLGTF--NLPNLLSESR 231
Query: 262 SMRYG-----YIGFTGIKLLKV---YVVVH-----STYGQYLGWIMAAIYTCSRIPQIWL 308
+M G ++ KLL+V V H S G +LGW MAAIY R+PQI L
Sbjct: 232 TMALGEGDRVFVVRAARKLLQVTSSNVAEHSGGESSRIGMFLGWAMAAIYMGGRLPQICL 291
Query: 309 NIKRGSVEGLNPFMFVFALIANTSYVGSILVRTTEFESIKANLPWLLDATVCVALDFFII 368
N++RG VEGLNP MF FAL+ N +YV SILV + E+ + NLPWL+DA CV LDF I+
Sbjct: 292 NMRRGHVEGLNPLMFFFALVGNMTYVASILVNSVEWLKLAPNLPWLVDAGGCVVLDFLIL 351
Query: 369 SQYIYYR 375
Q+ ++R
Sbjct: 352 LQFFHFR 358
>At4g20100 putative protein
Length = 288
Score = 112 bits (281), Expect = 3e-25
Identities = 57/128 (44%), Positives = 82/128 (63%), Gaps = 3/128 (2%)
Query: 250 LATAINLPLKGNSMRYGYIGFTGIKLLKVY---VVVHSTYGQYLGWIMAAIYTCSRIPQI 306
L+ + ++ L+G + G KLL+V + ++ G +LGW MAAIY R+PQI
Sbjct: 140 LSGSRSMDLRGKDRVFVVGGAGARKLLEVSSGNLGENNNIGMWLGWAMAAIYMGGRLPQI 199
Query: 307 WLNIKRGSVEGLNPFMFVFALIANTSYVGSILVRTTEFESIKANLPWLLDATVCVALDFF 366
+N++RG+VEGLNP MF FA I N +YV SILV + E+ I+ NLPWL+D+ C LDF
Sbjct: 200 CMNVRRGNVEGLNPLMFFFAFIGNVTYVASILVNSVEWSKIEPNLPWLVDSGGCAVLDFL 259
Query: 367 IISQYIYY 374
I+ Q+ Y+
Sbjct: 260 ILLQFFYF 267
Score = 109 bits (273), Expect = 2e-24
Identities = 49/91 (53%), Positives = 68/91 (73%)
Query: 28 LRDDISFSLGLMSLVSWGVAEIPQIITIFRNKSSHGISLAFLLTWVAGDICNLVGCLLEP 87
+RDD+S SLG++S++SW VAEIPQI+T + KS G+S+ FL TW+ GDI N+VGCL+EP
Sbjct: 2 IRDDLSLSLGIISVISWSVAEIPQIMTNYNQKSIEGVSITFLTTWMLGDIFNVVGCLMEP 61
Query: 88 ATLPTQFYTALTVINCSFMQALQSYYYCKLY 118
A+LP QFYTA+ + + +QS YY +Y
Sbjct: 62 ASLPVQFYTAVLYTLATLVLYVQSIYYGHIY 92
>At5g40670 unknown protein
Length = 270
Score = 35.0 bits (79), Expect = 0.071
Identities = 18/47 (38%), Positives = 28/47 (59%)
Query: 288 QYLGWIMAAIYTCSRIPQIWLNIKRGSVEGLNPFMFVFALIANTSYV 334
+ +GWI A ++ S PQ+ LN +R SV GLN + L ++SY+
Sbjct: 14 EIVGWIAFASWSISFYPQLILNFRRRSVVGLNFDFVMLNLTKHSSYM 60
Score = 30.0 bits (66), Expect = 2.3
Identities = 13/41 (31%), Positives = 26/41 (62%), Gaps = 1/41 (2%)
Query: 31 DISFSL-GLMSLVSWGVAEIPQIITIFRNKSSHGISLAFLL 70
+IS+ + G ++ SW ++ PQ+I FR +S G++ F++
Sbjct: 10 EISYEIVGWIAFASWSISFYPQLILNFRRRSVVGLNFDFVM 50
>At5g59470 unknown protein
Length = 148
Score = 32.3 bits (72), Expect = 0.46
Identities = 23/63 (36%), Positives = 31/63 (48%), Gaps = 3/63 (4%)
Query: 22 KDCLCNLRDDISFSLGLMSLVSWGVAEIPQIITIFRNKSSHGISLAFLLTWVAGDICNLV 81
