

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148098.5 + phase: 0
(189 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
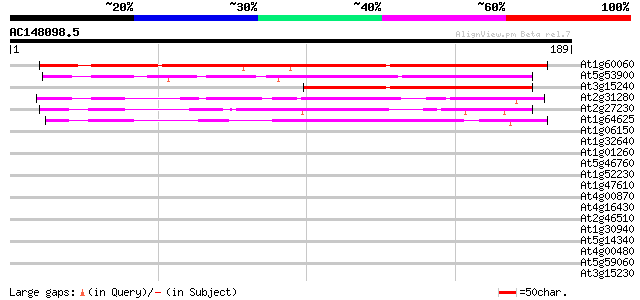
Score E
Sequences producing significant alignments: (bits) Value
At1g60060 unknown protein 192 9e-50
At5g53900 unknown protein 109 1e-24
At3g15240 unknown protein 80 8e-16
At2g31280 unknown protein 71 3e-13
At2g27230 unknown protein 51 4e-07
At1g64625 protein kinase, putative 47 6e-06
At1g06150 unknown protein 36 0.011
At1g32640 transcription factor like protein, BHLH6 32 0.15
At1g01260 transcription factor MYC7E, putative bHLH transcriptio... 32 0.15
At5g46760 putative transcription factor BHLH5 31 0.34
At1g52230 photosystem I subunit VI precursor (PsaH2) 31 0.45
At1g47610 unknown protein 31 0.45
At4g00870 putative bHLH transcription factor (bHLH014) 30 0.58
At4g16430 putative transcription factor BHLH3 30 1.00
At2g46510 putative bHLH transcription factor 28 2.2
At1g30940 F17F8.19 28 2.2
At5g14340 MYB40 - putative transcription factor 28 3.8
At4g00480 putative transcription factor BHLH12 28 3.8
At5g59060 putative protein 27 4.9
At3g15230 unknown protein 27 4.9
>At1g60060 unknown protein
Length = 386
Score = 192 bits (488), Expect = 9e-50
Identities = 95/189 (50%), Positives = 129/189 (67%), Gaps = 24/189 (12%)
Query: 11 LLQHTLRNLCSFPTSSTSSKWVYAVFWRVVPRIFPPPRWEFGGTSLDRSKGNKRNWIIVW 70
LLQHTLR+LC +S+WVYAVFWR++PR +PPP+W+ G + DRS+GN+RNWI+VW
Sbjct: 13 LLQHTLRSLCIHE----NSQWVYAVFWRILPRNYPPPKWD-GQGAYDRSRGNRRNWILVW 67
Query: 71 EDGFCDF-------NECEQRKSGCLNERFG-----------ADVFFKMSHEVYSYGEGLV 112
EDGFC+F + E G + +G ++FFKMSHE+Y+YGEGL+
Sbjct: 68 EDGFCNFAASAAEMSSGEGSGGGGGSAAYGNSDFQQYQGLQPELFFKMSHEIYNYGEGLI 127
Query: 113 GKVAADNGHKWVYSDTQNGCEANYVGPWNASIDHQPRAWEFQLNSGIQTIAVIAVKEGLV 172
GKVAAD+ HKW+Y + N E N++ W+ S D PR WE Q SGI+TIA+I+V+EG+V
Sbjct: 128 GKVAADHSHKWIYKE-PNDQEINFLSAWHNSADSYPRTWEAQFQSGIKTIALISVREGVV 186
Query: 173 QLGSFDKVL 181
QLG+ KV+
Sbjct: 187 QLGAVHKVI 195
>At5g53900 unknown protein
Length = 377
Score = 109 bits (272), Expect = 1e-24
Identities = 69/173 (39%), Positives = 89/173 (50%), Gaps = 25/173 (14%)
Query: 12 LQHTLRNLCSFPTSSTSSKWVYAVFWRVVPRIFPPPRWEFG-GTSLDRSKGNKRNWIIVW 70
L LR++C +S W+Y+VFW + PR PR G G + G+ +++W
