

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147959.10 + phase: 1 /pseudo
(378 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
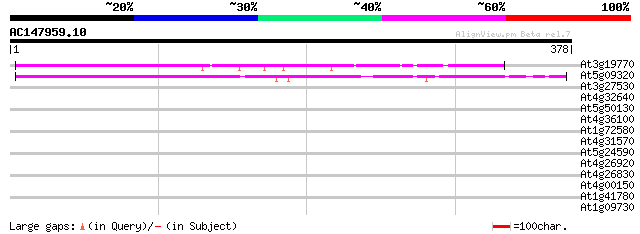
Score E
Sequences producing significant alignments: (bits) Value
At3g19770 unknown protein 197 1e-50
At5g09320 unknown protein 191 7e-49
At3g27530 unknown protein 32 0.75
At4g32640 putative protein 30 1.7
At5g50130 ribitol dehydrogenase-like 30 2.8
At4g36100 hypothetical protein 30 2.8
At1g72580 hypothetical protein 30 2.8
At4g31570 putative protein 29 3.7
At5g24590 NAC2-like protein 29 4.8
At4g26920 putative homeodomain protein 28 8.3
At4g26830 putative beta-1,3-glucanase 28 8.3
At4g00150 scarecrow-like 6 (SCL6) 28 8.3
At1g41780 hypothetical protein 28 8.3
At1g09730 unknown protein 28 8.3
>At3g19770 unknown protein
Length = 472
Score = 197 bits (500), Expect = 1e-50
Identities = 140/349 (40%), Positives = 191/349 (54%), Gaps = 28/349 (8%)
Query: 5 VQDFFLSMEVAIRNHPLWATATEEDIDCAMEGLEKYIMTKLFSRTFAASPEDAKIDHEIS 64
VQ+FF ME A R HPLW+ +EE++D A +GLEKY+MTKLF+R FA++ E+ D ++
Sbjct: 2 VQEFFSKMEAAFRAHPLWSGCSEEELDSAGDGLEKYVMTKLFTRVFASNTEEVIADEKLF 61
Query: 65 EKISLLQTFLKPEHLDIPPVLHNEASWLLAEKELQKINAFKAPQEKLSTIMNCCRVINNL 124
+K+SL+Q F+ PE+LDI P NE+SWLLA+KELQKIN +KAP++KL I+NCC+VINNL
Sbjct: 62 QKMSLVQQFISPENLDIQPTFQNESSWLLAQKELQKINMYKAPRDKLVCILNCCKVINNL 121
Query: 125 LLNA--AMSEYVPAGADDFIPVLIYVTIKAR------LALLVQTSNSLSCIEGK-----Q 171
LLNA A +E P GAD+F+PVLIYVTIKA L +Q S + G+
Sbjct: 122 LLNASIASNENAP-GADEFLPVLIYVTIKANPPQLHSNLLYIQRYRRESKLVGEAAYFFT 180
Query: 172 NSSQKLSIISQI---LCQLRHL*LN*ILNPSQLMKSNLRSACKQPNW----PRKSQVNYT 224
N S IS I L + ++ S L S Q P + +
Sbjct: 181 NILSAESFISNIDAKSISLDEAEFEKNMESARARISGLDSQTYQTGHGSAPPPRDESTLQ 240
Query: 225 LHVKLNKK*RTRAMFQTKCITNLILETK*TILGSLTKLSLKK*KIFEL*FQPLSQSHLNY 284
LN K R +FQ+K +L + + S T + K I +L + + L
Sbjct: 241 KTQSLNPK-RENTLFQSKSSDSLSGTNELLNINSETPMK-KAESISDL--ENKGATLLKD 296
Query: 285 HAEFHVLQHGTNYPYMEAESKDLAMEDVDILLNHYKDLVAKYTIICKAI 333
V Q YPY+ A + DL + DV+ LLN YK LV KY + K +
Sbjct: 297 TEPSKVFQ---EYPYIFASAGDLRIGDVEGLLNSYKQLVFKYVCLTKGL 342
>At5g09320 unknown protein
Length = 712
Score = 191 bits (484), Expect = 7e-49
Identities = 140/386 (36%), Positives = 206/386 (53%), Gaps = 37/386 (9%)