KDCL L IS LG + + ++PQI+ I NKS G+S+ V G +L
Sbjct: 23 KDCLLPL---ISKLLGYFLVAASMTVKLPQIMKIVDNKSVKGLSVVAFELEVIGYTISLA 79
Query: 82 GCL 84
CL
Sbjct: 80 YCL 82
Score = 28.1 bits (61), Expect = 8.6
Identities = 19/77 (24%), Positives = 34/77 (43%)
Query: 290 LGWIMAAIYTCSRIPQIWLNIKRGSVEGLNPFMFVFALIANTSYVGSILVRTTEFESIKA 349
LG+ + A ++PQI + SV+GL+ F +I T + L + F +
Sbjct: 34 LGYFLVAASMTVKLPQIMKIVDNKSVKGLSVVAFELEVIGYTISLAYCLNKDLPFSAFGE 93
Query: 350 NLPWLLDATVCVALDFF 366
L+ A + VA ++
Sbjct: 94 LAFLLIQALILVACIYY 110
>At4g07390 unknown protein
Length = 235
Score = 32.3 bits (72), Expect = 0.46
Identities = 27/92 (29%), Positives = 40/92 (43%), Gaps = 3/92 (3%)
Query: 22 KDCLCNLRDDISFSLGLMSLVSWGVAEIPQIITIFRNKSSHGISLAFLLTWVAGDICNLV 81
KDCL L IS LG + + ++PQI+ I ++KS G+S+ V G +L
Sbjct: 23 KDCLLPL---ISKLLGYCLVAASITVKLPQIMKIVQHKSVRGLSVVAFELEVVGYTISLA 79
Query: 82 GCLLEPATLPTQFYTALTVINCSFMQALQSYY 113
CL + A +I + A YY
Sbjct: 80 YCLHKGLPFSAFGEMAFLLIQALILVACIYYY 111
Score = 29.6 bits (65), Expect = 3.0
Identities = 23/79 (29%), Positives = 37/79 (46%), Gaps = 7/79 (8%)
Query: 296 AIYTCSRIPQIWLNIKRGSVEGLNPFMFVFALIANTSYVGSILVRTTEFESIKANLPWLL 355
AI+ +R+PQIW N K S L+ F ++ GSI+ T + KA + L
Sbjct: 152 AIFLFARLPQIWKNFKNKSTGELSFLTFFM------NFAGSIVRVFTSLQE-KAPISILT 204
Query: 356 DATVCVALDFFIISQYIYY 374
+ V + I++Q + Y
Sbjct: 205 GFALGVVTNGSILTQILLY 223
>At1g75940 ATA27
Length = 535
Score = 30.8 bits (68), Expect = 1.3
Identities = 19/63 (30%), Positives = 31/63 (49%), Gaps = 2/63 (3%)
Query: 193 ATPPSFTHLRAAKS-GPSALEFIHDSSDDDEASQVTSNISTTKPWSIPRSVDGRYGTFLA 251
A P F H R K + +++ HD D+ A+ +T + T W P+ ++ YG FL+
Sbjct: 117 AWPRIFPHGRMEKGISKAGVQYYHDLIDELLANGITPLV-TVFHWDTPQDLEDEYGGFLS 175
Query: 252 TAI 254
I
Sbjct: 176 DRI 178
>At3g09260 thioglucosidase 3D precursor
Length = 524
Score = 30.0 bits (66), Expect = 2.3
Identities = 20/63 (31%), Positives = 30/63 (46%), Gaps = 2/63 (3%)
Query: 193 ATPPSFTHLRAAKSGPSA-LEFIHDSSDDDEASQVTSNISTTKPWSIPRSVDGRYGTFLA 251
A P F H R K A ++F HD D+ + +T + T W P+ ++ YG FL+
Sbjct: 115 AWPRIFPHGRKEKGVSQAGVQFYHDLIDELIKNGITPFV-TVFHWDTPQDLEDEYGGFLS 173
Query: 252 TAI 254
I
Sbjct: 174 ERI 176
>At1g31000 F17F8.8
Length = 365
Score = 30.0 bits (66), Expect = 2.3
Identities = 17/52 (32%), Positives = 26/52 (49%), Gaps = 2/52 (3%)
Query: 219 DDDEASQVTSNISTTKPWSIPRSVDGRYGTFLATAINLPLKGNSMRYGYIGF 270
