

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
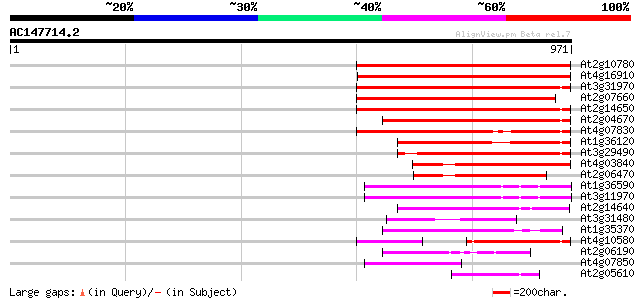
Query= AC147714.2 - phase: 0 /pseudo
(971 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At2g10780 pseudogene 376 e-104
At4g16910 retrotransposon like protein 365 e-101
At3g31970 hypothetical protein 354 2e-97
At2g07660 putative retroelement pol polyprotein 353 3e-97
At2g14650 putative retroelement pol polyprotein 348 6e-96
At2g04670 putative retroelement pol polyprotein 332 6e-91
At4g07830 putative reverse transcriptase 294 2e-79
At1g36120 putative reverse transcriptase gb|AAD22339.1 284 2e-76
At3g29490 hypothetical protein 278 1e-74
At4g03840 putative transposon protein 271 2e-72
At2g06470 putative retroelement pol polyprotein 238 2e-62
At1g36590 hypothetical protein 204 3e-52
At3g11970 hypothetical protein 203 4e-52
At2g14640 putative retroelement pol polyprotein 172 1e-42
At3g31480 hypothetical protein 167 3e-41
At1g35370 hypothetical protein 164 2e-40
At4g10580 putative reverse-transcriptase -like protein 140 3e-33
At2g06190 putative Ty3-gypsy-like retroelement pol polyprotein 124 3e-28
At4g07850 putative polyprotein 116 7e-26
At2g05610 putative retroelement pol polyprotein 89 1e-17
>At2g10780 pseudogene
Length = 1611
Score = 376 bits (966), Expect = e-104
Identities = 187/372 (50%), Positives = 253/372 (67%), Gaps = 2/372 (0%)
Query: 601 VESEHQNLLG*WVLLDVPXWKWDSISM--IL*RVAKYX*RDXRIGGILXR*RKSAHFLXX 658
V++EHQ G L +P WKWD I+M + + + ++ R KSAHF+
Sbjct: 1211 VKAEHQVPSGLLQNLPIPEWKWDHITMDFVTGLPTGIKSKHNAVWVVVDRLTKSAHFMAI 1270
Query: 659 NISFPVSQXXEIYIKEIVKCMGVPSSIVSDRDPRFTSRFWKSLQEALGSKLRLSSAYHPQ 718
+ E YI EIV+ G+P SIVSDRD RFTS+FWK+ Q+ALG+++ LS+AYHPQ
Sbjct: 1271 SDKDGAEIIAEKYIDEIVRLHGIPVSIVSDRDTRFTSKFWKAFQKALGTRVNLSTAYHPQ 1330
Query: 719 TDGQSERTIQSLEDLLRICVLEQGGTWDSHLPLIEFTYNNSYHSSIGMAPFEALYGRRCR 778
TD QSERTIQ+LED+LR CVL+ GG W+ +L L+EF YNNS+ +SIGM+P+EALYGR CR
Sbjct: 1331 TDEQSERTIQTLEDMLRACVLDWGGNWEKYLRLVEFAYNNSFQASIGMSPYEALYGRACR 1390
Query: 779 TPLCWFESGERVVLGPEIVQQTTEKVQMIQEKMKASQSRQKSYHDKRRKDLEFQEGDHVF 838
TPLCW GER + GP IV +TTE+++ ++ K+K +Q RQKSY +KRRK+LEFQ GD V+
Sbjct: 1391 TPLCWTPVGERRLFGPTIVDETTERMKFLKIKLKEAQDRQKSYANKRRKELEFQVGDLVY 1450
Query: 839 LRVTPMTGVXRALKSKKLTPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKY 898
L+ G R KKL+P+++GPY+++ERVG VAY++ LPP L+ HNVFHVSQLRK
Sbjct: 1451 LKAMTYKGAGRFTSRKKLSPRYVGPYKVIERVGAVAYKLDLPPKLNAFHNVFHVSQLRKC 1510
Query: 899 VPDPSHVIQSDDVQVRDNLTVETLPVRIDDRKVKMLRGKEIPLVRVVWTGATSESLTWEL 958
+ D ++ +++N+TVE PVRI DR K RGK L++V+W E TWE
Sbjct: 1511 LSDQEESVEDIPPGLKENMTVEAWPVRIMDRMTKGTRGKARDLLKVLWNCRGREEYTWET 1570
Query: 959 ESKMLESYPELF 970
E+KM ++PE F
Sbjct: 1571 ENKMKANFPEWF 1582
>At4g16910 retrotransposon like protein
Length = 687
Score = 365 bits (938), Expect = e-101
Identities = 181/371 (48%), Positives = 247/371 (65%), Gaps = 2/371 (0%)
Query: 602 ESEHQNLLG*WVLLDVPXWKWDSISM--IL*RVAKYX*RDXRIGGILXR*RKSAHFLXXN 659
+ EHQ G L +P WKWD I M + + + ++ R KSAHF+ +
Sbjct: 296 KEEHQVPSGMLQNLPIPEWKWDHIMMDFVTGLPTGIKSKHNAVWVVVDRLTKSAHFMAIS 355
Query: 660 ISFPVSQXXEIYIKEIVKCMGVPSSIVSDRDPRFTSRFWKSLQEALGSKLRLSSAYHPQT 719
E YI EIV+ G+P SIVSDRD RFTS+FWK Q+ LG+++ LS+AYHPQT
Sbjct: 356 DKDAAEIIAEKYIDEIVRLHGIPVSIVSDRDTRFTSKFWKPFQKVLGTRVNLSTAYHPQT 415
Query: 720 DGQSERTIQSLEDLLRICVLEQGGTWDSHLPLIEFTYNNSYHSSIGMAPFEALYGRRCRT 779
DGQSERTIQ+LED+LR CVL+ GG W+ +L L+EF YNNS+ +SIGM+P+EALYGR RT
