

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147714.16 - phase: 0
(322 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
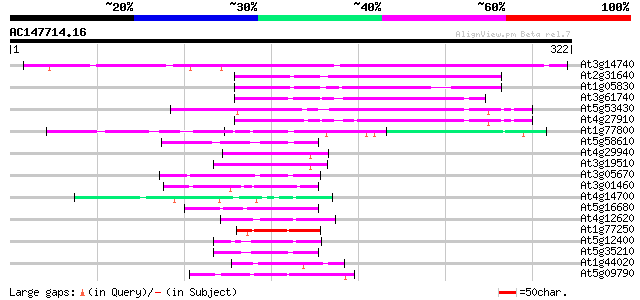
Sequences producing significant alignments: (bits) Value
At3g14740 unknown protein 305 2e-83
At2g31640 putative PHD-type zinc finger protein 120 1e-27
At1g05830 tri-thorax like protein (At1g05830) 94 8e-20
At3g61740 trithorax 3 (ATX3) 87 2e-17
At5g53430 trithorax 5 (TX5) 84 8e-17
At4g27910 trithorax 4 (TX4) 79 3e-15
At1g77800 putative phorbol ester / diacylglycerol binding protein 74 1e-13
At5g58610 putative protein 53 2e-07
At4g29940 pathogenesis related homeodomain protein (PRHA) 50 2e-06
At3g19510 putative homeobox protein, HAT3.1 48 6e-06
At3g05670 unknown protein 48 8e-06
At3g01460 unknown protein 47 1e-05
At4g14700 replication control protein 1 like 46 2e-05
At5g16680 putative protein 46 3e-05
At4g12620 origin recognition complex subunit 1 -like protein 45 5e-05
At1g77250 unknown protein 45 5e-05
At5g12400 putative protein 45 7e-05
At5g35210 putative protein 44 9e-05
At1g44020 hypothetical protein 44 9e-05
At5g09790 unknown protein 44 1e-04
>At3g14740 unknown protein
Length = 341
Score = 305 bits (781), Expect = 2e-83
Identities = 160/343 (46%), Positives = 207/343 (59%), Gaps = 42/343 (12%)
Query: 9 DLPPNKRLRFIYQ-----QQQQQEEQDLSHCSLLPTKKRKESRNSSLFHTPPSSPTPPPS 63
DLPP KRLR + + QQ Q Q LP KKRK++R + + + P+
Sbjct: 7 DLPPLKRLRLLQRDLEAAQQHQLPNQPEMKSLQLPAKKRKQTR---VDYDDDDAENSNPT 63
Query: 64 TYSLPTKKRITALQPHLHHHNNIPNDAVPLIDLNVEYSP-------------SLPSATPI 110
+ LP KKRI A+ P L P DLNVEY P ++ S+ +
Sbjct: 64 YHCLPAKKRIWAIDPDLLSGGGNPFSP---FDLNVEYKPYVEEKSIEKKSTLNVESSLEV 120
Query: 111 EKQSQKQDIE------------FEVDDDE-ILCCVCHSTDANAEDPIVFCDGCNLMVHAS 157
E+ K++I+ EV+D++ I+C VC STD + +PIVFCDGC+LMVHAS
Sbjct: 121 EEDDDKENIDPLGKGKALDLSDREVEDEDGIMCAVCQSTDGDPLNPIVFCDGCDLMVHAS 180
Query: 158 CYGNPLVKQIPDGDWFCDQCRFKNDIDTDTGPIRCSLCPTKEGAMKQTTDGKWVHLVCAL 217
CYGNPLVK IP+GDWFC QC + + C LC TK GAMK T DG+W H+ CAL
Sbjct: 181 CYGNPLVKAIPEGDWFCRQCLSSKNREKI---FSCCLCTTKGGAMKPTNDGRWAHITCAL 237