Sbjct: 21 LHEALRSVCF------NSDWIYSVFWTIRPR----PRVRGGNGCKIGDESGSL---MLMW 67
Query: 71 EDGFCDFNECEQRKSGCLN-------ERFGADVFFKMSHEVYSYGEGLVGKVAADNGHKW 123
EDGFC E CL E F KMS ++Y+YGEGL+GKVA+D HKW
Sbjct: 68 EDGFCGGGRSEDL---CLETDIEGHEEDLVRKAFSKMSIQLYNYGEGLMGKVASDKCHKW 124
Query: 124 VYSDTQNGCEANYVGPWNASIDHQPRAWEFQLNSGIQTIAVIAVKEGLVQLGS 176
V+ + E N W +S D P W Q SGIQTIAVI GL+QLGS
Sbjct: 125 VFKEPSES-EPNLANYWQSSFDALPPEWTDQFESGIQTIAVIQAGHGLLQLGS 176
>At3g15240 unknown protein
Length = 251
Score = 79.7 bits (195), Expect = 8e-16
Identities = 40/77 (51%), Positives = 51/77 (65%), Gaps = 1/77 (1%)
Query: 100 MSHEVYSYGEGLVGKVAADNGHKWVYSDTQNGCEANYVGPWNASIDHQPRAWEFQLNSGI 159
MS ++Y+YGEGL+GKVA+D HKWV+ + Q E+N W +S D P W Q SGI
Sbjct: 1 MSIQLYNYGEGLMGKVASDKCHKWVFKE-QTESESNASSYWQSSFDAIPSEWNDQFESGI 59
Query: 160 QTIAVIAVKEGLVQLGS 176
+TIAVI GL+QLGS
Sbjct: 60 RTIAVIQAGHGLLQLGS 76
>At2g31280 unknown protein
Length = 720
Score = 71.2 bits (173), Expect = 3e-13
Identities = 53/172 (30%), Positives = 84/172 (48%), Gaps = 40/172 (23%)
Query: 10 SLLQHTLRNLCSFPTSSTSSKWVYAVFWRVVPRIFPPPRWEFGGTSLDRSKGNKRNWIIV 69
S Q L++ C ++ W YAVFW++ +G++ ++
Sbjct: 3 STSQEILKSFCF------NTDWDYAVFWQL------------------NHRGSRM--VLT 36
Query: 70 WEDGFCDFNECEQRKSGCLNERFGADVFFKMSHEVYSYGEGLVGKVAADNGHKWVYSDTQ 129
ED + D + + ++ G V KMS+ VYS GEG+VG+VA H+WV+ +
Sbjct: 37 LEDAYYDHHGTNMHGA---HDPLGLAVA-KMSYHVYSLGEGIVGQVAVSGEHQWVFPENY 92
Query: 130 NGCEANYVGPWNASIDHQPRAWEFQLNSGIQTIAVIAVKE-GLVQLGSFDKV 180
N C N++ + WE Q+++GI+TI V+AV G+VQLGS KV
Sbjct: 93 NNC--------NSAFEFH-NVWESQISAGIKTILVVAVGPCGVVQLGSLCKV 135
>At2g27230 unknown protein
Length = 650
Score = 50.8 bits (120), Expect = 4e-07
Identities = 46/181 (25%), Positives = 78/181 (42%), Gaps = 57/181 (31%)
Query: 11 LLQHTLRNLCSFPTSSTSSKWVYAVFWRVVPRIFPPPRWEFGGTSLDRSKGNKRNWIIVW 70
LL+ LR++C +++W YAVFW++ G + + +++W
Sbjct: 4 LLREALRSMC------VNNQWSYAVFWKI---------------------GCQNSSLLIW 36
Query: 71 EDGFCDFNECEQRKSGCLNERFGADVF-----------FKMSHEVYSYGEGLVGKVAADN 119
E+ C +NE E + G D +++ + GEGLVG+ A
Sbjct: 37 EE--C-YNETESSSNPRRLCGLGVDTQGNEKVQLLTNRMMLNNRIILVGEGLVGRAAFTG 93
Query: 120 GHKWVYSDTQNGCEANYVGPWNASIDHQPRAWE---FQLNSGIQTIAVI-AVKEGLVQLG 175
H+W+ +++ +N + H P Q ++GIQT+AV V G+VQLG
Sbjct: 94 HHQWILANS-----------FNRDV-HPPEVINEMLLQFSAGIQTVAVFPVVPHGVVQLG 141
