

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148236.3 + phase: 0 /pseudo
(116 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
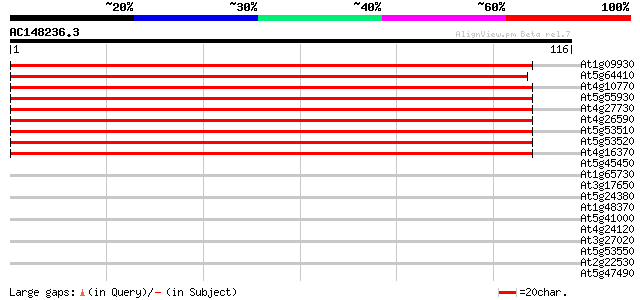
Score E
Sequences producing significant alignments: (bits) Value
At1g09930 unknown protein 192 2e-50
At5g64410 Isp4-like protein 192 3e-50
At4g10770 oligopeptide transporter like protein 163 1e-41
At5g55930 sexual differentiation process protein ISP4-like 162 3e-41
At4g27730 unknown protein 159 2e-40
At4g26590 isp4 like protein 153 2e-38
At5g53510 isp4 protein 152 2e-38
At5g53520 isp4 protein 150 1e-37
At4g16370 isp4 like protein 146 1e-36
At5g45450 putative protein 36 0.003
At1g65730 unknown protein 35 0.008
At3g17650 unknown protein 34 0.010
At5g24380 putative protein 34 0.013
At1g48370 unknown protein 34 0.013
At5g41000 unknown protein 33 0.018
At4g24120 unknown protein 32 0.051
At3g27020 unknown protein 32 0.051
At5g53550 EspB-like protein 31 0.087
At2g22530 unknown protein 30 0.15
At5g47490 putative protein 27 1.3
>At1g09930 unknown protein
Length = 734
Score = 192 bits (488), Expect = 2e-50
Identities = 87/108 (80%), Positives = 102/108 (93%)
Query: 1 MPWWALIFASGLALIFTLPVAIITATTNQTPGLNVITEYIMGVILPGRPIANVCFKTYGY 60
MPWW L+ AS +AL FT+PV+IITATTNQTPGLN+ITEY+MGV+LPGRPIANVCFKTYGY
Sbjct: 440 MPWWGLLLASFMALTFTVPVSIITATTNQTPGLNIITEYLMGVLLPGRPIANVCFKTYGY 499
Query: 61 MSMSQAISFLSDFKLGHYMKIPPRSMFIVQILGTLIAGTVDVGVAWWL 108
+SMSQAISFL+DFKLGHYMKIPPRSMF+VQ +GT+IAGTV++ VAW+L
Sbjct: 500 ISMSQAISFLNDFKLGHYMKIPPRSMFLVQFIGTVIAGTVNISVAWYL 547
>At5g64410 Isp4-like protein
Length = 729
Score = 192 bits (487), Expect = 3e-50
Identities = 82/107 (76%), Positives = 99/107 (91%)
Query: 1 MPWWALIFASGLALIFTLPVAIITATTNQTPGLNVITEYIMGVILPGRPIANVCFKTYGY 60
MPWW L+FAS +A +FTLP++IITATTNQTPGLN+ITEY MG+I PGRPIANVCFK YGY
Sbjct: 436 MPWWGLVFASAMAFVFTLPISIITATTNQTPGLNIITEYAMGLIYPGRPIANVCFKVYGY 495
Query: 61 MSMSQAISFLSDFKLGHYMKIPPRSMFIVQILGTLIAGTVDVGVAWW 107
MSM+QA+SFL+DFKLGHYMKIPPRSMF+VQ +GT++AGT+++ VAWW
Sbjct: 496 MSMAQAVSFLNDFKLGHYMKIPPRSMFLVQFIGTILAGTINITVAWW 542
>At4g10770 oligopeptide transporter like protein
Length = 766
Score = 163 bits (412), Expect = 1e-41
Identities = 72/108 (66%), Positives = 89/108 (81%)
Query: 1 MPWWALIFASGLALIFTLPVAIITATTNQTPGLNVITEYIMGVILPGRPIANVCFKTYGY 60
+PWW ++ A +A+IFTLP+ IITA TNQ PGLN+ITEYI+G I PG P+AN+CFK YGY
Sbjct: 475 LPWWGVLLACTVAIIFTLPIGIITAITNQAPGLNIITEYIIGYIYPGYPVANMCFKVYGY 534
Query: 61 MSMSQAISFLSDFKLGHYMKIPPRSMFIVQILGTLIAGTVDVGVAWWL 108
+SM QAI+FL DFKLGHYMKIPPR+MF+ QI+GTLI+ V + AWWL
