

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147960.5 - phase: 0
(617 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
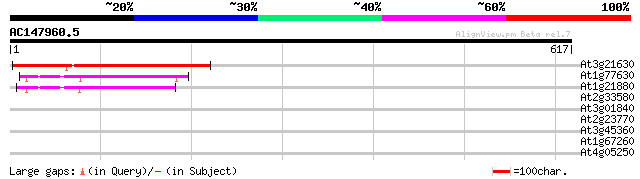
Score E
Sequences producing significant alignments: (bits) Value
At3g21630 receptor like protein kinase 219 3e-57
At1g77630 unknown protein with predicted GPI-anchor 58 1e-08
At1g21880 receptor-like GPI-anchored protein (lysM) 1 54 2e-07
At2g33580 putative protein kinase 37 0.032
At3g01840 putative protein kinase 35 0.12
At2g23770 putative protein kinase 31 1.8
At3g45360 putative protein 30 5.2
At1g67260 hypothetical protein 30 5.2
At4g05250 putative protein (ubiquitin like) 29 6.8
>At3g21630 receptor like protein kinase
Length = 617
Score = 219 bits (559), Expect = 3e-57
Identities = 109/221 (49%), Positives = 152/221 (68%), Gaps = 5/221 (2%)
Query: 4 IKFRLSFLFMLLASKSFIAESKCSKTCNIALASYYLQDDTNLTYVSNIMQSNLVTKPE-- 61
+K L +LL S F ESKC +C +ALASYYL++ T L+ ++ + S++ +
Sbjct: 3 LKISLIAPILLLFSFFFAVESKCRTSCPLALASYYLENGTTLSVINQNLNSSIAPYDQIN 62
Query: 62 --DIVSYNTDTITNKDFVQSFTRVNVPFPCDCIHDEFLGHIFQYQVATKDTYLSVASNNY 119
I+ YN++ I +KD +Q +RV VPFPC+C +FLGH F Y V +DTY VA +NY
Sbjct: 63 FDPILRYNSN-IKDKDRIQMGSRVLVPFPCECQPGDFLGHNFSYSVRQEDTYERVAISNY 121
Query: 120 SNLTTSEWLQNFNSYPSNDIPDTGTLNVTVNCSCGNSDVSKDYGLFITYPLRPEDSLELI 179
+NLTT E LQ N +P+ +IP + TLNV VNCSCG+ VSKD+GLF+TYPLRPEDSL I
Sbjct: 122 ANLTTMESLQARNPFPATNIPLSATLNVLVNCSCGDESVSKDFGLFVTYPLRPEDSLSSI 181
Query: 180 SNKTEIDAELLQKYNPGVNFSQGSGLVYIPGKDQNRNYVPF 220
+ + + A++LQ+YNPGVNF+ G+G+VY+PG+D N + PF
Sbjct: 182 ARSSGVSADILQRYNPGVNFNSGNGIVYVPGRDPNGAFPPF 222
>At1g77630 unknown protein with predicted GPI-anchor
Length = 423
Score = 58.2 bits (139), Expect = 1e-08
Identities = 54/207 (26%), Positives = 90/207 (43%), Gaps = 26/207 (12%)
Query: 11 LFMLLAS--------KSFIAESKCSKTCNIALASYYLQDDTNLTYVSNIMQSNLVTKPED 62
LF++LAS KS I TCN +L Y L D +T V+++ Q + P
Sbjct: 10 LFLILASSLASMATAKSTIEPCSSKDTCN-SLLGYTLYTDLKVTEVASLFQVD----PVS 64
Query: 63 IVSYNTDTITNKDF----VQSFTRVNVPFPCDCIHDEFLGHIFQYQVATKDTYLSVASNN 118
++ N+ I+ D + + + +P C C+ Y+ T DT S+A +
Sbjct: 65 MLLSNSIDISYPDVENHVLPAKLFLKIPITCSCVDGIRKSLSTHYKTRTSDTLGSIADSV 124
Query: 119 YSNLTTSEWLQNFNSYPSNDIPDTGT-LNVTVNCSCGNSDVSKDYGLFITYPLRPEDSLE 177