Query: 218 LVPEVFFVDPEGREGIDCSKIPKKRWLEKCYVCGCFDGCALVCSEQKCGLGFHITCGIKE 277
VPEV+F DPEGREGI CS++ KRW ++CY+C GC + CSE +C L FH+TCG+KE
Sbjct: 238 FVPEVYFEDPEGREGICCSEVLSKRWKDRCYLCKVRRGCVIECSEMRCKLAFHVTCGLKE 297
Query: 278 DLCIEYKEGKKGATVVAGFCKTHSQIWEKNKGSGKYKIVAVED 320
DLCIEY+EGKK +V GFC H+++WE+ SGKYKIVA E+
Sbjct: 298 DLCIEYREGKKSGGIVVGFCNEHTKLWERE--SGKYKIVAREE 338
>At2g31640 putative PHD-type zinc finger protein
Length = 178
Score = 120 bits (300), Expect = 1e-27
Identities = 59/154 (38%), Positives = 77/154 (49%), Gaps = 7/154 (4%)
Query: 130 CCVCHSTDANAEDPIVFCDGCNLMVHASCYGNPLVKQIPDGDWFCDQCRFKNDIDTDTGP 189
C VCH + + + CD C +MVHA CYG ++ W C+ CR P
Sbjct: 29 CNVCHMDEEYENNLFLQCDKCRMMVHAKCYGE--LEPCDGALWLCNLCR----PGAPDMP 82
Query: 190 IRCSLCPTKEGAMKQTTDGKWVHLVCALLVPEVFFVDPEGREGID-CSKIPKKRWLEKCY 248
RC LCP GAMK TTDG+W HL CA+ +PE D + E ID +K+ K RW C
Sbjct: 83 PRCCLCPVVGGAMKPTTDGRWAHLACAIWIPETCLSDVKKMEPIDGVNKVSKDRWKLMCT 142
Query: 249 VCGCFDGCALVCSEQKCGLGFHITCGIKEDLCIE 282
+CG G + CS C + +H C LC+E
Sbjct: 143 ICGVSYGACIQCSNNSCRVAYHPLCARAAGLCVE 176
>At1g05830 tri-thorax like protein (At1g05830)
Length = 1133
Score = 94.4 bits (233), Expect = 8e-20
Identities = 53/153 (34%), Positives = 70/153 (45%), Gaps = 17/153 (11%)
Query: 130 CCVCHSTDANAEDPIVFCDGCNLMVHASCYGNPLVKQIPDGDWFCDQCRFKNDIDTDTGP 189
C VCH + + + CD C +MVH CYG ++ W C+ CR + D P
Sbjct: 592 CNVCHMDEEYENNLFLQCDKCRMMVHTRCYGQ--LEPHNGILWLCNLCR---PVALDIPP 646
Query: 190 IRCSLCPTKEGAMKQTTDGKWVHLVCALLVPEVFFVDPEGREGIDCSKIPKKRWLEKCYV 249
RC LCP GAMK TTDG+W HL CA+ +PE +D + E ID K K
Sbjct: 647 -RCCLCPVVGGAMKPTTDGRWAHLACAIWIPETCLLDVKKMEPIDGVKKVSKN------- 698
Query: 250 CGCFDGCALVCSEQKCGLGFHITCGIKEDLCIE 282
+ CS C + +H C LC+E
Sbjct: 699 ----NDADFQCSNNTCRVAYHPLCARAAGLCVE 727
>At3g61740 trithorax 3 (ATX3)
Length = 900
Score = 86.7 bits (213), Expect = 2e-17
Identities = 51/146 (34%), Positives = 70/146 (47%), Gaps = 12/146 (8%)
Query: 130 CCVCHSTDANAEDPIVFCDGCNLMVHASCYGNPLVKQIPDGDWFCDQCRFKNDIDTDTGP 189
C VC + E+ ++ C+ C + VH CYG + K W C C DI+ D
Sbjct: 493 CAVCRWVEDWEENKMIICNRCQVAVHQECYG--VSKSQDLTSWVCRACE-TPDIERD--- 546
