

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
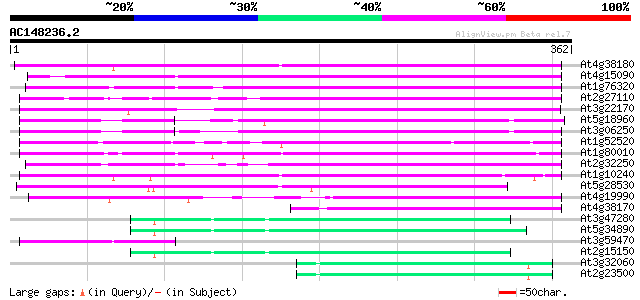
Query= AC148236.2 - phase: 0 /pseudo
(362 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At4g38180 unknown protein (At4g38180) 244 7e-65
At4g15090 unknown protein 202 2e-52
At1g76320 putative phytochrome A signaling protein 173 1e-43
At2g27110 Mutator-like transposase 169 2e-42
At3g22170 far-red impaired response protein, putative 167 1e-41
At5g18960 FAR1 - like protein 165 3e-41
At3g06250 unknown protein 164 7e-41
At1g52520 F6D8.26 149 3e-36
At1g80010 hypothetical protein 142 2e-34
At2g32250 Mutator-like transposase 139 2e-33
At1g10240 unknown protein 139 2e-33
At5g28530 far-red impaired response protein (FAR1) - like 127 9e-30
At4g19990 putative protein 118 4e-27
At4g38170 hypothetical protein 106 2e-23
At3g47280 putative protein 67 2e-11
At5g34890 putative protein 65 4e-11
At3g59470 unknown protein 65 6e-11
At2g15150 putative transposase of FARE2.2 (CDS1) 63 2e-10
At3g32060 hypothetical protein 62 6e-10
At2g23500 Mutator-like transposase 62 6e-10
>At4g38180 unknown protein (At4g38180)
Length = 788
Score = 244 bits (622), Expect = 7e-65
Identities = 129/358 (36%), Positives = 201/358 (56%), Gaps = 6/358 (1%)
Query: 4 DWNPKVDMFFCSLKNAWEFWNTYXGKXGFGVRKDYTHR-KKDGSVSSCRFVCCKEGMRKP 62
D P + F S + A F+N+Y + GF R + R ++DG++ +FVC KEG R
Sbjct: 70 DLEPYDGLEFESEEAAKAFYNSYARRIGFSTRVSSSRRSRRDGAIIQRQFVCAKEGFRNM 129
Query: 63 DKR---DYKTKNPRLETRTNCEARLGLKNVD-GKLMVHDFVEDHNHELLLPETTHMLSSQ 118
+++ D + K PR TR C+A L +K D GK +V FV+DHNHEL+ P+ H L S
Sbjct: 130 NEKRTKDREIKRPRTITRVGCKASLSVKMQDSGKWLVSGFVKDHNHELVPPDQVHCLRSH 189
Query: 119 RKVSEIHCQQIELADSVGVQQKKSFDLLSKEVGGRTNLGFIRLDQKNYLRKKRERSLVQG 178
R++S I+ + G+ ++ L KE GG + +GF +D +NY+R R++S ++G
Sbjct: 190 RQISGPAKTLIDTLQAAGMGPRRIMSALIKEYGGISKVGFTEVDCRNYMRNNRQKS-IEG 248
Query: 179 EAGYLLQYFQRKTVENPTFYHAYQLDIEDQITNVFWADARMLVDYSYFGDVVSLDSTYCT 238
E LL Y ++ +NP F+++ Q + + NVFWAD + ++D+++FGD V+ D+TY +
Sbjct: 249 EIQLLLDYLRQMNADNPNFFYSVQGSEDQSVGNVFWADPKAIMDFTHFGDTVTFDTTYRS 308
Query: 239 NSSHRPLAIISGFNHHRGAVIFGAALLYDETAESYEWLFETFLEAHKQKMPQTVFTDQAK 298
N P A +G NHH ++FG A + +ET S+ WLF T+L A P ++ TD
Sbjct: 309 NRYRLPFAPFTGVNHHGQPILFGCAFIINETEASFVWLFNTWLAAMSAHPPVSITTDHDA 368
Query: 299 SMAKALAEVMPEAYHGLCTWHLMQNGIKHLGNLMKGESYFLSDFEKCMYGYEDVEQFE 356
+ A+ V P A H C WH+++ + L ++ F SDF KC+ E VE FE
Sbjct: 369 VIRAAIMHVFPGARHRFCKWHILKKCQEKLSHVFLKHPSFESDFHKCVNLTESVEDFE 426
>At4g15090 unknown protein
Length = 768
Score = 202 bits (515), Expect = 2e-52
Identities = 109/347 (31%), Positives = 187/347 (53%), Gaps = 12/347 (3%)
Query: 12 FFCSLKNAWEFWNTYXGKXGFGVRKDYTHRKKDGSVSSCRFVCCKEGMRKPDKRDYKTKN 71
F +++N W F + K+ KK +F C + G+ + +
Sbjct: 7 FIRNMQNLWVFTTSI---------KNSRRSKKTKDFIDAKFACSRYGVTPESESSGSSSR 57
Query: 72 PRLETRTNCEARLGLKN-VDGKLMVHDFVEDHNHELLLPETTHMLSSQRKVSEIHCQQIE 130
+T+C+A + +K DGK ++H+FV+DHNHELL P + QR V I+
Sbjct: 58 RSTVKKTDCKASMHVKRRPDGKWIIHEFVKDHNHELL-PALAYHFRIQRNVKLAEKNNID 116
Query: 131 LADSVGVQQKKSFDLLSKEVGGRTNLG-FIRLDQKNYLRKKRERSLVQGEAGYLLQYFQR 189
+ +V + KK + +S++ GG N+G ++ D + + K R +L +G++ LL+YF+R
Sbjct: 117 ILHAVSERTKKMYVEMSRQSGGYKNIGSLLQTDVSSQVDKGRYLALEEGDSQVLLEYFKR 176