Query: 176 S 176
S
Sbjct: 142 S 142
>At1g64625 protein kinase, putative
Length = 527
Score = 47.0 bits (110), Expect = 6e-06
Identities = 43/170 (25%), Positives = 69/170 (40%), Gaps = 47/170 (27%)
Query: 13 QHTLRNLCSFPTSSTSSKWVYAVFWRVVPRIFPPPRWEFGGTSLDRSKGNKRNWIIVWED 72
+H L++LC S W YAVFWR P + I+ +E+
Sbjct: 6 KHILKSLC------LSHGWSYAVFWRYDPI---------------------NSMILRFEE 38
Query: 73 GFCDFNECEQRKSGCLNERFGADVFFKMSHEVYSYGEGLVGKVAADNGHKWVYSDTQNGC 132
+ N+ + M + G+G+VG+VA+ H+W++SDT
Sbjct: 39 AY--------------NDEQSVALVDDMVLQAPILGQGIVGEVASSGNHQWLFSDTLFQW 84
Query: 133 EANYVGPWNASIDHQPRAWEFQLNSGIQTIAVIAV-KEGLVQLGSFDKVL 181
E + + R + + QTIA+I + G+VQLGS K+L
Sbjct: 85 EHEFQNQFLCGFKILIRQFTY-----TQTIAIIPLGSSGVVQLGSTQKIL 129
>At1g06150 unknown protein
Length = 1288
Score = 36.2 bits (82), Expect = 0.011
Identities = 18/36 (50%), Positives = 25/36 (69%), Gaps = 1/36 (2%)
Query: 146 HQPRAWEFQLNSGIQTIAVIAVKE-GLVQLGSFDKV 180
H WE Q+++GI+TI ++AV G+VQLGS KV
Sbjct: 73 HVHNGWESQISAGIKTILIVAVGSCGVVQLGSLCKV 108
Score = 27.7 bits (60), Expect = 3.8
Identities = 12/28 (42%), Positives = 18/28 (63%), Gaps = 6/28 (21%)
Query: 12 LQHTLRNLCSFPTSSTSSKWVYAVFWRV 39
LQ LR++CS ++ W YAVFW++
Sbjct: 5 LQQILRSICS------NTDWNYAVFWKL 26
>At1g32640 transcription factor like protein, BHLH6
Length = 623
Score = 32.3 bits (72), Expect = 0.15
Identities = 48/217 (22%), Positives = 71/217 (32%), Gaps = 76/217 (35%)
Query: 5 LPLLHSLLQHTLRNLCSFPTSSTSSKWVYAVFWRVVPRIFPPPRWEFGGTSLDRSKGNKR 64
+P Q TL+ T W YA+FW+ P ++F G S
Sbjct: 57 IPAQAGFNQETLQQRLQALIEGTHEGWTYAIFWQ--------PSYDFSGAS--------- 99
Query: 65 NWIIVWEDGFCDFNECE---QRKSGC------------------LNERFGADV------- 96
++ W DG+ E + +R+S LN V
Sbjct: 100 --VLGWGDGYYKGEEDKANPRRRSSSPPFSTPADQEYRKKVLRELNSLISGGVAPSDDAV 157
Query: 97 ---------FFKMS-HEVYSYGEGLVGKVAADNGHKWVYSDTQ---NGCEANYVGPWNAS 143
FF +S + ++ G GL GK A WV Q +GCE G
Sbjct: 158 DEEVTDTEWFFLVSMTQSFACGAGLAGKAFATGNAVWVSGSDQLSGSGCERAKQGG---- 213
Query: 144 IDHQPRAWEFQLNSGIQTIAVIAVKEGLVQLGSFDKV 180
G+ TIA I G+V++GS + +
Sbjct: 214 ------------VFGMHTIACIPSANGVVEVGSTEPI 238
>At1g01260 transcription factor MYC7E, putative bHLH transcription
factor (AtbHLH013)
Length = 590
Score = 32.3 bits (72), Expect = 0.15