Sbjct: 416 DGQSERTIQTLEDMLRACVLDWGGNWEKYLRLVEFAYNNSFQASIGMSPYEALYGRAGRT 475
Query: 780 PLCWFESGERVVLGPEIVQQTTEKVQMIQEKMKASQSRQKSYHDKRRKDLEFQEGDHVFL 839
PLCW GER + GP +V +TT+K++ ++ K+K +Q RQKSY +KRRK+LEFQ GD V+L
Sbjct: 476 PLCWTPVGERRLFGPAVVDETTKKMKFLKIKLKEAQDRQKSYANKRRKELEFQVGDLVYL 535
Query: 840 RVTPMTGVXRALKSKKLTPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYV 899
+ G R KKL P+++GPY+++ERVG VAY++ LPP L HNVFHVSQLRK +
Sbjct: 536 KAMTYKGAGRFTSRKKLRPRYVGPYKVIERVGAVAYKLDLPPKLDAFHNVFHVSQLRKCL 595
Query: 900 PDPSHVIQSDDVQVRDNLTVETLPVRIDDRKVKMLRGKEIPLVRVVWTGATSESLTWELE 959
+ ++ +++N+TVE PVRI D+ K RGK + L++++W E TWE E
Sbjct: 596 SEQEESMEDVPPGLKENMTVEAWPVRIMDQMKKGTRGKSMDLLKILWNCGGREEYTWETE 655
Query: 960 SKMLESYPELF 970
+KM ++PE F
Sbjct: 656 TKMKANFPEWF 666
>At3g31970 hypothetical protein
Length = 1329
Score = 354 bits (908), Expect = 2e-97
Identities = 181/372 (48%), Positives = 252/372 (67%), Gaps = 4/372 (1%)
Query: 601 VESEHQNLLG*WVLLDVPXWKWDSISMIL*RVAKYX*RDXRIGGILXR*RKSAHFLXXNI 660
V++EHQ G L + WKWD I+M I I+ R KSAHFL
Sbjct: 954 VKAEHQVPGGLLQSLPILEWKWDFITMDFVVGLPVSRTKDAIWVIVDRLTKSAHFLAIRK 1013
Query: 661 SFPVSQXXEIYIKEIVKCMGVPSSIVSDRDPRFTSRFWKSLQEALGSKLRLSSAYHPQTD 720
+ + Y+ EIV+ GVP SIVSDRD +FTS FW++ Q +G+K+++S+AYHPQTD
Sbjct: 1014 TDGAVLLAKKYVSEIVELHGVPVSIVSDRDSKFTSAFWRAFQGEMGTKVQMSTAYHPQTD 1073
Query: 721 GQSERTIQSLEDLLRICVLEQGGTWDSHLPLIEFTYNNSYHSSIGMAPFEALYGRRCRTP 780
GQSERTIQ+LED+LR+CVL++GG W HL L+EF YNNSY +SI MAPFEALYGR CRTP
Sbjct: 1074 GQSERTIQTLEDMLRMCVLDRGGHWADHLSLVEFAYNNSYQASIRMAPFEALYGRPCRTP 1133
Query: 781 LCWFESGERVVLGPEIVQQTTEKVQMIQEKMKASQSRQKSYHDKRRKDLEFQEGDHVFLR 840
LCW + GER + G + V +TTE++++++ MK +Q RQ+SY DKRR++LEF+ GD V+L+
Sbjct: 1134 LCWTQVGERSIYGADYVLETTERIRVLKLNMKEAQDRQRSYADKRRRELEFEVGDRVYLK 1193
Query: 841 VTPMTGVXRALKSKKLTPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVP 900
+ + G R++ KLTP+++GP++I+ERVG VAYR+ LP + H VFHVS LRK +
Sbjct: 1194 MAMLRGPNRSISETKLTPRYMGPFRIVERVGPVAYRLELPDVMRAFHKVFHVSMLRKCLH 1253
Query: 901 DPSHVIQSDDVQVRDNLTVETLPVRIDDRKVKMLRGKEIPLVRVVWT--GATSESLTWEL 958
V+ ++ N+T+E PVRI +R++K LR K+IPL++V+W G T E TWE
Sbjct: 1254 KDDEVLAKILEDLQPNMTLEARPVRILERRIKELRRKKIPLIKVLWNCDGVTEE--TWEP 1311
Query: 959 ESKMLESYPELF 970
E++M S+ + F
Sbjct: 1312 EARMKASFKKWF 1323
>At2g07660 putative retroelement pol polyprotein
Length = 949
Score = 353 bits (905), Expect = 3e-97
Identities = 176/347 (50%), Positives = 239/347 (68%), Gaps = 2/347 (0%)
Query: 601 VESEHQNLLG*WVLLDVPXWKWDSISM--IL*RVAKYX*RDXRIGGILXR*RKSAHFLXX 658
V++EHQ G L + WKWD I+M + + + ++ R KSAHF+
Sbjct: 603 VKAEHQVPSGLLQNLPISEWKWDHITMDFVTRLPTGIKSKHNAVWVVVDRLTKSAHFMAI 662
Query: 659 NISFPVSQXXEIYIKEIVKCMGVPSSIVSDRDPRFTSRFWKSLQEALGSKLRLSSAYHPQ 718
+ E YI EI++ G+P SIVSDRD RFTS+FW + Q+ALG+++ LS+AYHPQ
Sbjct: 663 SDKDGAEIIAEKYIDEIMRLHGIPVSIVSDRDTRFTSKFWNAFQKALGTRVNLSTAYHPQ 722
Query: 719 TDGQSERTIQSLEDLLRICVLEQGGTWDSHLPLIEFTYNNSYHSSIGMAPFEALYGRRCR 778
TDGQSERTIQ+LED+LR CVL+ GG W+ +L LIEF YNNS+ +SIGM+P+EALYGR CR
Sbjct: 723 TDGQSERTIQTLEDMLRACVLDWGGNWEKYLRLIEFAYNNSFQASIGMSPYEALYGRACR 782
Query: 779 TPLCWFESGERVVLGPEIVQQTTEKVQMIQEKMKASQSRQKSYHDKRRKDLEFQEGDHVF 838
TPLCW GER + GP IV +TTE+++ ++ K+K +Q RQKSY +KRRK+LEFQ GD V+
Sbjct: 783 TPLCWTPVGERRLFGPTIVDETTERMKFLKIKLKEAQDRQKSYANKRRKELEFQVGDLVY 842
Query: 839 LRVTPMTGVXRALKSKKLTPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKY 898
L+ G R KKL+P+++GPY+++ERVG VAY++ LPP L+ HNVFHVSQLRKY