Query: 190 KTVENPTFYHAYQLDIEDQITNVFWADARMLVDYSYFGDVVSLDSTYCTNSSHRPLAIIS 249
ENP F++A L+ + ++ N+FWADA+ DY F DVVS D+TY + PLA+
Sbjct: 177 IKKENPKFFYAIDLNEDQRLRNLFWADAKSRDDYLSFNDVVSFDTTYVKFNDKLPLALFI 236
Query: 250 GFNHHRGAVIFGAALLYDETAESYEWLFETFLEAHKQKMPQTVFTDQAKSMAKALAEVMP 309
G NHH ++ G AL+ DE+ E++ WL +T+L A + P+ + TDQ K + A++E++P
Sbjct: 237 GVNHHSQPMLLGCALVADESMETFVWLIKTWLRAMGGRAPKVILTDQDKFLMSAVSELLP 296
Query: 310 EAYHGLCTWHLMQNGIKHLGNLMKGESYFLSDFEKCMYGYEDVEQFE 356
H WH+++ ++ ++MK FL F KC++ ++F+
Sbjct: 297 NTRHCFALWHVLEKIPEYFSHVMKRHENFLLKFNKCIFRSWTDDEFD 343
>At1g76320 putative phytochrome A signaling protein
Length = 670
Score = 173 bits (439), Expect = 1e-43
Identities = 108/349 (30%), Positives = 169/349 (47%), Gaps = 13/349 (3%)
Query: 11 MFFCSLKNAWEFWNTYXGKXGFGVRKDYTHRKKDGS-VSSCRFVCCKEGMRKPDKRDYKT 69
M F + ++A+ F+ Y GFG K + R + +F C + G ++
Sbjct: 1 MEFETHEDAYLFYKDYAKSVGFGTAKLSSRRSRASKEFIDAKFSCIRYGSKQQSD---DA 57
Query: 70 KNPRLETRTNCEARLGLKN-VDGKLMVHDFVEDHNHELLLPETTHMLSSQRKVSEIHCQQ 128
NPR + C+A + +K DGK V+ FV++HNH+LL PE H S R +
Sbjct: 58 INPRASPKIGCKASMHVKRRPDGKWYVYSFVKEHNHDLL-PEQAHYFRSHRNTELVKSND 116
Query: 129 IELADSVGVQQKKSFDLLS-KEVGGRTNLGFIRLDQKNYLRKKRERSLVQGEAGYLLQYF 187
L ++KK+ L K + +L FI +N K R L G+A LL++
Sbjct: 117 SRL------RRKKNTPLTDCKHLSAYHDLDFIDGYMRNQHDKGRRLVLDTGDAEILLEFL 170
Query: 188 QRKTVENPTFYHAYQLDIEDQITNVFWADARMLVDYSYFGDVVSLDSTYCTNSSHRPLAI 247
R ENP F+ A + + NVFW DA+ + DY F DVVS +++Y + PL +
Sbjct: 171 MRMQEENPKFFFAVDFSEDHLLRNVFWVDAKGIEDYKSFSDVVSFETSYFVSKYKVPLVL 230
Query: 248 ISGFNHHRGAVIFGAALLYDETAESYEWLFETFLEAHKQKMPQTVFTDQAKSMAKALAEV 307
G NHH V+ G LL D+T +Y WL +++L A + P+ + TDQ ++ A+A V
Sbjct: 231 FVGVNHHVQPVLLGCGLLADDTVYTYVWLMQSWLVAMGGQKPKVMLTDQNNAIKAAIAAV 290
Query: 308 MPEAYHGLCTWHLMQNGIKHLGNLMKGESYFLSDFEKCMYGYEDVEQFE 356
+PE H C WH++ ++L + F+ KC+Y E+F+
Sbjct: 291 LPETRHCYCLWHVLDQLPRNLDYWSMWQDTFMKKLFKCIYRSWSEEEFD 339
>At2g27110 Mutator-like transposase
Length = 851
Score = 169 bits (428), Expect = 2e-42
Identities = 112/350 (32%), Positives = 172/350 (49%), Gaps = 25/350 (7%)
Query: 7 PKVDMFFCSLKNAWEFWNTYXGKXGFGVRKDYTHRKKDGSVSSCRFVCCKEGMRKPDKRD 66
P V M F S K A F++ Y + GF + + DGSVS FVC R KR
Sbjct: 49 PCVGMEFNSEKEAKSFYDEYSRQLGFTSK---LLPRTDGSVSVREFVCSSSSKRS--KR- 102
Query: 67 YKTKNPRLETRTNCEARLGLKNVDGKLMVHDFVEDHNHELLLPETTHMLSSQRKVSEIHC 126
RL + R+ L+ + K +V FV++H H L H L +R +
Sbjct: 103 ------RLSESCDAMVRIELQGHE-KWVVTKFVKEHTHGLASSNMLHCLRPRRHFANSEK 155
Query: 127 QQIELADSVGVQQKKSFDLLSKEVGGRTNLGFIRLDQKNYLRKKRERSLVQGEAGYLLQY 186
+ GV +S + R +N + + +A LL+Y
Sbjct: 156 SSYQ----EGVNVPSGMMYVSMDANSR--------GARNASMATNTKRTIGRDAHNLLEY 203
Query: 187 FQRKTVENPTFYHAYQLDIEDQITNVFWADARMLVDYSYFGDVVSLDSTYCTNSSHRPLA 246
F+R ENP F++A QLD ++Q++NVFWAD+R V Y++FGD V+LD+ Y N P A
Sbjct: 204 FKRMQAENPGFFYAVQLDEDNQMSNVFWADSRSRVAYTHFGDTVTLDTRYRCNQFRVPFA 263
Query: 247 IISGFNHHRGAVIFGAALLYDETAESYEWLFETFLEAHKQKMPQTVFTDQAKSMAKALAE 306
+G NHH A++FG AL+ DE+ S+ WLF+TFL A + + P ++ TDQ +++ A +
Sbjct: 264 PFTGVNHHGQAILFGCALILDESDTSFIWLFKTFLTAMRDQPPVSLVTDQDRAIQIAAGQ 323
Query: 307 VMPEAYHGLCTWHLMQNGIKHLGNLMKGESYFLSDFEKCMYGYEDVEQFE 356
V P A H + W +++ G + L ++ F + C+ E +E+FE
Sbjct: 324 VFPGARHCINKWDVLREGQEKLAHVCLAYPSFQVELYNCINFTETIEEFE 373