Y L + E +Q NS + D GT L + + C+C N L+++Y +R D++
Sbjct: 125 YGGLVSPEQIQVANSETDLSVLDVGTKLVIPLPCACFNGTDESLPALYLSYVVRGIDTMA 184
Query: 178 LISNK--------TEIDAELLQKYNPG 196
I+ + T ++A NPG
Sbjct: 185 GIAKRFSTSVTDLTNVNAMGAPDINPG 211
>At1g21880 receptor-like GPI-anchored protein (lysM) 1
Length = 416
Score = 54.3 bits (129), Expect = 2e-07
Identities = 48/188 (25%), Positives = 84/188 (44%), Gaps = 18/188 (9%)
Query: 8 LSFLFMLLAS--------KSFIAESKCSKTCNIALASYYLQDDTNLTYVSNIMQSNLVTK 59
L F+ ++LAS KS I + TCN AL Y L D ++ V+++ Q +
Sbjct: 10 LIFVSLILASSLTFTATAKSTIEPCSSNDTCN-ALLGYTLYTDLKVSEVASLFQVD---- 64
Query: 60 PEDIVSYNTDTITNKD----FVQSFTRVNVPFPCDCIHDEFLGHIFQYQVATKDTYLSVA 115
P I+ N I+ D + S + +P C C+ Y+ D S+A
Sbjct: 65 PISILLANAIDISYPDVENHILPSKLFLKIPITCSCVDGIRKSVSTHYKTRPSDNLGSIA 124
Query: 116 SNNYSNLTTSEWLQNFNSYPSNDIPDTGT-LNVTVNCSCGNSDVSKDYGLFITYPLRPED 174
+ Y L ++E +Q NS + D GT L + + C+C N + ++++Y ++ D
Sbjct: 125 DSVYGGLVSAEQIQEANSVNDPSLLDVGTSLVIPLPCACFNGTDNSLPAVYLSYVVKEID 184
Query: 175 SLELISNK 182
+L I+ +
Sbjct: 185 TLVGIARR 192
>At2g33580 putative protein kinase
Length = 664
Score = 37.0 bits (84), Expect = 0.032
Identities = 26/108 (24%), Positives = 46/108 (42%), Gaps = 8/108 (7%)
Query: 81 TRVNVPFPCDCIHDEFLGHIFQYQVATK-------DTYLSVASNNYSNLTTSEWLQNFNS 133
TR V P +C G +Q+ +TY SVA++ Y L+T + + + N
Sbjct: 102 TRELVVIPANCSCSSSSGGFYQHNATYNLSGNRGDETYFSVANDTYQALSTCQAMMSQNR 161
Query: 134 YPSNDIPDTGTLNVTVNCSCGNS-DVSKDYGLFITYPLRPEDSLELIS 180
Y + L V + C+C + + + +TY + DS+ I+
Sbjct: 162 YGERQLTPGLNLLVPLRCACPTAKQTTAGFKYLLTYLVAMGDSISGIA 209
>At3g01840 putative protein kinase
Length = 654
Score = 35.0 bits (79), Expect = 0.12
Identities = 30/104 (28%), Positives = 48/104 (45%), Gaps = 7/104 (6%)
Query: 85 VPFPCDCIHDEFLGHIFQYQVATKDTYLSVASNNYSNLTTSEWLQNFNSYPSND-IPDTG 143
+P C C + + + V DT+ SV S + LTT ++ N + S D + D
Sbjct: 96 IPIECRCNGSIYEASLIKNCVKG-DTFRSV-SQSLQGLTTCLSIREKNPHISEDKLGDNI 153
Query: 144 TLNVTVNCSCGNSDVSKDYGLFITYPLRPEDSLELIS---NKTE 184
L + + CSC VS + +TYP+ DS+ ++ N TE
Sbjct: 154 KLRLAIRCSCPQEGVS-NASFLVTYPVGVRDSVSSLAVRFNTTE 196
>At2g23770 putative protein kinase
Length = 612
Score = 31.2 bits (69), Expect = 1.8
Identities = 16/49 (32%), Positives = 23/49 (46%), Gaps = 4/49 (8%)