Sbjct: 843 LKAMTYKGAGRFTSRKKLSPRYVGPYKVIERVGAVAYKLDLPPKLNVFHNVFHVSQLRKY 902
Query: 899 VPDPSHVIQSDDVQVRDNLTVETLPVRIDDRKVKMLRGKEIPLVRVV 945
+ D ++ +++N+TVE PVRI DR K RGK L++V+
Sbjct: 903 LSDQEESVEDIPPGLKENMTVEAWPVRIMDRMSKGTRGKSRDLLKVL 949
>At2g14650 putative retroelement pol polyprotein
Length = 1328
Score = 348 bits (894), Expect = 6e-96
Identities = 178/372 (47%), Positives = 251/372 (66%), Gaps = 4/372 (1%)
Query: 601 VESEHQNLLG*WVLLDVPXWKWDSISMIL*RVAKYX*RDXRIGGILXR*RKSAHFLXXNI 660
V++EHQ G L +P WKWD I++ I I+ R KSAHFL
Sbjct: 956 VKAEHQVPGGMLQSLPIPEWKWDFITIDFVVGLPVSRTKDAIWVIVDRLTKSAHFLAIRK 1015
Query: 661 SFPVSQXXEIYIKEIVKCMGVPSSIVSDRDPRFTSRFWKSLQEALGSKLRLSSAYHPQTD 720
+ + + Y+ EIVK GVP SIVSDRD +FTS FW++ Q +G+K+++S+AYHPQT
Sbjct: 1016 TDGAAVLAKKYVSEIVKLHGVPVSIVSDRDSKFTSAFWRAFQAEMGTKVQMSTAYHPQTY 1075
Query: 721 GQSERTIQSLEDLLRICVLEQGGTWDSHLPLIEFTYNNSYHSSIGMAPFEALYGRRCRTP 780
GQSERTIQ+LED+LR+CVL+ GG W HL L+EF YNNSY +SIGMAPFEALY R CRTP
Sbjct: 1076 GQSERTIQTLEDMLRMCVLDWGGHWADHLSLVEFAYNNSYPASIGMAPFEALYERPCRTP 1135
Query: 781 LCWFESGERVVLGPEIVQQTTEKVQMIQEKMKASQSRQKSYHDKRRKDLEFQEGDHVFLR 840
LC + GER + G + VQ+TTE++++++ MK +Q RQ+SY DKRR++LEF+ GD V+L+
Sbjct: 1136 LCLTQVGERSIYGADYVQETTERIRVLKLNMKEAQDRQRSYADKRRRELEFEVGDRVYLK 1195
Query: 841 VTPMTGVXRALKSKKLTPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVP 900
+ + G R++ KL+P+++GP++I+ERVG VAYR+ LP + H VFHVS LRK +
Sbjct: 1196 MAMLRGPNRSISETKLSPRYMGPFRIVERVGPVAYRLELPDVMRAFHKVFHVSMLRKCLH 1255
Query: 901 DPSHVIQSDDVQVRDNLTVETLPVRIDDRKVKMLRGKEIPLVRVVW--TGATSESLTWEL 958
V+ ++ N+T+E PVR+ +R++K LR K+IPL++V+W G T E TWE
Sbjct: 1256 KDDEVLAKIPEDLQPNMTLEARPVRVLERRIKELRRKKIPLIKVLWDCDGVTKE--TWEP 1313
Query: 959 ESKMLESYPELF 970
E++M + + F
Sbjct: 1314 EARMKARFKKWF 1325
>At2g04670 putative retroelement pol polyprotein
Length = 1411
Score = 332 bits (851), Expect = 6e-91
Identities = 163/328 (49%), Positives = 232/328 (70%), Gaps = 4/328 (1%)
Query: 645 ILXR*RKSAHFLXXNISFPVSQXXEIYIKEIVKCMGVPSSIVSDRDPRFTSRFWKSLQEA 704
I+ R KSAHFL + + + Y+ EIVK GVP SIVSDRD +FT FW++ Q
Sbjct: 1080 IMDRLTKSAHFLAIRKTDGAAVLAKKYVSEIVKLHGVPVSIVSDRDSKFTFAFWRAFQAK 1139
Query: 705 LGSKLRLSSAYHPQTDGQSERTIQSLEDLLRICVLEQGGTWDSHLPLIEFTYNNSYHSSI 764
+G+K+++S+AYHPQTDGQSERTIQ+LED+LR+CVL+ GG W HL L+EF YNNSY +SI
Sbjct: 1140 MGTKVQMSTAYHPQTDGQSERTIQTLEDMLRMCVLDWGGHWADHLSLVEFAYNNSYQASI 1199
Query: 765 GMAPFEALYGRRCRTPLCWFESGERVVLGPEIVQQTTEKVQMIQEKMKASQSRQKSYHDK 824
GMAPFEALYGR C TPL W + ER + G + VQ+TTE++++++ MK +Q+RQ+SY DK
Sbjct: 1200 GMAPFEALYGRPCWTPLRWTQVEERSIYGADYVQETTERIRVLKLNMKEAQARQRSYADK 1259
Query: 825 RRKDLEFQEGDHVFLRVTPMTGVXRALKSKKLTPKFIGPYQILERVGTVAYRVGLPPHLS 884
RR++LEF+ GD V+L++ + G R++ KL+P+++GP++I+ERVG VAYR+ LP +
Sbjct: 1260 RRRELEFEVGDRVYLKMAMLRGPNRSILETKLSPRYMGPFRIVERVGPVAYRLELPDVMR 1319
Query: 885 NLHNVFHVSQLRKYVPDPSHVIQSDDVQVRDNLTVETLPVRIDDRKVKMLRGKEIPLVRV 944
H VFHV LRK + V+ ++ N+T+E PVR+ +R++K LR K+IPL++V
Sbjct: 1320 AFHKVFHVLMLRKCLHKDDEVLVKIPEDLQPNMTLEARPVRVLERRIKELRRKKIPLIKV 1379
Query: 945 VW--TGATSESLTWELESKMLESYPELF 970
+W G T E TWE E++M + + F
Sbjct: 1380 LWDCDGVTEE--TWEPEARMKARFKKWF 1405
>At4g07830 putative reverse transcriptase
Length = 611
Score = 294 bits (752), Expect = 2e-79
Identities = 165/372 (44%), Positives = 229/372 (61%), Gaps = 25/372 (6%)
Query: 601 VESEHQNLLG*WVLLDVPXWKWDSISMIL*RVAKYX*RDXRIGGILXR*RKSAHFLXXNI 660
V+ EHQ L +P WKWD I+M I I+ R KSAHFL