>At3g22170 far-red impaired response protein, putative
Length = 814
Score = 167 bits (422), Expect = 1e-41
Identities = 106/364 (29%), Positives = 164/364 (44%), Gaps = 38/364 (10%)
Query: 7 PKVDMFFCSLKNAWEFWNTYXGKXGFGVRKDYTHRKKDG-SVSSCRFVCCKEGMRKPDKR 65
P M F S A+ F+ Y GF + R K +F C + G ++ +
Sbjct: 70 PLNGMEFESHGEAYSFYQEYSRAMGFNTAIQNSRRSKTTREFIDAKFACSRYGTKREYDK 129
Query: 66 DYKTKNPRLE-------------TRTNCEARLGLKN-VDGKLMVHDFVEDHNHELLLPET 111
+ R +T+C+A + +K DGK ++H FV +HNHELL
Sbjct: 130 SFNRPRARQSKQDPENMAGRRTCAKTDCKASMHVKRRPDGKWVIHSFVREHNHELLP--- 186
Query: 112 THMLSSQRKVSEIHCQQIELADSVGVQQKKSFDLLSKEVGGRTNLGFIRLDQKNYLRKKR 171
A +V Q +K + ++K+ + ++ D K+ K R
Sbjct: 187 --------------------AQAVSEQTRKIYAAMAKQFAEYKTVISLKSDSKSSFEKGR 226
Query: 172 ERSLVQGEAGYLLQYFQRKTVENPTFYHAYQLDIEDQITNVFWADARMLVDYSYFGDVVS 231
S+ G+ LL + R N F++A L + ++ NVFW DA+ +Y F DVVS
Sbjct: 227 TLSVETGDFKILLDFLSRMQSLNSNFFYAVDLGDDQRVKNVFWVDAKSRHNYGSFCDVVS 286
Query: 232 LDSTYCTNSSHRPLAIISGFNHHRGAVIFGAALLYDETAESYEWLFETFLEAHKQKMPQT 291
LD+TY N PLAI G N H ++ G AL+ DE+A +Y WL ET+L A + P+
Sbjct: 287 LDTTYVRNKYKMPLAIFVGVNQHYQYMVLGCALISDESAATYSWLMETWLRAIGGQAPKV 346
Query: 292 VFTDQAKSMAKALAEVMPEAYHGLCTWHLMQNGIKHLGNLMKGESYFLSDFEKCMYGYED 351
+ T+ M + E+ P H L WH++ ++LG ++K F+ FEKC+Y
Sbjct: 347 LITELDVVMNSIVPEIFPNTRHCLFLWHVLMKVSENLGQVVKQHDNFMPKFEKCIYKSGK 406
Query: 352 VEQF 355
E F
Sbjct: 407 DEDF 410
>At5g18960 FAR1 - like protein
Length = 788
Score = 165 bits (418), Expect = 3e-41
Identities = 112/357 (31%), Positives = 171/357 (47%), Gaps = 47/357 (13%)
Query: 7 PKVDMFFCSLKNAWEFWNTYXGKXGFGVRKDYTHRKK-DGSVSSCRFVCCKEGMRKPDKR 65
P + F S A +F+ Y GF VR R K DGS++S RFVC +EG + P
Sbjct: 211 PYAGLEFGSANEACQFYQAYAEVVGFRVRIGQLFRSKVDGSITSRRFVCSREGFQHP--- 267
Query: 66 DYKTKNPRLETRTNCEARLGLKNVD-GKLMVHDFVEDHNHELLLPETTHMLSSQRKVSEI 124
+R C A + +K D G +V +DHNH+L
Sbjct: 268 ----------SRMGCGAYMRIKRQDSGGWIVDRLNKDHNHDL------------------ 299
Query: 125 HCQQIELADSVGVQQKKSFDLLSKEVGGRTNLGFIRLDQ--KNYLRKKRERSLVQGEAGY 182
E KK D GG ++ I L+ N+++K RE + +
Sbjct: 300 -----EPGKKNDAGMKKIPD---DGTGGLDSVDLIELNDFGNNHIKKTRENRIGKEWYPL 351
Query: 183 LLQYFQRKTVENPTFYHAYQLDIED-QITNVFWADARMLVDYSYFGDVVSLDSTYCTNSS 241
LL YFQ + E+ F++A +LD+ + ++FWAD+R S FGD V D++Y S
Sbjct: 352 LLDYFQSRQTEDMGFFYAVELDVNNGSCMSIFWADSRARFACSQFGDSVVFDTSYRKGSY 411
Query: 242 HRPLAIISGFNHHRGAVIFGAALLYDETAESYEWLFETFLEAHKQKMPQTVFTDQAKSMA 301
P A I GFNHHR V+ G A++ DE+ E++ WLF+T+L A + P+++ DQ +
Sbjct: 412 SVPFATIIGFNHHRQPVLLGCAMVADESKEAFLWLFQTWLRAMSGRRPRSIVADQDLPIQ 471
Query: 302 KALAEVMPEAYHGLCTWHLMQNGIKHLGNLMKGESYFLSDFEKCMYGYEDVEQFEDV 358
+AL +V P A+H W + + K NL+ S F ++EKC+Y + + +F+ V
Sbjct: 472 QALVQVFPGAHHRYSAWQIRE---KERENLIPFPSEFKYEYEKCIYQTQTIVEFDSV 525
Score = 62.0 bits (149), Expect = 5e-10
Identities = 38/102 (37%), Positives = 55/102 (53%), Gaps = 15/102 (14%)
Query: 7 PKVDMFFCSLKNAWEFWNTYXGKXGFGVRKDYTHRKK-DGSVSSCRFVCCKEGMRKPDKR 65
P V + F + + A EF+N Y + GF VR +R + DG+VSS RFVC KEG
Sbjct: 43 PYVGLEFDTAEEAREFYNAYAARTGFKVRTGQLYRSRTDGTVSSRRFVCSKEGF------ 96
Query: 66 DYKTKNPRLETRTNCEARLGLKNVD-GKLMVHDFVEDHNHEL 106
+L +RT C A + ++ D GK ++ ++HNHEL
Sbjct: 97 -------QLNSRTGCTAFIRVQRRDTGKWVLDQIQKEHNHEL 131
>At3g06250 unknown protein
Length = 764