Query: 5 VQDFFLSMEVAIRNHPLWATATEEDIDCAMEGLEKYIMTKLFSRTFAASPEDAKIDHEIS 64
VQDFF ME A R HPLW+ +++++D A +GLEKY+MTKLF R FA++ ED D ++
Sbjct: 47 VQDFFYKMESAFRAHPLWSGCSDDELDNAGDGLEKYVMTKLFPRVFASNTEDVISDEKLF 106
Query: 65 EKISLLQTFLKPEHLDIPPVLHNEASWLLAEKELQKINAFKAPQEKLSTIMNCCRVINNL 124
+KISL+Q F+ PE+LDI P N+ SWLLA+KELQKIN + AP++KL I+ CC+VINNL
Sbjct: 107 QKISLVQQFISPENLDIQPTFQNQTSWLLAQKELQKINMYNAPRDKLMCILRCCKVINNL 166
Query: 125 LLNAAM-SEYVPAGADDFIPVLIYVTIKARLALLVQTSNSLSCIEGKQNSSQKLS----I 179
LLNA++ S GAD F+PVLIYVTIKA Q ++L I+ + S+ + +
Sbjct: 167 LLNASIASNQNEPGADQFLPVLIYVTIKANPP---QFHSNLLYIQRYRRQSKLVGEAGYL 223
Query: 180 ISQILCQ---LRHL*LN*ILNPSQLMKSNLRSACKQPNWPRKSQVNYTLHVKLNKK*RTR 236
+ IL + ++ + ++ ++SA + + P L T+
Sbjct: 224 FTNILSAESFISNIDAKSLSMDEADFETKMKSAHARLSGPGSQSYQTDHGAALPTAHNTK 283
Query: 237 AMFQTKCITNLILETK*TILGSLTKLSLKK*KIFEL*FQPLSQ-------SHLNYHAEFH 289
N++L TK T S T +L + I + P++ + LN +E
Sbjct: 284 R-------ENMLLHTKSTDSFSGTNETLSETPIKKA--DPITDLENKGAATLLNDRSE-- 332
Query: 290 VLQHGTNYPYMEAESKDLAMEDVDILLNHYKDLVAKYTIICKAINYLSMSEKEPLLHQLE 349
+ YPYM A DL + DV+ LLN+YK LV KY + K + + P + L
Sbjct: 333 ATKIFQEYPYMFASVGDLKIGDVEDLLNNYKQLVFKYVCLSKGLG--DATSLTPCISPL- 389
Query: 350 MQGEGSMLSECHGINTNTNDRTTSHE 375
+ S +SE H T ++D T E
Sbjct: 390 ---QASKVSENH--TTLSSDFQTKSE 410
>At3g27530 unknown protein
Length = 938
Score = 31.6 bits (70), Expect = 0.75
Identities = 27/105 (25%), Positives = 49/105 (45%), Gaps = 20/105 (19%)
Query: 84 VLHNEASWLLAE-----KELQKINAFKAPQEKLSTIM-------------NCCRVINNLL 125
V+ NEA LL +E+QKI F+ EK+ +I+ +C ++NNLL
Sbjct: 207 VIRNEALLLLTHLTREAEEIQKIVVFEGAFEKIFSIIKEEGGSDGDVVVQDCLELLNNLL 266
Query: 126 LNAAMSEYVPAGADDFIPVLIYVTIKARLALLVQ--TSNSLSCIE 168
+++ ++ + F P++ + ++ Q T N LS +E
Sbjct: 267 RSSSSNQILLRETMGFEPIISILKLRGITYKFTQQKTVNLLSALE 311
>At4g32640 putative protein
Length = 1069
Score = 30.4 bits (67), Expect = 1.7
Identities = 27/114 (23%), Positives = 47/114 (40%), Gaps = 5/114 (4%)
Query: 79 LDIPPVLHNEASWLLAEKE----LQKINAFKAPQEKLSTIMNCCRVINNLLLNAAMSEYV 134
L PP L+N+ W + + + ++ + Q + + C R+ ++ L A +
Sbjct: 728 LSDPPKLYNDLKWNITRPQGFEAVMRVRCSQGIQVQEYSGNFCKRIPTDIDLPAHDDKLQ 787
Query: 135 PAGADDFIPVLIYVTIKARLALLVQTSNSLSCIEGKQNSSQKLSIISQILCQLR 188
F L+Y TI + V T+ SLSC N + + SQ C L+