Sbjct: 535 ISMQQAITFLQDFKLGHYMKIPPRTMFMAQIVGTLISCFVYLTTAWWL 582
>At5g55930 sexual differentiation process protein ISP4-like
Length = 755
Score = 162 bits (409), Expect = 3e-41
Identities = 72/108 (66%), Positives = 87/108 (79%)
Query: 1 MPWWALIFASGLALIFTLPVAIITATTNQTPGLNVITEYIMGVILPGRPIANVCFKTYGY 60
+PWW LI A +AL FTLP+ +I ATTNQ GLNVITE I+G + PG+P+ANV FKTYGY
Sbjct: 460 LPWWGLILACAIALFFTLPIGVIQATTNQQMGLNVITELIIGYLYPGKPLANVAFKTYGY 519
Query: 61 MSMSQAISFLSDFKLGHYMKIPPRSMFIVQILGTLIAGTVDVGVAWWL 108
+SMSQA+ F+ DFKLGHYMKIPPRSMFIVQ++ T++A TV G WWL
Sbjct: 520 ISMSQALYFVGDFKLGHYMKIPPRSMFIVQLVATVVASTVCFGTTWWL 567
>At4g27730 unknown protein
Length = 736
Score = 159 bits (402), Expect = 2e-40
Identities = 67/108 (62%), Positives = 87/108 (80%)
Query: 1 MPWWALIFASGLALIFTLPVAIITATTNQTPGLNVITEYIMGVILPGRPIANVCFKTYGY 60
+PWW ++ A +A+ FT + +I ATTNQ PGLN+ITEY++G I P RP+AN+CFK YGY
Sbjct: 441 LPWWGVLLACAIAISFTPLIGVIAATTNQAPGLNIITEYVIGYIYPERPVANMCFKVYGY 500
Query: 61 MSMSQAISFLSDFKLGHYMKIPPRSMFIVQILGTLIAGTVDVGVAWWL 108
+SM+QA++F+SDFKLGHYMKIPPRSMF+ Q+ GTL+A V G AWWL
Sbjct: 501 ISMTQALTFISDFKLGHYMKIPPRSMFMAQVAGTLVAVVVYTGTAWWL 548
>At4g26590 isp4 like protein
Length = 753
Score = 153 bits (386), Expect = 2e-38
Identities = 66/108 (61%), Positives = 85/108 (78%)
Query: 1 MPWWALIFASGLALIFTLPVAIITATTNQTPGLNVITEYIMGVILPGRPIANVCFKTYGY 60
+PWW L+ A +A FTLP+ +I ATTNQ GLNVI+E I+G + PG+P+ANV FKTYG
Sbjct: 458 LPWWGLLLACAIAFTFTLPIGVILATTNQRMGLNVISELIIGFLYPGKPLANVAFKTYGS 517
Query: 61 MSMSQAISFLSDFKLGHYMKIPPRSMFIVQILGTLIAGTVDVGVAWWL 108
+S++QA+ F+ DFKLGHYMKIPPRSMFIVQ++ T++A TV G WWL
Sbjct: 518 VSIAQALYFVGDFKLGHYMKIPPRSMFIVQLVATIVASTVSFGTTWWL 565
>At5g53510 isp4 protein
Length = 741
Score = 152 bits (385), Expect = 2e-38
Identities = 64/108 (59%), Positives = 86/108 (79%)
Query: 1 MPWWALIFASGLALIFTLPVAIITATTNQTPGLNVITEYIMGVILPGRPIANVCFKTYGY 60
+PWW ++ A +A+ FT + +I ATTNQ PGLNVITEY++G + P RP+AN+CFK YGY
Sbjct: 446 LPWWGVLLACAIAVFFTPLIGVIAATTNQEPGLNVITEYVIGYLYPERPVANMCFKVYGY 505
Query: 61 MSMSQAISFLSDFKLGHYMKIPPRSMFIVQILGTLIAGTVDVGVAWWL 108
+SM+QA++F+ DFKLG YMKIPPRSMF+ Q++GTL++ V G AWWL
Sbjct: 506 ISMTQALTFIQDFKLGLYMKIPPRSMFMAQVVGTLVSVVVYTGTAWWL 553
>At5g53520 isp4 protein
Length = 733
Score = 150 bits (379), Expect = 1e-37
Identities = 66/108 (61%), Positives = 83/108 (76%)
Query: 1 MPWWALIFASGLALIFTLPVAIITATTNQTPGLNVITEYIMGVILPGRPIANVCFKTYGY 60
+PWW A +A+ FT V +I ATTNQ PGLN+ITEYI+G P RP+AN+CFKTYGY
Sbjct: 440 LPWWGAFLACLIAIFFTPLVGVIMATTNQAPGLNIITEYIIGYAYPERPVANICFKTYGY 499
Query: 61 MSMSQAISFLSDFKLGHYMKIPPRSMFIVQILGTLIAGTVDVGVAWWL 108
+SMSQ+++FLSD KLG YMKIPPR+MF+ Q++GTL+A G AWWL