Sbjct: 221 VKIEHQVSGSLLQSLPIPEWKWDFITMDFVVGLPVSRTKDAIWVIVDRLTKSAHFLAIRK 280
Query: 661 SFPVSQXXEIYIKEIVKCMGVPSSIVSDRDPRFTSRFWKSLQEALGSKLRLSSAYHPQTD 720
+ + + Y+ EIVK GVP SIVSDRD +FTS FW++ Q +G+K+++S+AYHPQTD
Sbjct: 281 TDGAAVLAKKYVSEIVKLHGVPVSIVSDRDSKFTSAFWRAFQAEMGTKVQMSTAYHPQTD 340
Query: 721 GQSERTIQSLEDLLRICVLEQGGTWDSHLPLIEFTYNNSYHSSIGMAPFEALYGRRCRTP 780
GQSERTIQ+LED+LR+CVL+ G W HL L+EF YNNSY +SIGMAPFE LYGR CRT
Sbjct: 341 GQSERTIQTLEDMLRMCVLDWRGHWADHLSLVEFAYNNSYQASIGMAPFEVLYGRPCRT- 399
Query: 781 LCWFESGERVVLGPEIVQQTTEKVQMIQEKMKASQSRQKSYHDKRRKDLEFQEGDHVFLR 840
LCW + GER + G + VQ+ TE++++++ MK +Q+RQ+SY DKRRK+LEF+ GD V
Sbjct: 400 LCWTQVGERSIYGADYVQEITERIRVLKLNMKEAQNRQRSYADKRRKELEFEVGDSV--- 456
Query: 841 VTPMTGVXRALKSKKLTPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVP 900
+ +S+ +RVG VA+R+ L + H VFHVS LRK +
Sbjct: 457 ----SQDGHVARSE-------------QRVGPVAFRLELSDVMRAFHKVFHVSMLRKCLH 499
Query: 901 DPSHVIQSDDVQVRDNLTVETLPVRIDDRKVKMLRGKEIPLVRVV--WTGATSESLTWEL 958
V+ ++ N+T+E PVR+ +R++K LR K+IPL++V+ G T E TWE
Sbjct: 500 KDDEVLAKIPEDLQPNMTLEARPVRVLERRIKELRRKKIPLIKVLRNCDGVTEE--TWEP 557
Query: 959 ESKMLESYPELF 970
E+++ + + F
Sbjct: 558 EARLKARFKKWF 569
>At1g36120 putative reverse transcriptase gb|AAD22339.1
Length = 1235
Score = 284 bits (726), Expect = 2e-76
Identities = 142/302 (47%), Positives = 202/302 (66%), Gaps = 33/302 (10%)
Query: 671 YIKEIVKCMGVPSSIVSDRDPRFTSRFWKSLQEALGSKLRLSSAYHPQTDGQSERTIQSL 730
Y+ EIVK GVP SI+S RD +FTS FW++ Q +G+K+++S+AYHPQTDGQSERTIQ+L
Sbjct: 959 YVSEIVKLHGVPVSILSHRDSKFTSAFWRAFQVEMGTKVQMSTAYHPQTDGQSERTIQTL 1018
Query: 731 EDLLRICVLEQGGTWDSHLPLIEFTYNNSYHSSIGMAPFEALYGRRCRTPLCWFESGERV 790
ED+L++CVL+ GG W HL L++F YNNSY +SIGMAPFEALYGR CRT LCW + GE+
Sbjct: 1019 EDMLQMCVLDWGGHWADHLSLVKFAYNNSYQASIGMAPFEALYGRPCRTLLCWTQVGEKS 1078
Query: 791 VLGPEIVQQTTEKVQMIQEKMKASQSRQKSYHDKRRKDLEFQEGDHVFLRVTPMTGVXRA 850
+ G + VQ+TTE++++++ MK +Q RQ+SY DKRR++LEF+ G
Sbjct: 1079 IYGADYVQETTERIRVLKLNMKEAQDRQRSYADKRRRELEFEVGT--------------- 1123
Query: 851 LKSKKLTPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPDPSHVIQSDD 910
+I+ERVG VAYR+ LP + HNVFHVS LRK + V+
Sbjct: 1124 --------------EIVERVGPVAYRLELPDVMRAFHNVFHVSMLRKCLHKDDEVLAKIP 1169
Query: 911 VQVRDNLTVETLPVRIDDRKVKMLRGKEIPLVRVVW--TGATSESLTWELESKMLESYPE 968
++ N+T+E PVR+ +R++K +R K+IP+++V+W G T E TWE E+++ + +
Sbjct: 1170 EDLQPNMTLEARPVRVLERRIKEVRRKKIPMIKVLWDCDGVTEE--TWEPEARIKARFKK 1227
Query: 969 LF 970
F
Sbjct: 1228 WF 1229
>At3g29490 hypothetical protein
Length = 438
Score = 278 bits (711), Expect = 1e-74
Identities = 138/302 (45%), Positives = 201/302 (65%), Gaps = 24/302 (7%)
Query: 671 YIKEIVKCMGVPSSIVSDRDPRFTSRFWKSLQEALGSKLRLSSAYHPQTDGQSERTIQSL 730
Y+ EIVK GVP+ +G+K+++S+ YHPQTDGQ ERTIQ+L
Sbjct: 56 YVSEIVKLHGVPAE--------------------MGTKVQMSTPYHPQTDGQFERTIQTL 95
Query: 731 EDLLRICVLEQGGTWDSHLPLIEFTYNNSYHSSIGMAPFEALYGRRCRTPLCWFESGERV 790
ED+LR+CVL+ GG W HL L+EF YNNSY + IGMAPFEALYGR CRTPLCW + GER
Sbjct: 96 EDMLRMCVLDWGGHWADHLSLVEFAYNNSYQAGIGMAPFEALYGRPCRTPLCWTQVGERS 155
Query: 791 VLGPEIVQQTTEKVQMIQEKMKASQSRQKSYHDKRRKDLEFQEGDHVFLRVTPMTGVXRA 850
+ G + VQ+TTE++++++ MK +Q RQ SY DKRR++LEF+ GD V+L++ + G R+
Sbjct: 156 IYGADYVQETTERIRVLKLNMKEAQDRQWSYADKRRRELEFEVGDRVYLKMAMLRGPNRS 215
Query: 851 LKSKKLTPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPDPSHVIQSDD 910
+ KL+ +++GP++I+ERVG VAY + LP + H VFHVS LRK + V+
Sbjct: 216 ISETKLSLRYMGPFRIVERVGPVAYMLELPDVMRAFHKVFHVSMLRKCLHKDDEVLAKIP 275