Sbjct: 788 DGAECAFQCALLYTTIYGERRIRV-TTLSLSCTNMLSNLFRAADLDSQFACMLK 840
>At5g50130 ribitol dehydrogenase-like
Length = 339
Score = 29.6 bits (65), Expect = 2.8
Identities = 23/94 (24%), Positives = 44/94 (46%), Gaps = 13/94 (13%)
Query: 26 TEEDIDC--AMEGLEKYIMTKLFSRTFAASPEDAKIDHEISEKISLLQTFLKPEHLDIPP 83
+EE I+ A L Y++T++ + E + I+ I S++ ++KP+ P
Sbjct: 133 SEEKIELTFATNFLGHYLLTEMLIEKMIDTAEKSGIEGRIINLSSVIHNWVKPDCFSFPK 192
Query: 84 VLHNEASWLLAEKELQKINAFKA-PQEKLSTIMN 116
+LH + + N +A Q KL+TI++
Sbjct: 193 LLH----------PISRYNGTRAYAQSKLATILH 216
>At4g36100 hypothetical protein
Length = 396
Score = 29.6 bits (65), Expect = 2.8
Identities = 19/73 (26%), Positives = 36/73 (49%), Gaps = 8/73 (10%)
Query: 53 SPEDAKIDHEISEKISLLQTF-------LKPEHLDIPPVLHNEASWLLAE-KELQKINAF 104
SP+D+++ EI E+ SLL+T+ L + LD E ++ + ++
Sbjct: 77 SPQDSRLAAEIQEQQSLLKTYEVMVKRSLMEQSLDAYDEKEKEMMMMIGSINRTELLSVL 136
Query: 105 KAPQEKLSTIMNC 117
KA + KL +++C
Sbjct: 137 KAKERKLLLLLSC 149
>At1g72580 hypothetical protein
Length = 98
Score = 29.6 bits (65), Expect = 2.8
Identities = 23/84 (27%), Positives = 38/84 (44%), Gaps = 3/84 (3%)
Query: 62 EISEKISLLQTFLKPEHLDIPPVLHN-EASWLLAEKELQKINAFKAPQEKLSTIMNCCRV 120
+++E +L QT E D+ VL E ++L EK EK + +MNC ++
Sbjct: 9 DVAELATLEQTKKNVE--DVEEVLKKLEQTYLGKEKVTDDTLLSDEEMEKFNQMMNCVKL 66
Query: 121 INNLLLNAAMSEYVPAGADDFIPV 144
+NN L S + G + + V
Sbjct: 67 LNNCLTKIGQSLPLKEGERELVEV 90
>At4g31570 putative protein
Length = 2712
Score = 29.3 bits (64), Expect = 3.7
Identities = 19/75 (25%), Positives = 39/75 (51%), Gaps = 14/75 (18%)
Query: 24 TATEEDIDCAMEGLEKYIMTKLFSRTFAASPEDAKIDHEISEKISLLQTFLKP-EHLDIP 82
T++E+D+ E L+ + + + ++ ++ + E+ +L+Q + K E++DIP
Sbjct: 1518 TSSEDDLRIKFEELK--------GKFYGLAEQNEMLEQSLMERNTLVQRWEKLLENIDIP 1569
Query: 83 PVLH-----NEASWL 92
P LH N+ WL
Sbjct: 1570 PQLHSMEVENKIEWL 1584
>At5g24590 NAC2-like protein
Length = 451
Score = 28.9 bits (63), Expect = 4.8
Identities = 19/64 (29%), Positives = 32/64 (49%), Gaps = 5/64 (7%)
Query: 25 ATEEDIDCAMEGLEKYIMTKLFSR----TFAASPEDAKIDHEISEKISLLQTFLKPEHLD 80
ATE+D+D G +++ KLF + AA PE++K E+ +S + E +
Sbjct: 141 ATEKDLDGTKSGQNPFVVCKLFKKQDIVNGAAEPEESK-SCEVEPAVSSPTVVDEVEMSE 199
Query: 81 IPPV 84
+ PV
Sbjct: 200 VSPV 203
>At4g26920 putative homeodomain protein
Length = 461