Identities = 44/204 (21%), Positives = 72/204 (34%), Gaps = 72/204 (35%)
Query: 12 LQHTLRNLCSFPTSSTSSKWVYAVFWRVVPRIFPPPRWEFGGTSLDRSKGNKRNWIIVWE 71
LQ+ L +L P +S S W YA+FW++ S+ + ++ W
Sbjct: 48 LQNKLSDLVERPNASNFS-WNYAIFWQI-------------------SRSKAGDLVLCWG 87
Query: 72 DGFCD-------------------------------------FNECEQRKSGCLNERFGA 94
DG+C F E+ +R
Sbjct: 88 DGYCREPKEGEKSEIVRILSMGREEETHQTMRKRVLQKLHDLFGGSEEENCALGLDRVTD 147
Query: 95 DVFFKMSHEVYSY--GEGLVGKVAADNGHKWVYSDTQNGCEANYVGPWNASIDHQPRAWE 152
F +S +S+ GEG GK A W+ SD N+ D+ R++
Sbjct: 148 TEMFLLSSMYFSFPRGEGGPGKCFASAKPVWL-SDVV-----------NSGSDYCVRSF- 194
Query: 153 FQLNSGIQTIAVIAVKEGLVQLGS 176
++GIQT+ ++ G+V+LGS
Sbjct: 195 LAKSAGIQTVVLVPTDLGVVELGS 218
>At5g46760 putative transcription factor BHLH5
Length = 592
Score = 31.2 bits (69), Expect = 0.34
Identities = 40/189 (21%), Positives = 73/189 (38%), Gaps = 37/189 (19%)
Query: 13 QHTLRNLCSFPTSSTSSKWVYAVFWRVVPRIFPPPRWEFGGTSLDRSKGNKRNWIIVWED 72
+ TL+ S W YA+FW++ D S G+ I+ W D
Sbjct: 49 EDTLQQRLQALIESAGENWTYAIFWQI-------------SHDFDSSTGD-NTVILGWGD 94
Query: 73 GFCDFNECEQRKSGCLNERFGADVFFKMSHEVYSYGEGLVGKVAADNGH-----KWVYSD 127
G+ E +++K N + ++ E+ S G +G N +W +
Sbjct: 95 GYYKGEEDKEKKKNNTNTA-EQEHRKRVIRELNSLISGGIGVSDESNDEEVTDTEWFFLV 153
Query: 128 TQNGCEANYVG-PWNASIDHQ---------------PRAWEFQLNSGIQTIAVIAVKEGL 171
+ N VG P + ++ + RA + Q+ G++T+ IA + G+
Sbjct: 154 SMTQSFVNGVGLPGESFLNSRVIWLSGSGALTGSGCERAGQGQI-YGLKTMVCIATQNGV 212
Query: 172 VQLGSFDKV 180
V+LGS + +
Sbjct: 213 VELGSSEVI 221
>At1g52230 photosystem I subunit VI precursor (PsaH2)
Length = 145
Score = 30.8 bits (68), Expect = 0.45
Identities = 22/80 (27%), Positives = 32/80 (39%), Gaps = 2/80 (2%)
Query: 93 GADVFFKMSHEVYSYGEGLVGKVAADNGHKWVYSDTQNGCEANYVGPWNASIDHQPRAWE 152
GA +F K S + + G V A G K VY D ++ N G W+ P +
Sbjct: 26 GAKLFIKPSRQSFKTKSTRAGAVVAKYGDKSVYFDLED--LGNTTGQWDVYGSDAPSPYN 83
Query: 153 FQLNSGIQTIAVIAVKEGLV 172
+ +T A K GL+
Sbjct: 84 PLQSKFFETFAAPFTKRGLL 103
>At1g47610 unknown protein
Length = 351
Score = 30.8 bits (68), Expect = 0.45
Identities = 22/78 (28%), Positives = 37/78 (47%), Gaps = 9/78 (11%)
Query: 105 YSYGEGLVGKVAADNGHKWVYSDTQN----GCEANYVGPWNASIDHQPRAWEFQLNSGIQ 160
YS G+ LVG++ + GH + + T + G + NY+ W + F+ NSG+
Sbjct: 10 YSDGDSLVGEIVREEGHIYSLAATNDLLYTGSDNNYIRVWKNLNEFS----GFKSNSGL- 64