Query: 135 PSNDIPDTGTLNVTVNCSCGNSDVSKDYGLFITYPLRPEDSLELISNKT 183
PS P + + + CSC D + ITY ++P DS I+N T
Sbjct: 92 PSTSFPSGQQVIIPLTCSCTGDDSQSN----ITYTIQPNDSYFAIANDT 136
>At3g45360 putative protein
Length = 126
Score = 29.6 bits (65), Expect = 5.2
Identities = 26/82 (31%), Positives = 37/82 (44%), Gaps = 6/82 (7%)
Query: 112 LSVASNNYSNLTTSEWLQNFNSYPSNDIPDTGTLNVTVNCSCGNSDVSKDYGLFITYPLR 171
+ ++ N S+ + SE F+ Y N I D G + C CG DV K G + L
Sbjct: 1 MPISFNGSSDSSPSEKSDGFDYY--NAIRD-GNWGIPTKCYCGR-DVEK--GNVVGGNLH 54
Query: 172 PEDSLELISNKTEIDAELLQKY 193
+ N TE+D E LQK+
Sbjct: 55 GKTRFRCPLNLTEVDGEHLQKW 76
>At1g67260 hypothetical protein
Length = 359
Score = 29.6 bits (65), Expect = 5.2
Identities = 17/69 (24%), Positives = 33/69 (47%), Gaps = 6/69 (8%)
Query: 164 LFITYPLRPEDSLELISNKTEIDAELLQKYNPGVNFSQGSGLVYIPGKDQNRNYVPFHTR 223
LF P+ D+ E I + ++ Q YN Q +G++ + + + NY F +
Sbjct: 269 LFYKEPIEEFDNQESILTNMTLPTKMGQSYN------QNNGILMLVDQSSSSNYNTFLPQ 322
Query: 224 SIRWSYNWN 232
++ +SY+ N
Sbjct: 323 NLDYSYDQN 331
>At4g05250 putative protein (ubiquitin like)
Length = 318
Score = 29.3 bits (64), Expect = 6.8
Identities = 28/104 (26%), Positives = 43/104 (40%), Gaps = 16/104 (15%)
Query: 110 TYLSVASNNYSNLTTSEWLQNFNSYPSNDIPDTGTLNVTVNCSCGNSD------------ 157
T + + SNN + E FNSY D ++ ++ NSD
Sbjct: 146 TSMELVSNNSDQVPQPEKSTPFNSYEEIDYIQDYPVSTSMELVSNNSDQVPPPEKSPPLN 205
Query: 158 VSKDYGLFITYPLRPEDSLELISNKTEIDAELLQKYNPGVNFSQ 201
SK+ F YP+ S+EL+SN ID + +P +N S+
Sbjct: 206 SSKEIDYFQEYPV--STSMELVSN--NIDQVPQPEKSPPLNSSK 245
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.351 0.153 0.577
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,920,678
Number of Sequences: 26719
Number of extensions: 510964
Number of successful extensions: 1875
Number of sequences better than 10.0: 9
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 1867
Number of HSP's gapped (non-prelim): 10
length of query: 617
length of database: 11,318,596
effective HSP length: 105
effective length of query: 512
effective length of database: 8,513,101
effective search space: 4358707712
effective search space used: 4358707712
T: 11
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.9 bits)
S2: 63 (28.9 bits)
Medicago: description of AC147960.5