Query: 190 IRCSLCPTKEGAMKQT-TDGKWVHLVCALLVPEVFFVDPEGRE-GIDCSKIPKKRWLEKC 247
C LCP K GA+K + +G WVH+ CA PEV F++ E E + KIP +L+ C
Sbjct: 547 --CCLCPVKGGALKPSDVEGLWVHVTCAWFRPEVGFLNHENMEPAVGLFKIPANSFLKVC 604
Query: 248 YVCGCFDGCALVCSEQKCGLGFHITC 273
+C G + C KC FH C
Sbjct: 605 TICKQTHGSCVHCC--KCATHFHAMC 628
Score = 35.8 bits (81), Expect = 0.032
Identities = 13/41 (31%), Positives = 25/41 (60%), Gaps = 2/41 (4%)
Query: 145 VFCDGCNLMVHASC--YGNPLVKQIPDGDWFCDQCRFKNDI 183
V CDGC++ VHA C N K++ +++C C+ ++++
Sbjct: 367 VCCDGCDVWVHAECDNITNERFKELEHNNYYCPDCKVQHEL 407
>At5g53430 trithorax 5 (TX5)
Length = 1043
Score = 84.3 bits (207), Expect = 8e-17
Identities = 62/223 (27%), Positives = 95/223 (41%), Gaps = 27/223 (12%)
Query: 93 LIDLNVEYSPSLPSATPIEKQSQKQDIEFEVDDDEIL--------CCVCHSTDANAEDPI 144
L + + + + P P KQ +++ + F + E + C VC + + I
Sbjct: 565 LAEFHANATAAKPPKRPSIKQRKQRLLSFLREKYEPVNVKWTTERCAVCRWVEDWDYNKI 624
Query: 145 VFCDGCNLMVHASCYGNPLVKQIPDGDWFCDQCRFKNDIDTDTGPIRCSLCPTKEGAMKQ 204
+ C+ C + VH CYG V+ W C C +T C LCP K GA+K
Sbjct: 625 IICNRCQIAVHQECYGTRNVRDFT--SWVCKAC------ETPEIKRECCLCPVKGGALKP 676
Query: 205 T-TDGKWVHLVCALLVPEVFFVDPEGRE-GIDCSKIPKKRWLEKCYVCGCFDGCALVCSE 262
T + WVH+ CA PEV F E E + IP +++ C +C G C
Sbjct: 677 TDVETLWVHVTCAWFQPEVCFASEEKMEPALGILSIPSSNFVKICVICKQIHGSCTQCC- 735
Query: 263 QKCGLGFHITC----GIKEDL-CIEYKEGKKGATVVAGFCKTH 300
KC +H C G + +L C+E K G++ T + +C H
Sbjct: 736 -KCSTYYHAMCASRAGYRMELHCLE-KNGRQ-ITKMVSYCSYH 775
Score = 35.4 bits (80), Expect = 0.042
Identities = 14/42 (33%), Positives = 24/42 (56%), Gaps = 2/42 (4%)
Query: 145 VFCDGCNLMVHASC--YGNPLVKQIPDGDWFCDQCRFKNDID 184
V CDGC + +H++C + K + + D++C CR K D +
Sbjct: 432 VRCDGCKVWIHSACDQISHKHFKDLGETDYYCPTCRTKFDFE 473
>At4g27910 trithorax 4 (TX4)
Length = 954
Score = 79.3 bits (194), Expect = 3e-15
Identities = 56/178 (31%), Positives = 80/178 (44%), Gaps = 20/178 (11%)
Query: 130 CCVCHSTDANAEDPIVFCDGCNLMVHASCYGNPLVKQIPDGDWFCDQCRFKNDIDTDTGP 189
C VC + + I+ C+ C + VH CYG V+ W C C + DI +
Sbjct: 523 CAVCRWVEDWDYNKIIICNRCQIAVHQECYGARHVRDFTS--WVCKACE-RPDIKRE--- 576
Query: 190 IRCSLCPTKEGAMKQT-TDGKWVHLVCALLVPEVFFVDPEGRE-GIDCSKIPKKRWLEKC 247