DDD+A +T NI T + P D R+G + + LK N +R+ + F
Sbjct: 291 DDDDAFLITGNIHTGEIIFAPYDCDVRFG--MTHVVYYDLKTNRLRFRTVAF 340
>At1g11040 DnaJ isolog
Length = 438
Score = 30.0 bits (66), Expect = 2.3
Identities = 25/99 (25%), Positives = 44/99 (44%), Gaps = 7/99 (7%)
Query: 151 EKRPLNPKPSQVYSGIAIPNGTQKEAARGEYYYMSARSLAGSATPPSFTHLRAAK-SGPS 209
+ RP++ S + S P G K+ G+ + S+ ++P S + + K S PS
Sbjct: 140 KSRPISNNTSPMTSPFTSPRG--KDQTLGDLF----GSVKKLSSPTSNSPINGVKQSSPS 193
Query: 210 ALEFIHDSSDDDEASQVTSNISTTKPWSIPRSVDGRYGT 248
++ D DE +S ST+ P+S +S G+
Sbjct: 194 SISKSASKRDKDERGSTSSATSTSLPYSKSKSTRDTAGS 232
>At5g05630 unknown protein
Length = 490
Score = 29.6 bits (65), Expect = 3.0
Identities = 19/86 (22%), Positives = 40/86 (46%), Gaps = 1/86 (1%)
Query: 267 YIGFTGIKLLKVYVVVHSTYGQYLGWIMAAIYTCSRIPQIWLNIKRGSVEGLNPFMFVFA 326
Y+ + G+ ++ V V+ + +M+ + P WL + + ++G+N +++
Sbjct: 182 YLNYRGLSIVGVAAVLLGVFSILPFVVMSFMSIPKLKPSRWLVVSK-KMKGVNWSLYLNT 240
Query: 327 LIANTSYVGSILVRTTEFESIKANLP 352
L N +Y S+ T E E+ LP
Sbjct: 241 LFWNLNYWDSVSTLTGEVENPSKTLP 266
>At1g16700 NADH-ubiquinone oxidoreductase like protein
Length = 222
Score = 29.3 bits (64), Expect = 3.9
Identities = 27/85 (31%), Positives = 42/85 (48%), Gaps = 7/85 (8%)
Query: 177 ARGEYYYMSARSLAGSATPPSFTHLRAAKSGPSALEFIHDSSDDDEASQVTSNISTTKPW 236
AR + + AR LA S +HL +S A+ + + DD+EA Q+ IS K W
Sbjct: 6 ARRSFSALRARHLAFSGQGLQGSHLCGLQS--RAISY-GSNKDDEEAEQLAKEIS--KDW 60
Query: 237 S--IPRSVDGRYGTFLATAINLPLK 259
S RS++ + T + ++L LK
Sbjct: 61 STVFERSMNTLFLTEMVRGLSLTLK 85
>At3g21370 beta-glucosidase, putative
Length = 527
Score = 28.9 bits (63), Expect = 5.1
Identities = 18/61 (29%), Positives = 27/61 (43%), Gaps = 2/61 (3%)
Query: 195 PPSFTHLRAAKS-GPSALEFIHDSSDDDEASQVTSNISTTKPWSIPRSVDGRYGTFLATA 253
P F H R K ++F HD D+ + +T + T W P ++ YG FL+
Sbjct: 115 PRIFPHGRMEKGISKEGVQFYHDLIDELLKNDITPLV-TVFHWDTPADLEDEYGGFLSER 173
Query: 254 I 254
I
Sbjct: 174 I 174
>At3g28300 At14a-2 protein
Length = 385
Score = 28.5 bits (62), Expect = 6.6
Identities = 25/86 (29%), Positives = 40/86 (46%), Gaps = 7/86 (8%)
Query: 205 KSGPSALEFIHDSSD--DDEASQVTSNISTTKPWSIPRSVDGRYGTFLATAIN---LPLK 259
K S LE + ++ + D+E + IS+ K WSI +V G F+A A+ L
Sbjct: 182 KEQESLLERVRETKEKLDEELKNIEMEISSRKKWSIISNV-LFIGAFVAVAVGSMVLVCT 240
Query: 260 GNSMRYGYIGFTGIKLLKV-YVVVHS 284
G G G + L+ + +V VH+