Score = 164 bits (415), Expect = 7e-41
Identities = 108/353 (30%), Positives = 165/353 (46%), Gaps = 46/353 (13%)
Query: 7 PKVDMFFCSLKNAWEFWNTYXGKXGFGVRKDYTHRKK-DGSVSSCRFVCCKEGMRKPDKR 65
P + F S A +F+ Y GF VR R K DGS++S RFVC KEG + P
Sbjct: 190 PYAGLEFNSANEACQFYQAYAEVVGFRVRIGQLFRSKVDGSITSRRFVCSKEGFQHP--- 246
Query: 66 DYKTKNPRLETRTNCEARLGLKNVD-GKLMVHDFVEDHNHELLLPETTHMLSSQRKVSEI 124
+R C A + +K D G +V +DHNH+L E + +K+++
Sbjct: 247 ----------SRMGCGAYMRIKRQDSGGWIVDRLNKDHNHDL---EPGKKNAGMKKITD- 292
Query: 125 HCQQIELADSVGVQQKKSFDLLSKEVGGRTNLGFIRL-DQKNYLRKKRERSLVQGEAGYL 183
GG ++ I L D N++ RE ++ + L
Sbjct: 293 -----------------------DVTGGLDSVDLIELNDLSNHISSTRENTIGKEWYPVL 329
Query: 184 LQYFQRKTVENPTFYHAYQLDIEDQITNVFWADARMLVDYSYFGDVVSLDSTYCTNSSHR 243
L YFQ K E+ F++A +LD ++FWAD+R S FGD V D++Y
Sbjct: 330 LDYFQSKQAEDMGFFYAIELDSNGSCMSIFWADSRSRFACSQFGDAVVFDTSYRKGDYSV 389
Query: 244 PLAIISGFNHHRGAVIFGAALLYDETAESYEWLFETFLEAHKQKMPQTVFTDQAKSMAKA 303
P A GFNHHR V+ G AL+ DE+ E++ WLF+T+L A + P+++ DQ + +A
Sbjct: 390 PFATFIGFNHHRQPVLLGGALVADESKEAFSWLFQTWLRAMSGRRPRSMVADQDLPIQQA 449
Query: 304 LAEVMPEAYHGLCTWHLMQNGIKHLGNLMKGESYFLSDFEKCMYGYEDVEQFE 356
+A+V P +H W + K NL + F ++EKC+Y + +F+
Sbjct: 450 VAQVFPGTHHRFSAWQIRS---KERENLRSFPNEFKYEYEKCLYQSQTTVEFD 499
Score = 58.2 bits (139), Expect = 7e-09
Identities = 35/102 (34%), Positives = 56/102 (54%), Gaps = 15/102 (14%)
Query: 7 PKVDMFFCSLKNAWEFWNTYXGKXGFGVRKDYTHRKK-DGSVSSCRFVCCKEGMRKPDKR 65
P V + F + + A +++N+Y + GF VR +R + DG+VSS RFVC KEG
Sbjct: 28 PYVGLEFDTAEEARDYYNSYATRTGFKVRTGQLYRSRTDGTVSSRRFVCSKEGF------ 81
Query: 66 DYKTKNPRLETRTNCEARLGLKNVD-GKLMVHDFVEDHNHEL 106
+L +RT C A + ++ D GK ++ ++HNH+L
Sbjct: 82 -------QLNSRTGCPAFIRVQRRDTGKWVLDQIQKEHNHDL 116
>At1g52520 F6D8.26
Length = 703
Score = 149 bits (375), Expect = 3e-36
Identities = 108/359 (30%), Positives = 166/359 (46%), Gaps = 30/359 (8%)
Query: 7 PKVDMFFCSLKNAWEFWNTYXGKXGFGVRKDYTHRKKDGSVSSCRFVCCKEGMRKPDKRD 66
P V M F S +A+ ++N Y + GF VR + K+ +CC + KR
Sbjct: 85 PAVGMEFESYDDAYNYYNCYASEVGFRVRVKNSWFKRRSKEKYGAVLCCSS---QGFKRI 141
Query: 67 YKTKNPRLETRTNCEARLGLKNVDGKLM-VHDFVEDHNHELLLPETTHMLSSQRKVSEIH 125
R ETRT C A + ++ VD K V + DHNH LL + + +RK
Sbjct: 142 NDVNRVRKETRTGCPAMIRMRQVDSKRWRVVEVTLDHNH-LLGCKLYKSVKRKRK----- 195
Query: 126 CQQIELADSVGVQQKKSFDLLSKEVGGRTNLGFIRLDQKNYLRKKRERS--------LVQ 177
C ++D+ + K + + G N + L KK + S L +
Sbjct: 196 CVSSPVSDAKTI---KLYRACVVDNGSNVN-------PNSTLNKKFQNSTGSPDLLNLKR 245
Query: 178 GEAGYLLQYFQRKTVENPTFYHAYQLDIEDQITNVFWADARMLVDYSYFGDVVSLDSTYC 237
G++ + YF R + NP F++ ++ E Q+ NVFWADA V SYFGDV+ +DS+Y
Sbjct: 246 GDSAAIYNYFCRMQLTNPNFFYLMDVNDEGQLRNVFWADAFSKVSCSYFGDVIFIDSSYI 305
Query: 238 TNSSHRPLAIISGFNHHRGAVIFGAALLYDETAESYEWLFETFLEAHKQKMPQTVFTDQA 297
+ PL +G NHH + L ET ESY WL + +L K + PQT+ TD+
Sbjct: 306 SGKFEIPLVTFTGVNHHGKTTLLSCGFLAGETMESYHWLLKVWLSVMK-RSPQTIVTDRC 364
Query: 298 KSMAKALAEVMPEAYHGLCTWHLMQNGIKHLGNLMKGESYFLSDFEKCMYGYEDVEQFE 356
K + A+++V P ++ H+M+ + LG L ++ F K +Y V +FE
Sbjct: 365 KPLEAAISQVFPRSHQRFSLTHIMRKIPEKLGGLHNYDA-VRKAFTKAVYETLKVVEFE 422
>At1g80010 hypothetical protein
Length = 696
Score = 142 bits (359), Expect = 2e-34
Identities = 104/360 (28%), Positives = 171/360 (46%), Gaps = 17/360 (4%)