Query: 911 VQVRDNLTVETLPVRIDDRKVKMLRGKEIPLVRVVW--TGATSESLTWELESKMLESYPE 968
++ N+T+E VR+ +R++K L+ K+I L++V+W G T E TW+ E++M + +
Sbjct: 276 EDLQPNMTLEARQVRVLERRIKELQRKKISLIKVLWDCDGVTEE--TWQPEARMKARFKK 333
Query: 969 LF 970
F
Sbjct: 334 WF 335
>At4g03840 putative transposon protein
Length = 973
Score = 271 bits (692), Expect = 2e-72
Identities = 133/274 (48%), Positives = 182/274 (65%), Gaps = 19/274 (6%)
Query: 697 FWKSLQEALGSKLRLSSAYHPQTDGQSERTIQSLEDLLRICVLEQGGTWDSHLPLIEFTY 756
FWK+ Q+ALG+++ LS+AYHPQTDGQSERTIQ+LED+LR C L+ GG W+ +L L
Sbjct: 690 FWKAFQKALGTRVNLSTAYHPQTDGQSERTIQTLEDMLRACALDWGGNWEKYLRL----- 744
Query: 757 NNSYHSSIGMAPFEALYGRRCRTPLCWFESGERVVLGPEIVQQTTEKVQMIQEKMKASQS 816
ALYGR CRTPLCW GER + GP IV +TTE+++ ++ K+K +Q
Sbjct: 745 --------------ALYGRACRTPLCWTPVGERRLFGPIIVDETTERMKFLKIKLKEAQD 790
Query: 817 RQKSYHDKRRKDLEFQEGDHVFLRVTPMTGVXRALKSKKLTPKFIGPYQILERVGTVAYR 876
RQKSY +KRRK+LEFQ D V+L+ G R KKL+P+++GPY+++ERVG VAY+
Sbjct: 791 RQKSYANKRRKELEFQVEDLVYLKAMTYKGAGRFTSRKKLSPRYVGPYKVIERVGAVAYK 850
Query: 877 VGLPPHLSNLHNVFHVSQLRKYVPDPSHVIQSDDVQVRDNLTVETLPVRIDDRKVKMLRG 936
+ LPP L+ HNVFHVSQLRK + + ++ +++N+TVE PV+I DR K RG
Sbjct: 851 LDLPPKLNAFHNVFHVSQLRKCLSNQEESVEDVPPGLKENMTVEAWPVQIMDRMTKGTRG 910
Query: 937 KEIPLVRVVWTGATSESLTWELESKMLESYPELF 970
K L++V+W E TWE E+KM ++ E F
Sbjct: 911 KSRDLLKVLWNCGGREQYTWETENKMKANFSEWF 944
Score = 29.6 bits (65), Expect = 8.6
Identities = 13/27 (48%), Positives = 17/27 (62%)
Query: 601 VESEHQNLLG*WVLLDVPXWKWDSISM 627
V++EHQ G L +P WKWD I+M
Sbjct: 661 VKAEHQVPSGLLQNLPIPEWKWDHITM 687
>At2g06470 putative retroelement pol polyprotein
Length = 899
Score = 238 bits (606), Expect = 2e-62
Identities = 116/229 (50%), Positives = 157/229 (67%), Gaps = 19/229 (8%)
Query: 700 SLQEALGSKLRLSSAYHPQTDGQSERTIQSLEDLLRICVLEQGGTWDSHLPLIEFTYNNS 759
+ Q+ALG+++ LS+AYHPQTDGQSERTIQ+LED+LR CVL+ GG W+ +L L
Sbjct: 690 AFQKALGTRVNLSTAYHPQTDGQSERTIQTLEDMLRACVLDWGGNWEKYLTL-------- 741
Query: 760 YHSSIGMAPFEALYGRRCRTPLCWFESGERVVLGPEIVQQTTEKVQMIQEKMKASQSRQK 819
ALYGR CRTPLCW GER + GP IV +TTE+++ ++ K+K + RQK
Sbjct: 742 -----------ALYGRACRTPLCWTPVGERRLFGPTIVDETTERMKFLKIKLKEAHDRQK 790
Query: 820 SYHDKRRKDLEFQEGDHVFLRVTPMTGVXRALKSKKLTPKFIGPYQILERVGTVAYRVGL 879
SY +KRRK+LEFQ GD V+L+ G R KKL+P+++GPY+++ERVG VAY++ L
Sbjct: 791 SYANKRRKELEFQVGDLVYLKAMTYKGAGRFTSRKKLSPRYVGPYKVIERVGAVAYKLDL 850
Query: 880 PPHLSNLHNVFHVSQLRKYVPDPSHVIQSDDVQVRDNLTVETLPVRIDD 928
PP L+ HNVFHVSQLRK + D ++ +++N+TVE PVRI D
Sbjct: 851 PPKLNAFHNVFHVSQLRKCLSDQEESVEDVPPGLKENMTVEAWPVRIMD 899
Score = 29.6 bits (65), Expect = 8.6
Identities = 13/27 (48%), Positives = 17/27 (62%)
Query: 601 VESEHQNLLG*WVLLDVPXWKWDSISM 627
V++EHQ G L +P WKWD I+M
Sbjct: 661 VKAEHQVPSGLLQNLPIPEWKWDHITM 687
>At1g36590 hypothetical protein
Length = 1499
Score = 204 bits (518), Expect = 3e-52
Identities = 121/360 (33%), Positives = 194/360 (53%), Gaps = 9/360 (2%)
Query: 615 LDVPXWKWDSISMIL*RVAKYX*RDXRIGGILXR*RKSAHFLXXNISFPVSQXXEIYIKE 674
L +P W +SM I ++ R K+AHF+ + + + Y+
Sbjct: 1146 LPIPDTIWSEVSMDFIEGLPVSGGKTVIMVVVDRLSKAAHFIALSHPYSALTVAQAYLDN 1205
Query: 675 IVKCMGVPSSIVSDRDPRFTSRFWKSLQEALGSKLRLSSAYHPQTDGQSERTIQSLEDLL 734
+ K G P+SIVSDRD FTS FW+ G L+L+SAYHPQ+DGQ+E + LE L
Sbjct: 1206 VFKLHGCPTSIVSDRDVVFTSEFWREFFTLQGVALKLTSAYHPQSDGQTEVVNRCLETYL 1265
Query: 735 RICVLEQGGTWDSHLPLIEFTYNNSYHSSIGMAPFEALYGRRCRTPLCWFESGERVVLGP 794
