

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148241.10 - phase: 1 /pseudo
(611 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
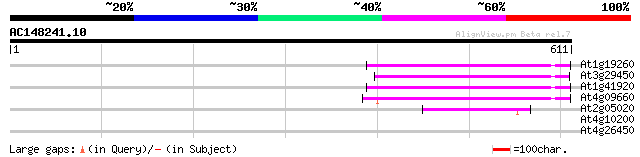
Score E
Sequences producing significant alignments: (bits) Value
At1g19260 hypothetical protein 151 9e-37
At3g29450 hypothetical protein 150 2e-36
At1g41920 hypothetical protein 149 3e-36
At4g09660 putative protein 146 4e-35
At2g05020 hypothetical protein 72 1e-12
At4g10200 putative protein 42 0.001
At4g26450 unknown protein 30 3.0
>At1g19260 hypothetical protein
Length = 769
Score = 151 bits (382), Expect = 9e-37
Identities = 76/222 (34%), Positives = 133/222 (59%), Gaps = 3/222 (1%)
Query: 389 MTASREVASIHKFFEKLIFVVNVVGSSTKRHDELQAAQSKEIENLLETGEIVTGKGKNQV 448
M +++ + +FF + ++NVVG+S KR D+++ +++E + GEI TG G NQ
Sbjct: 373 MAVAKKHVEVGEFFYMISVLLNVVGASCKRKDKIREIHRQKVEEKISNGEIKTGTGLNQE 432
Query: 449 GTVKRAGDTRWGSHFNSICSLISMYGATCPVLKIIAKDAKKIAQRADADSSYNHLKSFDF 508
+++R G+TRWGSH+ ++ L ++ + VL+ I + +R A + +FDF
Sbjct: 433 LSLQRPGNTRWGSHYKTLLRLEELFSSIVIVLEYIQDEGTDTTKRQQAYGILKYFHTFDF 492
Query: 509 IFILHLMKEIMGTTDLLCQALQKQSQDVVNAVLLVRSTKALIQDLRENGWDKLFANVVSF 568
+F L LM +MG TD L +ALQ++ QD++NA+ LV++TK +Q +R++GWD A V SF
Sbjct: 493 VFYLELMLLVMGLTDSLSKALQRKDQDILNAISLVKTTKCQLQKVRDDGWDAFMAKVSSF 552
Query: 569 CEKHDIEVPDLNDSHSTTRFGRSHLEENQVTIEHYFRVDFFY 610
EK++ + + + +R R ++ +T H+++VD FY
Sbjct: 553 SEKNNTGMLKMEEEFVDSRRPR---KKTGITNLHHYKVDCFY 591
>At3g29450 hypothetical protein
Length = 522
Score = 150 bits (380), Expect = 2e-36
Identities = 78/212 (36%), Positives = 127/212 (59%), Gaps = 3/212 (1%)
Query: 398 IHKFFEKLIFVVNVVGSSTKRHDELQAAQSKEIENLLETGEIVTGKGKNQVGTVKRAGDT 457
I FF+ + ++NVVG+S KR D ++ K++E + GEI TGKG NQ +++R G+T
Sbjct: 266 IGDFFDMISVLINVVGASCKRKDRVRDEFRKKLEERINQGEIKTGKGLNQKLSLQRPGNT 325
Query: 458 RWGSHFNSICSLISMYGATCPVLKIIAKDAKKIAQRADADSSYNHLKSFDFIFILHLMKE 517
RWG+H+ ++ L+ ++ VL+ I D +R A+ + +FDF+F L LM
Sbjct: 326 RWGTHYTTLLRLVDLFSVIIKVLEWIEDDGTDSTKRRQANGLLKYFNTFDFVFYLQLMLL 385
Query: 518 IMGTTDLLCQALQKQSQDVVNAVLLVRSTKALIQDLRENGWDKLFANVVSFCEKHDIEVP 577
I+G T+ L ALQ++ QD++NA+ LV+STK + LR++GWD V SFC+ HDIE
Sbjct: 386 ILGLTNSLSVALQRKDQDILNAMSLVKSTKQQLFKLRDDGWDSFLNEVFSFCKDHDIEFV 445
Query: 578 DLNDSHSTTRFGRSHLEENQVTIEHYFRVDFF 609
++ R R +++ +T H+++V+ F
Sbjct: 446 IMDGEFVDPRKPR---KKSNMTNLHHYQVECF 474
>At1g41920 hypothetical protein
Length = 496
Score = 149 bits (377), Expect = 3e-36
Identities = 77/222 (34%), Positives = 131/222 (58%), Gaps = 3/222 (1%)
Query: 389 MTASREVASIHKFFEKLIFVVNVVGSSTKRHDELQAAQSKEIENLLETGEIVTGKGKNQV 448
+ +++ + FF+ + ++NVVG+S K D ++ KEIE + GEI TGKG NQ
Sbjct: 198 VAVAKKQFEVGDFFDMISALLNVVGASCKGKDRIREEYLKEIEEGINQGEIKTGKGLNQE 257
Query: 449 GTVKRAGDTRWGSHFNSICSLISMYGATCPVLKIIAKDAKKIAQRADADSSYNHLKSFDF 508
+++R G+TRWG+H+ ++ L ++ +L+ + ++ +R A+ + +FDF
Sbjct: 258 LSLQRPGNTRWGTHYTTLHRLAHLFSVIIKLLEFVEEEGTDSTKRRQANGLLKYFNTFDF 317
Query: 509 IFILHLMKEIMGTTDLLCQALQKQSQDVVNAVLLVRSTKALIQDLRENGWDKLFANVVSF 568
F L LM ++G T+ L ALQK+ QD++N + LV+STK + LR++GWD L V SF
Sbjct: 318 AFYLQLMLLLLGLTNSLSVALQKKDQDILNPMSLVKSTKQQLCKLRDDGWDSLVNEVFSF 377
Query: 569 CEKHDIEVPDLNDSHSTTRFGRSHLEENQVTIEHYFRVDFFY 610