Query: 7 PKVDMFFCSLKNAWEFWNTYXGKXGFGVRKDYTHRKKDGSVSSCRFVCCK-EGMRKPDKR 65
P M F S +A+ F+N+Y + GF +R + K++ +CC +G + +
Sbjct: 66 PTPGMEFESYDDAYSFYNSYARELGFAIRVKSSWTKRNSKEKRGAVLCCNCQGFKL--LK 123
Query: 66 DYKTKNPRLETRTNCEARLGLKNVDGKLMVHDFVE-DHNHELLLPETTHMLSSQRKVSEI 124
D ++ R ETRT C+A + L+ + D V+ DHNH P+ H S +K S
Sbjct: 124 DAHSR--RKETRTGCQAMIRLRLIHFDRWKVDQVKLDHNHSFD-PQRAHNSKSHKKSSSS 180
Query: 125 HCQQI----ELADSVGVQQKKSFDLLSKE----VGGRTNLGFIRLDQKNYLRKKRERSLV 176
E V V+ K + L+ + +G + G ++ + R L
Sbjct: 181 ASPATKTNPEPPPHVQVRTIKLYRTLALDTPPALGTSLSSGETSDLSLDHFQSSRRLEL- 239
Query: 177 QGEAGYLLQYFQRKTVENPTFYHAYQLDIEDQITNVFWADARMLVDYSYFGDVVSLDSTY 236
+G L +F + + +P F + L + + NVFW DAR YS+FGDV+ D+T
Sbjct: 240 RGGFRALQDFFFQIQLSSPNFLYLMDLADDGSLRNVFWIDARARAAYSHFGDVLLFDTTC 299
Query: 237 CTNSSHRPLAIISGFNHHRGAVIFGAALLYDETAESYEWLFETFLEAHKQKMPQTVFTDQ 296
+N+ PL G NHH ++ G LL D++ E+Y WLF +L + PQ T+Q
Sbjct: 300 LSNAYELPLVAFVGINHHGDTILLGCGLLADQSFETYVWLFRAWLTCMLGRPPQIFITEQ 359
Query: 297 AKSMAKALAEVMPEAYHGLCTWHLMQNGIKHLGNLMKGESYFLSDFEKCMYGYEDVEQFE 356
K+M A++EV P A+H L H++ N + + L + + ++ + +YG VE+FE
Sbjct: 360 CKAMRTAVSEVFPRAHHRLSLTHVLHNICQSVVQLQDSDLFPMA-LNRVVYGCLKVEEFE 418
>At2g32250 Mutator-like transposase
Length = 684
Score = 139 bits (351), Expect = 2e-33
Identities = 98/349 (28%), Positives = 158/349 (45%), Gaps = 47/349 (13%)
Query: 11 MFFCSLKNAWEFWNTYXGKXGFGVRKDYTHR-KKDGSVSSCRFVCCKEGMRKPDKRDYKT 69
M F S + A+ F+ Y GFG+ + R K+ G + C + G KR+ T
Sbjct: 42 MDFESKEAAYYFYREYARSVGFGITIKASRRSKRSGKFIDVKIACSRFGT----KREKAT 97
Query: 70 K-NPRLETRTNCEARLGLKNV-DGKLMVHDFVEDHNHELLLPETTHMLSSQRKVSEIHCQ 127
NPR +T C+A L +K D K ++++FV++HNHE+ P+ +
Sbjct: 98 AINPRSCPKTGCKAGLHMKRKEDEKWVIYNFVKEHNHEIC-PDDFY-------------- 142
Query: 128 QIELADSVGVQQKKSFDLLSKEVGGRTNLGFIRLDQKNYLRKKRERSLVQGEAGYLLQYF 187
V V+ K +K G ++K + +L + + LL++F
Sbjct: 143 -------VSVRGK------NKPAGALA------------IKKGLQLALEEEDLKLLLEHF 177
Query: 188 QRKTVENPTFYHAYQLDIEDQITNVFWADARMLVDYSYFGDVVSLDSTYCTNSSHRPLAI 247
+ P F++A D + ++ NVFW DA+ DY F DVV D+ Y N P A
Sbjct: 178 MEMQDKQPGFFYAVDFDSDKRVRNVFWLDAKAKHDYCSFSDVVLFDTFYVRNGYRIPFAP 237
Query: 248 ISGFNHHRGAVIFGAALLYDETAESYEWLFETFLEAHKQKMPQTVFTDQAKSMAKALAEV 307
G +HHR V+ G AL+ + + +Y WLF T+L+A + P + TDQ K ++ + EV
Sbjct: 238 FIGVSHHRQYVLLGCALIGEVSESTYSWLFRTWLKAVGGQAPGVMITDQDKLLSDIVVEV 297
Query: 308 MPEAYHGLCTWHLMQNGIKHLGNLMKGESYFLSDFEKCMYGYEDVEQFE 356
P+ H C W ++ + L + + F+ F C+ E FE
Sbjct: 298 FPDVRHIFCLWSVLSKISEMLNPFVSQDDGFMESFGNCVASSWTDEHFE 346
>At1g10240 unknown protein
Length = 680
Score = 139 bits (350), Expect = 2e-33
Identities = 101/360 (28%), Positives = 161/360 (44%), Gaps = 13/360 (3%)
Query: 7 PKVDMFFCSLKNAWEFWNTYXGKXGFGVRKDYTHRKKD-GSVSSCRFVCCKEGMRKPDKR 65
P + F + A+EF++T+ + GF +R+ T K G + R+ C P K
Sbjct: 48 PYLGQIFLTHDTAYEFYSTFAKRCGFSIRRHRTEGKDGVGKGLTRRYFVCHRAGNTPIKT 107
Query: 66 --DYKTKNPRLETRTNCEARLGLKNV----DGKLMVHDFVEDHNHELLLPETTHMLSSQR 119
+ K + R +R C+A L + + + V F HNHELL P L + R
Sbjct: 108 LSEGKPQRNRRSSRCGCQAYLRISKLTELGSTEWRVTGFANHHNHELLEPNQVRFLPAYR 167
Query: 120 KVSEIHCQQIELADSVGVQQKKSFDLLSKEVGGRTN-LGFIRLDQKNYLRKKRERSLVQG 178
+S+ +I + G+ ++ LL E L F D +N L+ ++ +
Sbjct: 168 SISDADKSRILMFSKTGISVQQMMRLLELEKCVEPGFLPFTEKDVRNLLQSFKKLD-PED 226