C LCP K GA+K T + WVH+ CA PEV F E E + IP +++ C
Sbjct: 577 --CCLCPVK-GALKPTDVETLWVHVTCAWFQPEVCFASEEKMEPAVGILSIPSTNFVKIC 633
Query: 248 YVCGCFDGCALVCSEQKCGLGFHITC----GIKEDL-CIEYKEGKKGATVVAGFCKTH 300
+C G C KC +H C G + +L C+E K G++ T + +C H
Sbjct: 634 VICKQIHGSCTQCC--KCSTYYHAMCASRAGYRMELHCLE-KNGQQ-ITKMVSYCAYH 687
>At1g77800 putative phorbol ester / diacylglycerol binding protein
Length = 1506
Score = 73.9 bits (180), Expect = 1e-13
Identities = 56/195 (28%), Positives = 79/195 (39%), Gaps = 20/195 (10%)
Query: 124 DDDEILCCVCHSTDANAEDPIVFCDGCNLMVHASCYGNPLVKQIPDGDWFCDQCRFKNDI 183
+ + +LC C ++ C C VH CYG + + W C C +N
Sbjct: 279 EGNALLCDFC----CTGHHQLIVCTSCKATVHKKCYG---LLEDSGKPWLCSWCELENGR 331
Query: 184 DTDTGPIRCSLCPTKEGAMK----QTTDG---KWVHLVCALLVPEVFFVDPEGREGI-DC 235
P C LCP K G +K +T +G ++ HL C+L +PEV+ D + E I +
Sbjct: 332 ADSERP--CLLCPKKGGILKPVLSKTENGGPAEFAHLFCSLWMPEVYIEDLKKMEPILNF 389
Query: 236 SKIPKKRWLEKCYVCGCFDGCALVCSEQKCGLGFHITCGIKEDLCIEYKEGKKGATVV-- 293
I + R C +C G + C C FH C + +E GK G V
Sbjct: 390 PGIKETRRKLLCNLCKVKSGACIRCCNGTCRTSFHPICAREAGNRLEV-WGKHGCDTVEL 448
Query: 294 AGFCKTHSQIWEKNK 308
FC HS I E K
Sbjct: 449 RAFCSKHSDIQESGK 463
Score = 63.5 bits (153), Expect = 1e-10
Identities = 50/201 (24%), Positives = 83/201 (40%), Gaps = 27/201 (13%)
Query: 22 QQQQQEEQDLSHCSLLPTKKRKESRNSSLFHTPPSSPTPPPSTYSLPTKKRITALQPHLH 81
+++Q+ +Q + + SRN+SL + P+ + T +R HL
Sbjct: 948 RKEQRNKQAQAVLAAATAAAATSSRNTSL----RKDMSEEPAQQEMSTSRRKVVGSSHL- 1002
Query: 82 HHNNIPNDAVPLIDLNVEYSPSLPSATPIEKQSQKQDIEFEVDDDEILCCVCHSTDANAE 141
+P L+ + V S P EK+S +F V++ C +C ++
Sbjct: 1003 ----VPQTKESLLKMAV-------SGPPSEKRSDHHTPDFLVENPRT-CDICRRSET-IW 1049
Query: 142 DPIVFCDGCNLMVHASCYGNPLVKQIPDGDWFCDQCRFK------NDIDTDTGPIRCSLC 195
+ IV C C + VH CY + G W+C+ C N + C+LC
Sbjct: 1050 NLIVVCSSCKVAVHIDCYK---CAKESTGPWYCELCAESSSEPSFNFGEKPNSSTECTLC 1106
Query: 196 PTKEGAMKQTTDGKWVHLVCA 216
GA ++TT+G+WVH CA
Sbjct: 1107 GGTTGAFRKTTNGQWVHAFCA 1127
>At5g58610 putative protein
Length = 1065
Score = 53.1 bits (126), Expect = 2e-07
Identities = 27/90 (30%), Positives = 41/90 (45%), Gaps = 12/90 (13%)