Query: 161 TIAVIAVKEGLVQLGSFD 178
A++ +E V G D
Sbjct: 65 VKAIVISREAKVFTGHQD 82
>At4g00870 putative bHLH transcription factor (bHLH014)
Length = 423
Score = 30.4 bits (67), Expect = 0.58
Identities = 44/186 (23%), Positives = 69/186 (36%), Gaps = 58/186 (31%)
Query: 11 LLQHTLRNLCSFPTSSTSSKWVYAVFWRVVPRIFPPPRWEFGGTSLDRSKGNKRNWIIVW 70
+LQ LR F ++ +W Y +FW+ + F S DRS +VW
Sbjct: 34 VLQQKLR----FVVETSPDRWAYVIFWQKM----------FDDQS-DRS-------YLVW 71
Query: 71 EDG-FCDFN-------------ECEQRKSGCLNERFGADVFFKMSHEVYSYGEGLVG-KV 115
DG FC ECE G G D+ ++ + YGE K
Sbjct: 72 VDGHFCGNKNNNSQENYTTNSIECELMMDG------GDDL--ELFYAASFYGEDRSPRKE 123
Query: 116 AADNGHKWVYSDTQNGCEANYVGPWNASIDHQPRAWEFQLNSGIQTIAVIAVKEGLVQLG 175
+D W+ GP + RA E + G+ T+ I + G+++LG
Sbjct: 124 VSDESLVWL------------TGPDELRFSNYERAKEAGFH-GVHTLVSIPINNGIIELG 170
Query: 176 SFDKVL 181
S + ++
Sbjct: 171 SSESII 176
>At4g16430 putative transcription factor BHLH3
Length = 467
Score = 29.6 bits (65), Expect = 1.00
Identities = 42/200 (21%), Positives = 66/200 (33%), Gaps = 64/200 (32%)
Query: 6 PLLHSLLQHTLRNLCSFPTSSTSSKWVYAVFWRVVPRIFPPPRWEFGGTSLDRSKGNKRN 65
P S LQ LR++ S W YA+FW ++++ S G
Sbjct: 44 PPSDSNLQQGLRHVVE------GSDWDYALFWLA--------------SNVNSSDG---- 79
Query: 66 WIIVWEDGFCDFNECEQRKSGCLNERFGADVFFKMSHEVYSYGEGLVGKVAADNGHKWVY 125
+++W DG C + + + V K+ + V +D H+ V
Sbjct: 80 CVLIWGDGHCRVKKGASGEDYSQQDEIKRRVLRKLH----------LSFVGSDEDHRLVK 129
Query: 126 SDTQNG--------------CEANYVGPWNASIDHQPRAWEFQLNS-------------- 157
S C+ N GP + +P W L S
Sbjct: 130 SGALTDLDMFYLASLYFSFRCDTNKYGPAGTYVSGKP-LWAADLPSCLSYYRVRSFLARS 188
Query: 158 -GIQTIAVIAVKEGLVQLGS 176
G QT+ + V G+V+LGS
Sbjct: 189 AGFQTVLSVPVNSGVVELGS 208
>At2g46510 putative bHLH transcription factor
Length = 662
Score = 28.5 bits (62), Expect = 2.2
Identities = 22/83 (26%), Positives = 44/83 (52%), Gaps = 14/83 (16%)
Query: 95 DVFFKMS-HEVYSYGEGLVGKVAADNGHKWVYSDTQNGCEANYVGPWNASIDHQPRAWEF 153
++FF S + +++GEG G+ + H W+ SD N + D+ R++
Sbjct: 152 EIFFLASMYFFFNHGEGGPGRCYSSGKHVWL-SDAVN-----------SESDYCFRSFMA 199
Query: 154 QLNSGIQTIAVIAVKEGLVQLGS 176
+ ++GI+TI ++ G+++LGS
Sbjct: 200 K-SAGIRTIVMVPTDAGVLELGS 221
>At1g30940 F17F8.19
Length = 461
Score = 28.5 bits (62), Expect = 2.2
Identities = 21/62 (33%), Positives = 28/62 (44%), Gaps = 4/62 (6%)