Query: 179 EAGYLLQYFQRKTVENPTFYHAYQLDIEDQITNVFWADARMLVDYSYFGDVVSLDSTYCT 238
E L+ Q ++P F + LD D++ N+ W+ A + Y FGD V D+T+
Sbjct: 227 ENIDFLRMCQSIKEKDPNFKFEFTLDANDKLENIAWSYASSIQSYELFGDAVVFDTTHRL 286
Query: 239 NSSHRPLAIISGFNHHRGAVIFGAALLYDETAESYEWLFETFLEAHKQKMPQTVFTDQAK 298
++ PL I G N++ FG LL DE S+ W + F K PQT+ TD
Sbjct: 287 SAVEMPLGIWVGVNNYGVPCFFGCVLLRDENLRSWSWALQAFTGFMNGKAPQTILTDHNM 346
Query: 299 SMAKALAEVMPEAYHGLCTWHLMQNGIKHLGNLMKGESY--FLSDFEKCMYGYEDVEQFE 356
+ +A+A MP H LC W ++ N GE Y + ++F + +Y E VE+FE
Sbjct: 347 CLKEAIAGEMPATKHALCIW-MVVGKFPSWFNAGLGERYNDWKAEFYR-LYHLESVEEFE 404
>At5g28530 far-red impaired response protein (FAR1) - like
Length = 700
Score = 127 bits (319), Expect = 9e-30
Identities = 90/334 (26%), Positives = 154/334 (45%), Gaps = 19/334 (5%)
Query: 5 WNPKVDMFFCSLKNAWEFWNTYXGKXGFGVRKDYTHRKKDGSVSSCRFVCCKEGMRKPDK 64
+ P V F + A+E+++T+ K GF +RK + ++ V FVC + G +P K
Sbjct: 68 FTPYVGQIFTTDDEAFEYYSTFARKSGFSIRKARSTESQNLGVYRRDFVCYRSGFNQPRK 127
Query: 65 R-DYKTKNPRLETRTNCEARLGLKN--VDG--KLMVHDFVEDHNHELLLPETTHMLSSQR 119
+ + + R R C+ +L L VDG V F HNHELL + +L + R
Sbjct: 128 KANVEHPRERKSVRCGCDGKLYLTKEVVDGVSHWYVSQFSNVHNHELLEDDQVRLLPAYR 187
Query: 120 KVSEIHCQQIELADSVGVQQKKSFDLLSKEVGGRTN-LGFIRLDQKNYLRKKRERSLVQG 178
K+ + ++I L G + LL E G + L FI D +N++R ++ VQ
Sbjct: 188 KIQQSDQERILLLSKAGFPVNRIVKLLELEKGVVSGQLPFIEKDVRNFVRACKKS--VQE 245
Query: 179 EAGYLLQYFQRKTVE-----------NPTFYHAYQLDIEDQITNVFWADARMLVDYSYFG 227
++ + + T+E + F + D ++ N+ WA + YS FG
Sbjct: 246 NDAFMTEKRESDTLELLECCKGLAERDMDFVYDCTSDENQKVENIAWAYGDSVRGYSLFG 305
Query: 228 DVVSLDSTYCTNSSHRPLAIISGFNHHRGAVIFGAALLYDETAESYEWLFETFLEAHKQK 287
DVV D++Y + L + G +++ A++ G LL DE+ S+ W +TF+ + +
Sbjct: 306 DVVVFDTSYRSVPYGLLLGVFFGIDNNGKAMLLGCVLLQDESCRSFTWALQTFVRFMRGR 365
Query: 288 MPQTVFTDQAKSMAKALAEVMPEAYHGLCTWHLM 321
PQT+ TD + A+ MP H + H++
Sbjct: 366 HPQTILTDIDTGLKDAIGREMPNTNHVVFMSHIV 399
>At4g19990 putative protein
Length = 672
Score = 118 bits (296), Expect = 4e-27
Identities = 91/362 (25%), Positives = 155/362 (42%), Gaps = 72/362 (19%)
Query: 13 FCSLKNAWEFWNTYXGKXGFG-VRKDYTHRKKDGSVSSCRFVCCKEGMRKPD-------- 63
F S + A+EF+ Y GF + K + G +FVC + G +K D
Sbjct: 27 FESKEEAFEFYKEYANSVGFTTIIKASRRSRMTGKFIDAKFVCTRYGSKKEDIDTGLGTD 86
Query: 64 -----KRDYKTKNPRLETRTNCEARLGLKN-VDGKLMVHDFVEDHNHELLLPETTHM--L 115
+ + + R ++T+C+A L +K DG+ +V V++HNHE+ + + L
Sbjct: 87 GFNIPQARKRGRINRSSSKTDCKAFLHVKRRQDGRWVVRSLVKEHNHEIFTGQADSLREL 146
Query: 116 SSQRKVSEIHCQQIELADSVGVQQKKSFDLLSKEVGGRTNLGFIRLDQKNYLRKKRERSL 175
S +RK+ +++ + KEV + R L
Sbjct: 147 SGRRKLEKLN------------------GAIVKEV--------------------KSRKL 168
Query: 176 VQGEAGYLLQYFQRKTVENPTFYHAYQLDIEDQITNVFWADARMLVDYSYFGDVVSLDST 235
G+ LL +F D++ + N+FW DA+ DY+ F DVVS+D+T
Sbjct: 169 EDGDVERLLNFFT---------------DMQS-LRNIFWVDAKGRFDYTCFSDVVSIDTT 212
Query: 236 YCTNSSHRPLAIISGFNHHRGAVIFG-AALLYDETAESYEWLFETFLEAHKQKMPQTVFT 294
+ N PL +G NHH ++ G LL DE+ + WLF +L+A P+ + T
Sbjct: 213 FIKNEYKLPLVAFTGVNHHGQFLLLGFGLLLTDESKSGFVWLFRAWLKAMHGCRPRVILT 272
Query: 295 DQAKSMAKALAEVMPEAYHGLCTWHLMQNGIKHLGNLMKGESYFLSDFEKCMYGYEDVEQ 354
+ + +A+ EV P + H W + + LG++++ E + + +YG E
Sbjct: 273 KHDQMLKEAVLEVFPSSRHCFYMWDTLGQMPEKLGHVIRLEKKLVDEINDAIYGSCQSED 332
Query: 355 FE 356
FE