Sbjct: 500 ISMSQSLTFLSDLKLGTYMKIPPRTMFMAQVVGTLVAVIAYAGTAWWL 547
>At4g16370 isp4 like protein
Length = 637
Score = 146 bits (369), Expect = 1e-36
Identities = 62/108 (57%), Positives = 85/108 (78%)
Query: 1 MPWWALIFASGLALIFTLPVAIITATTNQTPGLNVITEYIMGVILPGRPIANVCFKTYGY 60
+PWW ++FA LA I TLP+ +I ATTNQ PG ++I ++I+G ILPG+PIAN+ FK YG
Sbjct: 368 LPWWGMLFAFALAFIVTLPIGVIQATTNQQPGYDIIGQFIIGYILPGKPIANLIFKIYGR 427
Query: 61 MSMSQAISFLSDFKLGHYMKIPPRSMFIVQILGTLIAGTVDVGVAWWL 108
+S A+SFL+D KLGHYMKIPP M+ Q++GT++AG V++GVAWW+
Sbjct: 428 ISTVHALSFLADLKLGHYMKIPPPCMYTAQLVGTVVAGVVNLGVAWWM 475
>At5g45450 putative protein
Length = 216
Score = 36.2 bits (82), Expect = 0.003
Identities = 19/52 (36%), Positives = 28/52 (53%), Gaps = 2/52 (3%)
Query: 57 TYGYMS--MSQAISFLSDFKLGHYMKIPPRSMFIVQILGTLIAGTVDVGVAW 106
T G+M +S+A DFK G+ P+SMF+ Q++GT + V V W
Sbjct: 7 TCGFMMNIVSRASDLTQDFKTGYLTLSSPKSMFVSQVIGTAMGCVVSPCVFW 58
>At1g65730 unknown protein
Length = 688
Score = 34.7 bits (78), Expect = 0.008
Identities = 16/44 (36%), Positives = 23/44 (51%)
Query: 63 MSQAISFLSDFKLGHYMKIPPRSMFIVQILGTLIAGTVDVGVAW 106
+S A + DFK G+ PRSMF+ Q +GT + + V W
Sbjct: 482 VSTASDLMQDFKTGYMTLASPRSMFLSQAIGTAMGCVISPCVFW 525
Score = 25.4 bits (54), Expect = 4.8
Identities = 13/31 (41%), Positives = 18/31 (57%)
Query: 5 ALIFASGLALIFTLPVAIITATTNQTPGLNV 35
ALI + LA++FT V + TT P LN+
Sbjct: 48 ALIVSFILAILFTFVVMKLNLTTGIIPSLNI 78
>At3g17650 unknown protein
Length = 714
Score = 34.3 bits (77), Expect = 0.010
Identities = 16/44 (36%), Positives = 23/44 (51%)
Query: 63 MSQAISFLSDFKLGHYMKIPPRSMFIVQILGTLIAGTVDVGVAW 106
+S A DFK G+ P+SMF+ Q++GT + V V W
Sbjct: 507 VSTASDLTQDFKTGYLTLSSPKSMFVSQVIGTAMGCVVSPCVFW 550
>At5g24380 putative protein
Length = 652
Score = 33.9 bits (76), Expect = 0.013
Identities = 15/38 (39%), Positives = 21/38 (54%)
Query: 63 MSQAISFLSDFKLGHYMKIPPRSMFIVQILGTLIAGTV 100
+S + + DFK GH + PRSM + Q +GT I V
Sbjct: 459 VSVSADLMHDFKTGHLTQTSPRSMLVAQAIGTAIGCVV 496
>At1g48370 unknown protein
Length = 724
Score = 33.9 bits (76), Expect = 0.013
Identities = 16/44 (36%), Positives = 23/44 (51%)
Query: 63 MSQAISFLSDFKLGHYMKIPPRSMFIVQILGTLIAGTVDVGVAW 106
+S A DFK G+ PR+MF+ Q++GT + V V W
Sbjct: 517 VSTASDLTQDFKTGYLTLSSPRAMFVSQVIGTAMGCLVSPCVFW 560
>At5g41000 unknown protein
Length = 670
Score = 33.5 bits (75), Expect = 0.018
Identities = 16/46 (34%), Positives = 26/46 (55%), Gaps = 1/46 (2%)
Query: 63 MSQAISFLSDFKLGHYMKIPPRSMFIVQILGTLIAGTVDVGVAWWL 108
+S A + DFK G+ +SMF+ Q+LGT + G + + +WL
Sbjct: 478 VSTAADLMQDFKTGYLTLSSAKSMFVTQLLGTAM-GCIIAPLTFWL 522
>At4g24120 unknown protein