Query: 55 SLDRSKGNKRNWIIVWEDGFCDFNECEQRKSGCLNERFGADVFFKMSHEV----YSYGEG 110
S+ RSK R I V +DG F Q ++ C N+ AD K S + SY G
Sbjct: 92 SVARSKARPRLLIGVQQDGEWRFYSTPQSQNPCENKLSPADFHMKFSKGISNYSCSYASG 151
Query: 111 LV 112
L+
Sbjct: 152 LI 153
>At5g14340 MYB40 - putative transcription factor
Length = 263
Score = 27.7 bits (60), Expect = 3.8
Identities = 14/38 (36%), Positives = 19/38 (49%), Gaps = 2/38 (5%)
Query: 138 GPWNASIDHQPRAWEFQLNSGIQTIAVIAVKEGLVQLG 175
GPW DH R F LN+GI ++ GL++ G
Sbjct: 15 GPWTIEEDH--RLMNFILNNGIHCWRIVPKLAGLLRCG 50
>At4g00480 putative transcription factor BHLH12
Length = 526
Score = 27.7 bits (60), Expect = 3.8
Identities = 19/58 (32%), Positives = 30/58 (50%), Gaps = 8/58 (13%)
Query: 55 SLDRSKGNKRNW------IIVWEDG--FCDFNECEQRKSGCLNERFGADVFFKMSHEV 104
S DRS N +N I+ E G F + +CEQ+ SG + ++ +V K+ H+V
Sbjct: 265 SADRSGKNDKNIRHRQPNIVTSEPGSSFLRWKQCEQQVSGFVQKKKSQNVLRKILHDV 322
>At5g59060 putative protein
Length = 110
Score = 27.3 bits (59), Expect = 4.9
Identities = 12/38 (31%), Positives = 25/38 (65%), Gaps = 1/38 (2%)
Query: 5 LPLLHSLLQHTLRN-LCSFPTSSTSSKWVYAVFWRVVP 41
+P++ + +LR+ L SFP S+S+ ++ AV++ +P
Sbjct: 71 IPVIRKSIDRSLRDRLLSFPAPSSSAPFLLAVYFGCIP 108
>At3g15230 unknown protein
Length = 47
Score = 27.3 bits (59), Expect = 4.9
Identities = 15/42 (35%), Positives = 20/42 (46%), Gaps = 10/42 (23%)
Query: 12 LQHTLRNLCSFPTSSTSSKWVYAVFWRVVPRIFPPPRWEFGG 53
L LR +C ++ W Y+VFW + PR PR GG
Sbjct: 4 LHDALRTVC------LNTDWTYSVFWSIRPR----PRVRGGG 35
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.321 0.138 0.457
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,001,781
Number of Sequences: 26719
Number of extensions: 220704
Number of successful extensions: 435
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 9
Number of HSP's successfully gapped in prelim test: 15
Number of HSP's that attempted gapping in prelim test: 405
Number of HSP's gapped (non-prelim): 31
length of query: 189
length of database: 11,318,596
effective HSP length: 94
effective length of query: 95
effective length of database: 8,807,010
effective search space: 836665950
effective search space used: 836665950
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 57 (26.6 bits)
Medicago: description of AC148098.5