Query: 88 NDAVPLIDLNVEYSPSLPSATPIEKQSQKQDIEFEVDDDEILCCVCHSTDANAEDPIVFC 147
+D L++ VE A P + K ++++ C VCH ++ C
Sbjct: 659 DDGRSLLECQVEAYKKRKKAQPPDMLKMK----LRQGENDVFCSVCHYGGK-----LILC 709
Query: 148 DGCNLMVHASCYGNPLVKQIPDGDWFCDQC 177
DGC HA+C G ++ +PDGDWFC C
Sbjct: 710 DGCPSAFHANCLG---LEDVPDGDWFCQSC 736
>At4g29940 pathogenesis related homeodomain protein (PRHA)
Length = 796
Score = 50.1 bits (118), Expect = 2e-06
Identities = 26/65 (40%), Positives = 34/65 (52%), Gaps = 4/65 (6%)
Query: 123 VDDDEILCCVCHSTDANAEDPIVFCDG-CNLMVHASCYGNPL-VKQIPDGD--WFCDQCR 178
+ D I C C+S +A ++ I+ CDG CN H C PL + IP GD WFC C
Sbjct: 186 IHHDHIFCAECNSREAFPDNDIILCDGTCNRAFHQKCLDPPLETESIPPGDQGWFCKFCD 245
Query: 179 FKNDI 183
K +I
Sbjct: 246 CKIEI 250
>At3g19510 putative homeobox protein, HAT3.1
Length = 723
Score = 48.1 bits (113), Expect = 6e-06
Identities = 25/69 (36%), Positives = 36/69 (51%), Gaps = 4/69 (5%)
Query: 118 DIEFEVDDDEILCCVCHSTDANAEDPIVFCDG-CNLMVHASCYGNPLVKQ-IPDGD--WF 173
D + E+ ++I C C S D + ++ I+ CDG C+ H C PL K+ IP D W
Sbjct: 256 DTDGEISSEDIFCAKCGSKDLSVDNDIILCDGFCDRGFHQYCLEPPLRKEDIPPDDEGWL 315
Query: 174 CDQCRFKND 182
C C K+D
Sbjct: 316 CPGCDCKDD 324
>At3g05670 unknown protein
Length = 883
Score = 47.8 bits (112), Expect = 8e-06
Identities = 24/92 (26%), Positives = 47/92 (51%), Gaps = 5/92 (5%)
Query: 87 PNDAVPLIDLNVEYSPSLPSATPIEKQSQKQDIEFEVDDDEILCCVCHSTDANAEDPIVF 146
P + P +DL P +P + + ++++ + + I+C CH D + ++
Sbjct: 464 PARSTPGVDLREVVIP-VPERDQVYQPTEEELRSYLDPYENIICTECHQGDDDGL--MLL 520
Query: 147 CDGCNLMVHASCYGNPLVKQIPDGDWFCDQCR 178
CD C+ H C G L +++P+G+W+C+ CR
Sbjct: 521 CDLCDSSAHTYCVG--LGREVPEGNWYCEGCR 550
>At3g01460 unknown protein
Length = 2176
Score = 47.4 bits (111), Expect = 1e-05
Identities = 29/95 (30%), Positives = 46/95 (47%), Gaps = 11/95 (11%)
Query: 89 DAVPLIDLNVEYSPSLPSATPIEKQSQKQDIEFEVDD------DEILCCVCHSTDANAED 142
+ VPL+ +Y E + + +DI V+ DE +C VC D + +D
Sbjct: 1245 EVVPLVQKLKDYRKL--ECLSAEMKKEIKDIVVSVNKLPKAPWDEGVCKVC-GVDKD-DD 1300
Query: 143 PIVFCDGCNLMVHASCYGNPLVKQIPDGDWFCDQC 177
++ CD C+ H C PL++ IPDG+W+C C