Sbjct: 333 FE 334
>At4g38170 hypothetical protein
Length = 531
Score = 106 bits (264), Expect = 2e-23
Identities = 56/176 (31%), Positives = 95/176 (53%), Gaps = 5/176 (2%)
Query: 182 YLLQYFQRKTVENPTFYHAYQLDIEDQITNVFWADARMLVDYSYFGDVVSLDSTYCTNSS 241
++L Y +R+ +ENP F +A IED NVFWAD ++Y+YFGD + D+TY
Sbjct: 6 HVLNYLKRRQLENPGFLYA----IEDDCGNVFWADPTCRLNYTYFGDTLVFDTTYRRGKR 61
Query: 242 HR-PLAIISGFNHHRGAVIFGAALLYDETAESYEWLFETFLEAHKQKMPQTVFTDQAKSM 300
++ P A +GFNHH V+FG AL+ +E+ S+ WLF+T+L+A P ++ + + +
Sbjct: 62 YQVPFAAFTGFNHHGQPVLFGCALILNESESSFAWLFQTWLQAMSAPPPPSITVEPDRLI 121
Query: 301 AKALAEVMPEAYHGLCTWHLMQNGIKHLGNLMKGESYFLSDFEKCMYGYEDVEQFE 356
A++ V + + + + L ++ + F S+F C+ E +FE
Sbjct: 122 QVAVSRVFSQTRLRFSQPLIFEETEEKLAHVFQAHPTFESEFINCVTETETAAEFE 177
>At3g47280 putative protein
Length = 739
Score = 66.6 bits (161), Expect = 2e-11
Identities = 56/247 (22%), Positives = 100/247 (39%), Gaps = 5/247 (2%)
Query: 79 NCEARLGLKNVDGK--LMVHDFVEDHNHELLLPETTHMLSSQRKVSEIHCQQIELADSVG 136
NC RL G ++ +V H+ + L +H +S R + + E +
Sbjct: 216 NCSWRLRATRAGGSESYVIRKYVSHHSCDSSLRNVSHRQASARTLGRLISNHFE-GGKLP 274
Query: 137 VQQKKSFDLLSKEVGGRTNLGFIRLDQKNYLRKKRERSLVQGEAGYLLQYFQRKTVENPT 196
++ K+ ++ K+ G N QK+ + R L L ++F R V NP
Sbjct: 275 LRPKQLIEIFRKDHGVGINYSKAWRVQKH--AAELARGLPDDSFEVLPRWFHRVQVTNPG 332
Query: 197 FYHAYQLDIEDQITNVFWADARMLVDYSYFGDVVSLDSTYCTNSSHRPLAIISGFNHHRG 256
++ D ++ F A + Y V+S+D + T+ L S + +
Sbjct: 333 SITFFKKDSANKFKYAFLAFGASIRGYKLMRKVISIDGAHLTSKFKGTLLGASAQDGNFN 392
Query: 257 AVIFGAALLYDETAESYEWLFETFLEAHKQKMPQTVFTDQAKSMAKALAEVMPEAYHGLC 316
A++ E S++W + L + +D+A S+A L+ P A+HGLC
Sbjct: 393 LYPIAFAIVDSENDASWDWFLKCLLNIIPDENDLVFVSDRAASIASRLSGNYPLAHHGLC 452
Query: 317 TWHLMQN 323
T+HL +N
Sbjct: 453 TFHLQKN 459
>At5g34890 putative protein
Length = 681
Score = 65.5 bits (158), Expect = 4e-11
Identities = 57/257 (22%), Positives = 103/257 (39%), Gaps = 5/257 (1%)
Query: 79 NCEARLGLKNVDGK--LMVHDFVEDHNHELLLPETTHMLSSQRKVSEIHCQQIELADSVG 136
NC RL G ++ +V H+ + L +H +S R + + E +
Sbjct: 199 NCSWRLRATRAGGSESYVIRKYVSHHSCDSSLRNVSHRQASARTLGRLISNHFE-GGKLP 257
Query: 137 VQQKKSFDLLSKEVGGRTNLGFIRLDQKNYLRKKRERSLVQGEAGYLLQYFQRKTVENPT 196
+ K+ ++ K+ G N Q++ + R L L ++F R V NP
Sbjct: 258 LIPKQLIEIFRKDHGVGINYSKAWRVQEH--AAELARGLRDDSFEVLPRWFHRVQVTNPG 315
Query: 197 FYHAYQLDIEDQITNVFWADARMLVDYSYFGDVVSLDSTYCTNSSHRPLAIISGFNHHRG 256
++ D ++ F A + Y V+S+D + T+ L S + +
Sbjct: 316 SITFFKKDSANKFKYAFLAFGASIRGYKLMRKVISIDGAHLTSKFKGTLLGASAQDGNFN 375
Query: 257 AVIFGAALLYDETAESYEWLFETFLEAHKQKMPQTVFTDQAKSMAKALAEVMPEAYHGLC 316
A++ E S++W + L + +D+A S+A L+ P A+HGLC
Sbjct: 376 LYPIAFAIVDSENDASWDWFLKCLLNITPDENDLVFVSDRAASIASGLSGNYPLAHHGLC 435
Query: 317 TWHLMQNGIKHLGNLMK 333
T+HL +N H ++ K
Sbjct: 436 TFHLQKNLETHFRDVRK 452
>At3g59470 unknown protein
Length = 251
Score = 65.1 bits (157), Expect = 6e-11
Identities = 42/103 (40%), Positives = 54/103 (51%), Gaps = 3/103 (2%)
Query: 7 PKVDMFFCSLKNAWEFWNTYXGKXGFGVRKDYTHRKK-DGSVSSCRFVCCKEGMRKPDKR 65
P V F S A F+N Y K GF +R R + DGS + VC KEG R P KR
Sbjct: 70 PYVGQEFESEAAAHGFYNAYATKVGFVIRVSKLSRSRHDGSPIGRQLVCNKEGYRLPSKR 129
Query: 66 DYKTKNPRLETRTNCEARLGL-KNVDGKLMVHDFVEDHNHELL 107
D K R ETR C+A + + K GK ++ FV++HNH L+
Sbjct: 130 D-KVIRQRAETRVGCKAMILIRKENSGKWVITKFVKEHNHSLM 171