R ++ W L L E+ YN +YHSS M PFE +YG+ L + +V +
Sbjct: 1266 RCMCHDRPQLWSKWLALAEYWYNTNYHSSSRMTPFEIVYGQVPPVHLPYLPGESKVAVVA 1325
Query: 795 EIVQQTTEKVQMIQEKMKASQSRQKSYHDKRRKDLEFQEGDHVFLRVTP---MTGVXRAL 851
+Q+ + + ++ + +Q R K + D+ R + EF+ GD+V++++ P + V RA
Sbjct: 1326 RSLQEREDMLLFLKFHLMRAQHRMKQFADQHRTEREFEIGDYVYVKLQPYRQQSVVMRA- 1384
Query: 852 KSKKLTPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPDPSHVIQSDDV 911
++KL+PK+ GPY+I++R G VAY++ LP + S +H VFHVSQL+ V + S + V
Sbjct: 1385 -NQKLSPKYFGPYKIIDRCGEVAYKLALPSY-SQVHPVFHVSQLKVLVGNVSTTVHLPSV 1442
Query: 912 QVRDNLTVETLPVRIDDRKVKMLRGKEIPLVRVVWTGATSESLTWELESKMLESYPELFA 971
++D E +P ++ +RK+ +GK + V V W+ E TWE + +++PE A
Sbjct: 1443 -MQD--VFEKVPEKVVERKMVNRQGKAVTKVLVKWSNEPLEEATWEFLFDLQKTFPEFEA 1499
>At3g11970 hypothetical protein
Length = 1499
Score = 203 bits (516), Expect = 4e-52
Identities = 121/360 (33%), Positives = 193/360 (53%), Gaps = 9/360 (2%)
Query: 615 LDVPXWKWDSISMIL*RVAKYX*RDXRIGGILXR*RKSAHFLXXNISFPVSQXXEIYIKE 674
L +P W +SM I ++ R K+AHF+ + + Y+
Sbjct: 1146 LPIPDTIWSEVSMDFIEGLPVSGGKTVIMVVVDRLSKAAHFIALSHPYSALTVAHAYLDN 1205
Query: 675 IVKCMGVPSSIVSDRDPRFTSRFWKSLQEALGSKLRLSSAYHPQTDGQSERTIQSLEDLL 734
+ K G P+SIVSDRD FTS FW+ G L+L+SAYHPQ+DGQ+E + LE L
Sbjct: 1206 VFKLHGCPTSIVSDRDVVFTSEFWREFFTLQGVALKLTSAYHPQSDGQTEVVNRCLETYL 1265
Query: 735 RICVLEQGGTWDSHLPLIEFTYNNSYHSSIGMAPFEALYGRRCRTPLCWFESGERVVLGP 794
R ++ W L L E+ YN +YHSS M PFE +YG+ L + +V +
Sbjct: 1266 RCMCHDRPQLWSKWLALAEYWYNTNYHSSSRMTPFEIVYGQVPPVHLPYLPGESKVAVVA 1325
Query: 795 EIVQQTTEKVQMIQEKMKASQSRQKSYHDKRRKDLEFQEGDHVFLRVTP---MTGVXRAL 851
+Q+ + + ++ + +Q R K + D+ R + EF+ GD+V++++ P + V RA
Sbjct: 1326 RSLQEREDMLLFLKFHLMRAQHRMKQFADQHRTEREFEIGDYVYVKLQPYRQQSVVMRA- 1384
Query: 852 KSKKLTPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPDPSHVIQSDDV 911
++KL+PK+ GPY+I++R G VAY++ LP + S +H VFHVSQL+ V + S + V
Sbjct: 1385 -NQKLSPKYFGPYKIIDRCGEVAYKLALPSY-SQVHPVFHVSQLKVLVGNVSTTVHLPSV 1442
Query: 912 QVRDNLTVETLPVRIDDRKVKMLRGKEIPLVRVVWTGATSESLTWELESKMLESYPELFA 971
++D E +P ++ +RK+ +GK + V V W+ E TWE + +++PE A
Sbjct: 1443 -MQD--VFEKVPEKVVERKMVNRQGKAVTKVLVKWSNEPLEEATWEFLFDLQKTFPEFEA 1499
>At2g14640 putative retroelement pol polyprotein
Length = 945
Score = 172 bits (435), Expect = 1e-42
Identities = 107/302 (35%), Positives = 165/302 (54%), Gaps = 9/302 (2%)
Query: 671 YIKEIVKCMGVPSSIVSDRDPRFTSRFWKSLQEALGSKLRLSSAYHPQTDGQSERTIQSL 730
++ +VK G+P SIVSD DP F S FW+ + +KL +S+AYHPQTDGQ+E + +
Sbjct: 638 FVSHVVKLHGIPRSIVSDCDPIFMSLFWQEFWKLSRTKLWMSTAYHPQTDGQTEVVNRCI 697
Query: 731 EDLLRICVLEQGGTWDSHLPLIEFTYNNSYHSSIGMAPFEALYGRRCRTPLCWFESGERV 790
E LR V W S +P E+ YN ++H+S GM PF+ALYGR +P+ +E G V
Sbjct: 698 EQFLRCFVHYHPKQWSSFIPWAEYWYNTTFHASTGMTPFQALYGRP-PSPIPAYELGS-V 755
Query: 791 VLGPEIVQQTTEKVQMIQE---KMKASQSRQKSYHDKRRKDLEFQEGDHVFLRVTPMTGV 847
V G E+ +Q + +++ E + + + K D R +D+ FQ GD V LR+ P
Sbjct: 756 VCG-ELNEQMAARDELLAELKQHLVTANNCMKQQADSRLRDVSFQVGDWVLLRIQPYRQK 814
Query: 848 XRALK-SKKLTPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPDPSHVI 906
+ S+KL+ +F GP+Q+ + G VAYR+ LP + +H VFHVS L+ +V D
Sbjct: 815 TLFRRSSQKLSHRFYGPFQVASKHGEVAYRLTLPEG-TRIHPVFHVSLLKPWVGD-GEPD 872
Query: 907 QSDDVQVRDNLTVETLPVRIDDRKVKMLRGKEIPLVRVVWTGATSESLTWELESKMLESY 966
+R+N ++ P + + + + K + + V W G E TWE ++ S+
Sbjct: 873 MGQLPPLRNNGELKLQPTAVLEVRWRSQDKKRVADLLVQWEGLHIEDATWEEYDQLAASF 932
Query: 967 PE 968
PE
Sbjct: 933 PE 934
>At3g31480 hypothetical protein