Sbjct: 1301 SVLLCDTCDAEYHTYCLNPPLIR-IPDGNWYCPSC 1334
Score = 37.0 bits (84), Expect = 0.014
Identities = 25/92 (27%), Positives = 35/92 (37%), Gaps = 16/92 (17%)
Query: 87 PNDAVPLIDLNVEYSPSLPSATPIEKQSQKQDIEFEVDDDEILCCVCHSTDANAEDPIVF 146
P + V I N + +P P+ P D + C C ++ + +V
Sbjct: 56 PVEVVRSIHDNPDPAPGAPAEVP-------------EPDRDASCGACGRPESI--ELVVV 100
Query: 147 CDGCNLMVHASCYGNPLVKQIPDGDWFCDQCR 178
CD C H SC N V+ P DW C CR
Sbjct: 101 CDACERGFHMSCV-NDGVEAAPSADWMCSDCR 131
>At4g14700 replication control protein 1 like
Length = 771
Score = 46.2 bits (108), Expect = 2e-05
Identities = 49/195 (25%), Positives = 72/195 (36%), Gaps = 58/195 (29%)
Query: 38 PTKKRKESRNSSLFHTPPSSPTPPPSTYSLPTKKRITALQPHLHHHNNIPNDAVPL---- 93
PTK + S +S TPP + +L +R + H+ N+ ND + L
Sbjct: 15 PTKTPTKMYRKSYLSPSSTSLTPPQTPETLTPLRRSSR---HVSRKINLGNDPIDLPGKE 71
Query: 94 ----------------------------IDLNVEYSPSLPSATPIEKQSQKQDI------ 119
ID V +SP P + +K +K+ +
Sbjct: 72 SVEEINLIRKPRKRTNDIVVAEKSKKKKIDPEVSFSPVSPIRSETKKTKKKKRVYYNKVE 131
Query: 120 ----EFEVDDDEILCCVCHSTDANA-----EDPIVFCDGCNLMVHASCYGNPLVKQIPDG 170
EFE+ DD V + DAN EDP + + C H +C PL K++P+G
Sbjct: 132 FDETEFEIGDDVY---VKRTEDANPDEEEEEDPEI--EDCGF--HLNCLKPPL-KEVPEG 183
Query: 171 DWFCDQCRFKNDIDT 185
DW C C K T
Sbjct: 184 DWICQFCEVKKSGQT 198
>At5g16680 putative protein
Length = 1280
Score = 45.8 bits (107), Expect = 3e-05
Identities = 23/77 (29%), Positives = 39/77 (49%), Gaps = 3/77 (3%)
Query: 101 SPSLPSATPIEKQSQKQDIEFEVDDDEILCCVCHSTDANAEDPIVFCDGCNLMVHASCYG 160
S S S + +S+ D E V+ D +C +C DA ED + C GC+ +
Sbjct: 248 SKSSSSNSSAVSESESDDSEM-VEHDVKVCDICG--DAGREDLLAICSGCSDGAEHTYCM 304
Query: 161 NPLVKQIPDGDWFCDQC 177
++ ++P+GDW C++C
Sbjct: 305 REMLDEVPEGDWLCEEC 321
>At4g12620 origin recognition complex subunit 1 -like protein
Length = 813
Score = 45.1 bits (105), Expect = 5e-05
Identities = 24/67 (35%), Positives = 33/67 (48%), Gaps = 5/67 (7%)
Query: 122 EVDDDEILCC-VCHSTDANAEDPIVFCDGCNLMVHASCYGNPLVKQIPDGDWFCDQCRFK 180
E +D EI C +C +D N ++ CD C H C PL K++P+GDW C C K
Sbjct: 160 EEEDPEIEDCQICFKSDTNI---MIECDDCLGGFHLKCLKPPL-KEVPEGDWICQFCEVK 215
Query: 181 NDIDTDT 187
+ T