>At2g15150 putative transposase of FARE2.2 (CDS1)
Length = 607
Score = 63.2 bits (152), Expect = 2e-10
Identities = 54/247 (21%), Positives = 99/247 (39%), Gaps = 5/247 (2%)
Query: 79 NCEARLGLKNVDGK--LMVHDFVEDHNHELLLPETTHMLSSQRKVSEIHCQQIELADSVG 136
NC RL G + +V H+ + L +H +S R + + E +
Sbjct: 93 NCSWRLRATRAGGSESYAIRKYVSHHSCDSSLRNVSHRQASARTLGRLISNHFE-GGKLP 151
Query: 137 VQQKKSFDLLSKEVGGRTNLGFIRLDQKNYLRKKRERSLVQGEAGYLLQYFQRKTVENPT 196
++ K+ ++ K+ G N Q++ + R L L ++F R V NP
Sbjct: 152 LRPKQLIEIFRKDHGVGINYSKAWRVQEH--AAELARGLPDDSFEVLPRWFHRVQVTNPG 209
Query: 197 FYHAYQLDIEDQITNVFWADARMLVDYSYFGDVVSLDSTYCTNSSHRPLAIISGFNHHRG 256
++ D ++ F A + Y V+S+D + T+ L S + +
Sbjct: 210 SITFFKKDSANKFKYAFLAFGASIRGYKLMRKVISIDGAHLTSKFKGTLLGASAQDGNFN 269
Query: 257 AVIFGAALLYDETAESYEWLFETFLEAHKQKMPQTVFTDQAKSMAKALAEVMPEAYHGLC 316
A++ E S++W + L + +++A S+A L+ P A+HGLC
Sbjct: 270 LYPIAFAIVDSENDASWDWFLKCLLNIIPDENDLVFVSERAASIASGLSGNYPLAHHGLC 329
Query: 317 TWHLMQN 323
T+HL +N
Sbjct: 330 TFHLQKN 336
>At3g32060 hypothetical protein
Length = 487
Score = 61.6 bits (148), Expect = 6e-10
Identities = 45/175 (25%), Positives = 70/175 (39%), Gaps = 12/175 (6%)
Query: 186 YFQRKTVENPTFYHAYQLDIEDQITNVFWADARMLVDYSYFGDVVSLDSTYCTNSSHRPL 245
Y +K + Y +LD D+ VF A + + V+ +D+T+ N L
Sbjct: 225 YMLKKVNDGTVTY--LKLDENDKFQYVFVALGASIEGFRVMRKVLIVDATHLKNGYGGVL 282
Query: 246 AIISGFNHHRGAVIFGAALLYDETAESYEWLFETFLEAHKQKMPQTVFTDQAKSMAKALA 305
S + +R I A+L E S+EW FE TD+ S+ KA+
Sbjct: 283 VFASAQDPNRHHYIIAFAVLDGENDASWEWFFEKLKTVVPDTSELVFMTDRNASLIKAIR 342
Query: 306 EVMPEAYHGLCTWHLMQNGIKHLGNLMK----------GESYFLSDFEKCMYGYE 350
V A+HG C WHL QN H + + Y ++DF + G++
Sbjct: 343 NVYTAAHHGYCIWHLSQNVKGHATHTNRDVLAWKFQELSRVYVVADFNRAYDGFK 397
>At2g23500 Mutator-like transposase
Length = 784
Score = 61.6 bits (148), Expect = 6e-10
Identities = 45/175 (25%), Positives = 70/175 (39%), Gaps = 12/175 (6%)
Query: 186 YFQRKTVENPTFYHAYQLDIEDQITNVFWADARMLVDYSYFGDVVSLDSTYCTNSSHRPL 245
Y +K + Y +LD D+ VF A + + V+ +D+T+ N L
Sbjct: 355 YMLKKVNDGTVTY--LKLDENDKFQYVFVALGASIEGFRVMRKVLIVDATHLKNGYGGVL 412
Query: 246 AIISGFNHHRGAVIFGAALLYDETAESYEWLFETFLEAHKQKMPQTVFTDQAKSMAKALA 305
S + +R I A+L E S+EW FE TD+ S+ KA+
Sbjct: 413 VFASAQDPNRHHYIIAFAVLDGENDASWEWFFEKLKTVVPDTSELVFMTDRNASLIKAIR 472
Query: 306 EVMPEAYHGLCTWHLMQNGIKHLGNLMK----------GESYFLSDFEKCMYGYE 350
V A+HG C WHL QN H + + Y ++DF + G++
Sbjct: 473 NVYTAAHHGYCIWHLSQNVKGHATHTNRDVLAWKFQELSRVYVVADFNRAYDGFK 527
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.320 0.136 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,355,889
Number of Sequences: 26719
Number of extensions: 344731
Number of successful extensions: 953
Number of sequences better than 10.0: 82
Number of HSP's better than 10.0 without gapping: 65
Number of HSP's successfully gapped in prelim test: 17
Number of HSP's that attempted gapping in prelim test: 834
Number of HSP's gapped (non-prelim): 94
length of query: 362
length of database: 11,318,596
effective HSP length: 101
effective length of query: 261
effective length of database: 8,619,977
effective search space: 2249813997
effective search space used: 2249813997
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC148236.2