Length = 338
Score = 167 bits (422), Expect = 3e-41
Identities = 88/225 (39%), Positives = 131/225 (58%), Gaps = 42/225 (18%)
Query: 652 SAHFLXXNISFPVSQXXEIYIKEIVKCMGVPSSIVSDRDPRFTSRFWKSLQEALGSKLRL 711
SA FL + V+ Y+ EIVK GV SIVSD+D +FTS FW + Q +G+K+++
Sbjct: 26 SADFLAIRKTDGVAVLANKYVSEIVKLHGVHVSIVSDKDSKFTSAFWIAFQAEMGTKVQI 85
Query: 712 SSAYHPQTDGQSERTIQSLEDLLRICVLEQGGTWDSHLPLIEFTYNNSYHSSIGMAPFEA 771
S+AYHP+TDG+ E+TIQ+LED+LR+
Sbjct: 86 STAYHPKTDGKFEKTIQTLEDMLRM----------------------------------- 110
Query: 772 LYGRRCRTPLCWFESGERVVLGPEIVQQTTEKVQMIQEKMKASQSRQKSYHDKRRKDLEF 831
TPLCW + GER + G V +TTE++Q+++ MK + RQ+SY D RR+++EF
Sbjct: 111 -------TPLCWTQVGERSIYGANYVHETTERIQVLKLNMKEAHDRQRSYADMRRREVEF 163
Query: 832 QEGDHVFLRVTPMTGVXRALKSKKLTPKFIGPYQILERVGTVAYR 876
+ GD V+L++ + R + KL+PK++GP++I+ERVG VAYR
Sbjct: 164 EVGDRVYLKMDMLQSPKRFILETKLSPKYMGPFRIVERVGPVAYR 208
>At1g35370 hypothetical protein
Length = 1447
Score = 164 bits (415), Expect = 2e-40
Identities = 100/314 (31%), Positives = 157/314 (49%), Gaps = 28/314 (8%)
Query: 645 ILXR*RKSAHFLXXNISFPVSQXXEIYIKEIVKCMGVPSSIVSDRDPRFTSRFWKSLQEA 704
++ R K+AHF+ + + ++ + K G P+SIVSDRD FTS FWK +
Sbjct: 1147 VVDRLSKAAHFVALAHPYSALTVAQAFLDNVYKHHGCPTSIVSDRDVLFTSDFWKEFFKL 1206
Query: 705 LGSKLRLSSAYHPQTDGQSERTIQSLEDLLRICVLEQGGTWDSHLPLIEFTYNNSYHSSI 764
G +LR+SSAYHPQ+DGQ+E + LE+ LR + W+ LPL E+ YN +YHSS
Sbjct: 1207 QGVELRMSSAYHPQSDGQTEVVNRCLENYLRCMCHARPHLWNKWLPLAEYWYNTNYHSSS 1266
Query: 765 GMAPFEALYGRRCRTPLCWFESGERVVLGPEIVQQTTEKVQMIQEKMKASQSRQKSYHDK 824
M PFE +YG+ L + +V + +Q+ + ++ + +Q R K + D+
Sbjct: 1267 QMTPFELVYGQAPPIHLPYLPGKSKVAVVARSLQERENMLLFLKFHLMRAQHRMKQFADQ 1326
Query: 825 RRKDLEFQEGDHVFLRVTPMTGVXRALK-SKKLTPKFIGPYQILERVGTVAYRVGLPPHL 883
R + F GD V++++ P L+ ++KL+PK+ GPY+I+E+ G V
Sbjct: 1327 HRTERTFDIGDFVYVKLQPYRQQSVVLRVNQKLSPKYFGPYKIIEKCGEVM--------- 1377
Query: 884 SNLHNVFHVSQLRKYVPDPSHVIQSDDVQVRDNLTVETLPVRIDDRKVKMLRGKEIPLVR 943
+ NV +QL +PD E P I +RK+ +G+ +V
Sbjct: 1378 --VGNVTTSTQLPSVLPD----------------IFEKAPEYILERKLVKRQGRAATMVL 1419
Query: 944 VVWTGATSESLTWE 957
V W G E TW+
Sbjct: 1420 VKWIGEPVEEATWK 1433
>At4g10580 putative reverse-transcriptase -like protein
Length = 1240
Score = 140 bits (353), Expect = 3e-33
Identities = 73/182 (40%), Positives = 115/182 (63%), Gaps = 5/182 (2%)
Query: 791 VLGPEIVQQTTEKVQMIQEKMKASQSRQKSYHDKRRKDLEFQEGDHVFLRVTPMTGVXRA 850
VL + V + K+ + MK +Q RQ+SY DKRR++LEF+ GD V+L++ + G R+
Sbjct: 1056 VLAKKFVSEIV-KLHGVPLNMKEAQDRQRSYADKRRRELEFEVGDRVYLKMAMLRGPNRS 1114
Query: 851 LKSKKLTPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPDPSHVIQSDD 910
+ KL+P+++GP++I+ERV VAYR+ LP + H VFHVS LRK + +
Sbjct: 1115 ISETKLSPRYMGPFKIVERVEPVAYRLELPDVMRAFHKVFHVSMLRKCLHKDDEALAKIP 1174
Query: 911 VQVRDNLTVETLPVRIDDRKVKMLRGKEIPLVRVVW--TGATSESLTWELESKMLESYPE 968
++ N+T+E PVR+ +R++K LR K+IPL++V+W G T E TWE E++M + +
Sbjct: 1175 EDLQPNMTLEARPVRVLERRIKELRQKKIPLIKVLWDCDGVTEE--TWEPEARMKARFKK 1232
Query: 969 LF 970
F
Sbjct: 1233 WF 1234
Score = 53.1 bits (126), Expect = 7e-07
Identities = 37/116 (31%), Positives = 57/116 (48%), Gaps = 2/116 (1%)
Query: 601 VESEHQNLLG*WVLLDVPXWKWDSISMIL*RVAKYX*RDXRIGGILXR*RKSAHFLXXNI 660
V++EHQ L G L +P WKWD I+M L + I I+ R KSAHFL
Sbjct: 991 VKAEHQVLGGMLQSLPIPEWKWDFITMDLVVGLRVSRTKDAIWVIVDRLTKSAHFLAIRK 1050
Query: 661 SFPVSQXXEIYIKEIVKCMGVPSSI--VSDRDPRFTSRFWKSLQEALGSKLRLSSA 714
+ + + ++ EIVK GVP ++ DR + + + L+ +G ++ L A