Sbjct: 241 GVGAGVGVAGLLSLPLIAIGWVGVHT 266
>At3g28290 At14a-1 protein
Length = 385
Score = 28.5 bits (62), Expect = 6.6
Identities = 25/86 (29%), Positives = 40/86 (46%), Gaps = 7/86 (8%)
Query: 205 KSGPSALEFIHDSSD--DDEASQVTSNISTTKPWSIPRSVDGRYGTFLATAIN---LPLK 259
K S LE + ++ + D+E + IS+ K WSI +V G F+A A+ L
Sbjct: 182 KEQESLLERVRETKEKLDEELKNIEMEISSRKKWSIISNV-LFIGAFVAVAVGSMVLVCT 240
Query: 260 GNSMRYGYIGFTGIKLLKV-YVVVHS 284
G G G + L+ + +V VH+
Sbjct: 241 GVGAGVGVAGLLSLPLIAIGWVGVHT 266
>At2g44490 putative beta-glucosidase
Length = 560
Score = 28.5 bits (62), Expect = 6.6
Identities = 14/44 (31%), Positives = 26/44 (58%), Gaps = 1/44 (2%)
Query: 211 LEFIHDSSDDDEASQVTSNISTTKPWSIPRSVDGRYGTFLATAI 254
++F +D D+ A+++T + T W IP+ ++ YG FL+ I
Sbjct: 114 IKFYNDVIDELLANEITPLV-TIFHWDIPQDLEDEYGGFLSEQI 156
>At4g38480 unknown protein
Length = 471
Score = 28.1 bits (61), Expect = 8.6
Identities = 28/105 (26%), Positives = 44/105 (41%), Gaps = 16/105 (15%)
Query: 129 IVRMLRVNCYFYNCIILQDNEEEKRPLNPK--PSQVYSGIAI--PNGTQKEAARGEYYYM 184
++R + + + NCI E P P S + + I I P GT+K + G
Sbjct: 344 LLRAMEADRHVVNCI-------ESHPHMPLMCSSGIDTDIKIWTPGGTEKPLSPG----- 391
Query: 185 SARSLAGSATPPSFTHLRAAKSGPSALEFIHDSSDDDEASQVTSN 229
+A+ +G P F S E +SSDDD++S+ N
Sbjct: 392 NAKQASGFGNPRWFDGYYVDDDDDSDDESSEESSDDDDSSEEEEN 436
>At2g40630 unknown protein
Length = 535
Score = 28.1 bits (61), Expect = 8.6
Identities = 17/46 (36%), Positives = 19/46 (40%)
Query: 202 RAAKSGPSALEFIHDSSDDDEASQVTSNISTTKPWSIPRSVDGRYG 247
R SG AL H DD +V NIS KP SV + G
Sbjct: 129 RVVNSGKQALLENHTVKTDDSKCRVVKNISLLKPRETTESVVSQRG 174
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.322 0.137 0.421
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,005,173
Number of Sequences: 26719
Number of extensions: 374041
Number of successful extensions: 874
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 12
Number of HSP's that attempted gapping in prelim test: 854
Number of HSP's gapped (non-prelim): 30
length of query: 390
length of database: 11,318,596
effective HSP length: 101
effective length of query: 289
effective length of database: 8,619,977
effective search space: 2491173353
effective search space used: 2491173353
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 61 (28.1 bits)
Medicago: description of AC148241.11