C+KHDIE+ ++ R R + + +T H+++V+ FY
Sbjct: 378 CKKHDIELVIMDGEFVNPRNPR---KRSNMTNFHHYQVECFY 416
>At4g09660 putative protein
Length = 664
Score = 146 bits (368), Expect = 4e-35
Identities = 78/232 (33%), Positives = 137/232 (58%), Gaps = 9/232 (3%)
Query: 385 EENGMTAS--REVASIH----KFFEKLIFVVNVVGSSTKRHDELQAAQSKEIENLLETGE 438
E NG+ + +E+A H +FF+ + ++NVVG+S R D+++ +++E + GE
Sbjct: 307 EFNGLRSLILKEIAKKHVEVGEFFDMISVLLNVVGASCTRKDKIREIHRQKVEEKISNGE 366
Query: 439 IVTGKGKNQVGTVKRAGDTRWGSHFNSICSLISMYGATCPVLKIIAKDAKKIAQRADADS 498
I TG NQ +++R G+TRWGSH+ ++ L ++ + VL+ I + +R A
Sbjct: 367 IKTGTRLNQELSLQRPGNTRWGSHYKTLLRLEELFSSIVIVLEYIQDEGTDTTKRQQAYG 426
Query: 499 SYNHLKSFDFIFILHLMKEIMGTTDLLCQALQKQSQDVVNAVLLVRSTKALIQDLRENGW 558
+ +FDF+F L LM +MG TD L +ALQ++ QD++N + LV++TK +Q +R++GW
Sbjct: 427 ILKYFHTFDFVFYLELMLLVMGLTDSLSKALQRKDQDILNVISLVKTTKCQLQKVRDDGW 486
Query: 559 DKLFANVVSFCEKHDIEVPDLNDSHSTTRFGRSHLEENQVTIEHYFRVDFFY 610
D A V SF EK++ + + + +R R +++ +T H+++VD FY
Sbjct: 487 DAFMAEVSSFSEKNNTAMLKMEEEFVDSRRPR---KKSGITNLHHYKVDCFY 535
>At2g05020 hypothetical protein
Length = 370
Score = 71.6 bits (174), Expect = 1e-12
Identities = 37/124 (29%), Positives = 67/124 (53%), Gaps = 6/124 (4%)
Query: 450 TVKRAGDTRWGSHFNSICSLISMYGATCPVLKIIAKDAKKIAQRADADSSYNHLKSFDFI 509
++++ +TRWG+H+ ++ +I + ++ I + +R A+ + +FDF
Sbjct: 127 SLQKPANTRWGTHYKTLLRIIEFFSCIIEYIEYIQDEDVDNIKRRQANGLLKYFHTFDFA 186
Query: 510 FILHLMKEIMGTTDLLCQALQKQSQDVVNAVLLVRSTKALIQ------DLRENGWDKLFA 563
F L LM I+G D+L QALQ++ QD++N + LV STK + D++ ++ LF
Sbjct: 187 FYLQLMLHILGFADILSQALQRRDQDILNVMSLVASTKREVDYYYNVLDMQLQAFNDLFD 246
Query: 564 NVVS 567
V S
Sbjct: 247 EVNS 250
>At4g10200 putative protein
Length = 733
Score = 42.0 bits (97), Expect = 0.001
Identities = 25/100 (25%), Positives = 52/100 (52%), Gaps = 4/100 (4%)
Query: 480 LKIIAKDAKKIAQRADAD----SSYNHLKSFDFIFILHLMKEIMGTTDLLCQALQKQSQD 535
L +A++ R+DAD S + + F+F+F + + ++ T + + +ALQ ++ D
Sbjct: 426 LDYLAENCDDPKARSDADCLATSETHRIGGFEFLFGMVIWYNLLFTMNTVSKALQSENID 485
Query: 536 VVNAVLLVRSTKALIQDLRENGWDKLFANVVSFCEKHDIE 575
+ A++ ++ + +Q+ RE G+ + A E DIE
Sbjct: 486 IELALVQLKGLVSYLQNYRETGFQEAKAEATLIAESMDIE 525
>At4g26450 unknown protein
Length = 489
Score = 30.4 bits (67), Expect = 3.0
Identities = 17/58 (29%), Positives = 31/58 (53%)
Query: 529 LQKQSQDVVNAVLLVRSTKALIQDLRENGWDKLFANVVSFCEKHDIEVPDLNDSHSTT 586
++++SQ V +V +++S+ L +G+D F V F D+E+ D S S+T
Sbjct: 125 VEQESQGSVGSVNMLKSSGDGFDILGSSGYDSRFVAGVGFSAGMDLEIDDDRSSKSST 182
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.359 0.157 0.535
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,742,056
Number of Sequences: 26719
Number of extensions: 377803
Number of successful extensions: 2091
Number of sequences better than 10.0: 7
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 2082
Number of HSP's gapped (non-prelim): 7
length of query: 611
length of database: 11,318,596
effective HSP length: 105
effective length of query: 506
effective length of database: 8,513,101
effective search space: 4307629106
effective search space used: 4307629106
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 63 (28.9 bits)
Medicago: description of AC148241.10