Sbjct: 1051 TDGAAVLAKKFVSEIVKLHGVPLNMKEAQDRQRSYADKRRRELEFEVGDRVYLKMA 1106
>At2g06190 putative Ty3-gypsy-like retroelement pol polyprotein
Length = 280
Score = 124 bits (310), Expect = 3e-28
Identities = 80/257 (31%), Positives = 126/257 (48%), Gaps = 20/257 (7%)
Query: 645 ILXR*RKSAHFLXXNISFPVSQXXEIYIKEIVKCMGVPSSIVSDRDPRFTSRFWKSLQEA 704
++ R K HF+ + S +++ KE+V+ GVP SI SDRD +F S FW +L
Sbjct: 20 VVDRFSKMTHFIACKKTADASNIAKLFFKEVVRLHGVPKSITSDRDTKFLSHFWSTLWRM 79
Query: 705 LGSKLRLSSAYHPQTDGQSERTIQSLEDLLRICVLEQGGTWDSHLPLIEFTYNNSYHSSI 764
G+ L SS H Q DGQ+E T ++L +++R + WD LP IEF YN+ H +
Sbjct: 80 FGTALNRSSTPHTQIDGQTEVTNRTLGNMVRSICGDNPKQWDLALPQIEFAYNSVVH-VV 138
Query: 765 GMAPFEALYGRRCRTPLCWFESGERVVLGPEIVQQTTEKVQMIQEKMKASQSRQKSYHDK 824
+ G T E +++ E+V + K++A+ + K DK
Sbjct: 139 DLVKLPKALGASAET------MAEEILVVKEVV----------KAKLEATGKKNKVAADK 182
Query: 825 RRKDLEFQEGDHVFLRVTPMTGVXRALKSKKLTPKFIGPYQILERVGTVAYRVGLPPHLS 884
RR+ F+EGD V V G K+ P+ GP+++L ++ AY V LP +
Sbjct: 183 RRRFKVFKEGDDVM--VLLRKGRFAVGTYNKVKPRKYGPFKVLRKINDNAYVVALPKSM- 239
Query: 885 NLHNVFHVSQLRKYVPD 901
N+ N F+V+ + +Y D
Sbjct: 240 NISNTFNVADIHEYHAD 256
>At4g07850 putative polyprotein
Length = 1138
Score = 116 bits (290), Expect = 7e-26
Identities = 63/168 (37%), Positives = 96/168 (56%), Gaps = 1/168 (0%)
Query: 615 LDVPXWKWDSISM-IL*RVAKYX*RDXRIGGILXR*RKSAHFLXXNISFPVSQXXEIYIK 673
L +P W+ ISM + + + I ++ R K AHF+ + + ++ +
Sbjct: 823 LPIPLHPWNDISMDFVVGLPRTRTGKDSIFVVVDRFSKMAHFIPCHKTDDAMHIANLFFR 882
Query: 674 EIVKCMGVPSSIVSDRDPRFTSRFWKSLQEALGSKLRLSSAYHPQTDGQSERTIQSLEDL 733
E+V+ G+P +IVSDRD +F S FWK+L LG+KL S+ HPQTDGQ+E ++L L
Sbjct: 883 EVVRLHGMPKTIVSDRDTKFLSYFWKTLWSKLGTKLLFSTTCHPQTDGQTEVVNRTLSTL 942
Query: 734 LRICVLEQGGTWDSHLPLIEFTYNNSYHSSIGMAPFEALYGRRCRTPL 781
LR + + TW+ LP +EF YN+S HS+ +PF+ +YG TPL
Sbjct: 943 LRALIKKNLKTWEDCLPHVEFAYNHSVHSATKFSPFQIVYGFNPITPL 990
>At2g05610 putative retroelement pol polyprotein
Length = 780
Score = 89.0 bits (219), Expect = 1e-17
Identities = 50/156 (32%), Positives = 90/156 (57%), Gaps = 5/156 (3%)
Query: 766 MAPFEALYGRRCRTPLCWFESGERVVLGPEIVQQTTEKVQMIQEKMKASQSRQKSYHDKR 825
M P+EA+YG+ L + +V + +Q+ + ++ + +Q R K D+
Sbjct: 625 MTPYEAVYGQPPPLHLPYLPGESKVAVVARSMQERESMILFLKFHLMRAQHRMKQLADQH 684
Query: 826 RKDLEFQEGDHVFLRVTPMTGVXRALKS-KKLTPKFIGPYQILERVGTVAYRVGLPPHLS 884
+ EF+ GD+VF+++ P ++S +KL+PK+ GPY++++R G VAY++ LP + S
Sbjct: 685 ITEREFEVGDYVFVKLQPYRQQSVVMRSTQKLSPKYFGPYKVIDRCGEVAYKLQLPAN-S 743
Query: 885 NLHNVFHVSQLRKY---VPDPSHVIQSDDVQVRDNL 917
+H VFHVSQLR V +H+++ + +R+NL
Sbjct: 744 QVHPVFHVSQLRVLVGTVTTSTHLLRCYLMSLRENL 779
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.350 0.155 0.524
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,340,909
Number of Sequences: 26719
Number of extensions: 628479
Number of successful extensions: 2465
Number of sequences better than 10.0: 53
Number of HSP's better than 10.0 without gapping: 41
Number of HSP's successfully gapped in prelim test: 12
Number of HSP's that attempted gapping in prelim test: 2372
Number of HSP's gapped (non-prelim): 65
length of query: 971
length of database: 11,318,596
effective HSP length: 109
effective length of query: 862
effective length of database: 8,406,225
effective search space: 7246165950
effective search space used: 7246165950
T: 11
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.9 bits)
S2: 65 (29.6 bits)
Medicago: description of AC147714.2