Length = 665
Score = 32.0 bits (71), Expect = 0.051
Identities = 12/34 (35%), Positives = 21/34 (61%)
Query: 63 MSQAISFLSDFKLGHYMKIPPRSMFIVQILGTLI 96
+S + + DFK HY P++MF Q++GT++
Sbjct: 473 VSVSCILMQDFKTAHYTMTSPKAMFASQMIGTVV 506
Score = 24.6 bits (52), Expect = 8.1
Identities = 12/51 (23%), Positives = 27/51 (52%), Gaps = 1/51 (1%)
Query: 53 VCFKTYGYMSMSQAISFLSDFKLGHYMKIPPRSMFIVQILGTLIAGTVDVG 103
+C+ +G+ + + L+ K+G +M + P +M + ++G A + VG
Sbjct: 560 MCYGFFGFAVLVNVVRDLTPAKIGRFMPL-PTAMAVPFLVGAYFAIDMCVG 609
>At3g27020 unknown protein
Length = 676
Score = 32.0 bits (71), Expect = 0.051
Identities = 16/46 (34%), Positives = 26/46 (55%), Gaps = 1/46 (2%)
Query: 63 MSQAISFLSDFKLGHYMKIPPRSMFIVQILGTLIAGTVDVGVAWWL 108
+S A + DFK G+ +SMF+ Q++GT + G V + +WL
Sbjct: 481 VSTAADLMQDFKTGYLTLSSAKSMFVSQLVGTAM-GCVIAPLTFWL 525
>At5g53550 EspB-like protein
Length = 669
Score = 31.2 bits (69), Expect = 0.087
Identities = 15/38 (39%), Positives = 20/38 (52%)
Query: 63 MSQAISFLSDFKLGHYMKIPPRSMFIVQILGTLIAGTV 100
+S + + DFK GH PRSM + Q +GT I V
Sbjct: 473 VSISSDLMHDFKTGHLTLTSPRSMLVSQAIGTAIGCVV 510
>At2g22530 unknown protein
Length = 897
Score = 30.4 bits (67), Expect = 0.15
Identities = 21/60 (35%), Positives = 30/60 (50%), Gaps = 9/60 (15%)
Query: 55 FKTYGYMSMSQAISFLSDFKLGHYMKIPPRSMFIVQILGTLIAGTVDVGVAW-WLPRFNK 113
FKT +S+ ++ L DFK G S+F + I G L+ G GV W +LP +K
Sbjct: 557 FKTAKSFKISKGMNILRDFKFG--------SIFSLLISGRLLRGWHQGGVNWTYLPDISK 608
>At5g47490 putative protein
Length = 1361
Score = 27.3 bits (59), Expect = 1.3
Identities = 13/66 (19%), Positives = 28/66 (41%)
Query: 19 PVAIITATTNQTPGLNVITEYIMGVILPGRPIANVCFKTYGYMSMSQAISFLSDFKLGHY 78
P +I A Q ++ + + + ++PG P+ +C G + + SD
Sbjct: 723 PALVIAAQLGQQFYVDTVKQMALRQLVPGSPLRTLCLLVAGQPAEVFSTGSTSDISFPGS 782
Query: 79 MKIPPR 84
+ +PP+
Sbjct: 783 VNLPPQ 788
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.328 0.142 0.459
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,427,000
Number of Sequences: 26719
Number of extensions: 83144
Number of successful extensions: 274
Number of sequences better than 10.0: 29
Number of HSP's better than 10.0 without gapping: 23
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 248
Number of HSP's gapped (non-prelim): 32
length of query: 116
length of database: 11,318,596
effective HSP length: 92
effective length of query: 24
effective length of database: 8,860,448
effective search space: 212650752
effective search space used: 212650752
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 52 (24.6 bits)
Medicago: description of AC148236.3