Score = 28.1 bits (61), Expect = 8.3
Identities = 21/82 (25%), Positives = 36/82 (43%), Gaps = 7/82 (8%)
Query: 107 PQEKLSTIMNCCRVINNLLLNAAMSEYVPAGADDFIPVLIYVT---IKARLALLVQTSNS 163
P K+ + C R+ N+ + A +S Y+ + +DD P + I +A + + +NS
Sbjct: 180 PTRKVKVLRYCHRIANDTWIIADISMYLSSYSDDLRPEFLRFPSGFIIKHVARIFRVTNS 239
Query: 164 LSCIEGKQNSSQKLSIISQILC 185
GK N Q + I C
Sbjct: 240 ----AGKNNLLQASKRLVHIFC 257
>At4g26830 putative beta-1,3-glucanase
Length = 448
Score = 28.1 bits (61), Expect = 8.3
Identities = 16/44 (36%), Positives = 25/44 (56%)
Query: 121 INNLLLNAAMSEYVPAGADDFIPVLIYVTIKARLALLVQTSNSL 164
I++ + +A++ P + F P LI +K LALL QTS+ L
Sbjct: 151 ISSPIALSALANSYPPSSGSFKPELIEPVVKPMLALLQQTSSYL 194
>At4g00150 scarecrow-like 6 (SCL6)
Length = 558
Score = 28.1 bits (61), Expect = 8.3
Identities = 15/55 (27%), Positives = 26/55 (47%)
Query: 310 EDVDILLNHYKDLVAKYTIICKAINYLSMSEKEPLLHQLEMQGEGSMLSECHGIN 364
E ++ LL++ + Y++I K Y S SE P+L ++L HG +
Sbjct: 251 EALNNLLHNVSQTLNPYSLIFKIAAYKSFSEISPVLQFANFTSNQALLESFHGFH 305
>At1g41780 hypothetical protein
Length = 246
Score = 28.1 bits (61), Expect = 8.3
Identities = 19/68 (27%), Positives = 29/68 (41%), Gaps = 10/68 (14%)
Query: 23 ATATEEDIDCAMEGLEKYIMTKLFSRTFAASPEDAKID----------HEISEKISLLQT 72
A ++ +IDC +E T+ S + PE +I+ H S KIS+ +
Sbjct: 107 AVSSVPNIDCILEVYHNTPFTRKISNAIISDPEKLRIEYFSGSSDPKGHLKSFKISVARA 166
Query: 73 FLKPEHLD 80
KPE D
Sbjct: 167 KFKPEESD 174
>At1g09730 unknown protein
Length = 865
Score = 28.1 bits (61), Expect = 8.3
Identities = 12/23 (52%), Positives = 15/23 (65%)
Query: 206 LRSACKQPNWPRKSQVNYTLHVK 228
L+ A K+ NWP K Q +LHVK
Sbjct: 309 LKIAVKEHNWPNKQQKINSLHVK 331
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.328 0.139 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,615,249
Number of Sequences: 26719
Number of extensions: 287743
Number of successful extensions: 848
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 838
Number of HSP's gapped (non-prelim): 17
length of query: 378
length of database: 11,318,596
effective HSP length: 101
effective length of query: 277
effective length of database: 8,619,977
effective search space: 2387733629
effective search space used: 2387733629
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC147959.10