Sbjct: 216 KSGQSQT 222
>At1g77250 unknown protein
Length = 522
Score = 45.1 bits (105), Expect = 5e-05
Identities = 20/50 (40%), Positives = 30/50 (60%), Gaps = 4/50 (8%)
Query: 131 CVCHS--TDANAEDPIVFCDGCNLMVHASCYGNPLVKQIPDGDWFCDQCR 178
C+C + TD + +D IV CDGC+ H C P + +P+G+WFC C+
Sbjct: 403 CLCRNCLTDKD-DDKIVLCDGCDDAYHIYCM-RPPCESVPNGEWFCTACK 450
Score = 34.3 bits (77), Expect = 0.093
Identities = 27/83 (32%), Positives = 35/83 (41%), Gaps = 6/83 (7%)
Query: 129 LCCVCHSTDANAEDPIVFCDGCNLMVHASCYGNPLVKQIPDGDWFCDQCRFKNDIDTDTG 188
+C +C A A D + CD C M H SC P K +P W+C C K
Sbjct: 242 ICKLC-GEKAEARDCLA-CDHCEDMYHVSC-AQPGGKGMPTHSWYCLDCTSKGIGSPHEN 298
Query: 189 PIRCSLCPTKEGAMKQTTDGKWV 211
+ C T +G MK TD + V
Sbjct: 299 CVVCEKMKT-QGMMK--TDNRSV 318
>At5g12400 putative protein
Length = 1595
Score = 44.7 bits (104), Expect = 7e-05
Identities = 20/62 (32%), Positives = 31/62 (49%), Gaps = 8/62 (12%)
Query: 118 DIEFEVDDDEILCCVCHSTDANAEDPIVFCDGCNLMVHASCYGNPLVKQIPDGDWFCDQC 177
D F+ + D+ CC C + ++ CDGC H+ C G +P+GDW+C +C
Sbjct: 601 DTSFDRNSDD--CCFC-----KMDGSLLCCDGCPAAYHSKCVGLAS-HLLPEGDWYCPEC 652
Query: 178 RF 179
F
Sbjct: 653 AF 654
>At5g35210 putative protein
Length = 1606
Score = 44.3 bits (103), Expect = 9e-05
Identities = 21/61 (34%), Positives = 33/61 (53%), Gaps = 8/61 (13%)
Query: 118 DIEFEVDDDEILCCVCHSTDANAEDPIVFCDGCNLMVHASCYGNPLVKQ-IPDGDWFCDQ 176
++ ++D + C +C + ++ CDGC L H+ C G +VK IPDG WFC +
Sbjct: 402 EVSSDLDGNSDECRIC-----GMDGTLLCCDGCPLAYHSRCIG--VVKMYIPDGPWFCPE 454
Query: 177 C 177
C
Sbjct: 455 C 455
Score = 35.4 bits (80), Expect = 0.042
Identities = 23/76 (30%), Positives = 32/76 (41%), Gaps = 14/76 (18%)
Query: 105 PSATPIEKQSQKQDIEFEVDDDEILCCVCHSTDANAEDPIVFCDGCNLMVHASCYGNPLV 164
P P EK+ E +DD C C D P + C C L++H+ C +P
Sbjct: 1347 PQFEPTEKE--------ECEDDMGPCQRCLQMDPA---PDLLCTVCGLLIHSHC--SPW- 1392
Query: 165 KQIPDGDWFCDQCRFK 180
+P W C QCR +
Sbjct: 1393 SALPGSSWSCGQCRIR 1408
>At1g44020 hypothetical protein
Length = 654
Score = 44.3 bits (103), Expect = 9e-05
Identities = 27/78 (34%), Positives = 35/78 (44%), Gaps = 17/78 (21%)
Query: 128 ILCCVCHSTDANAEDPIVFCDGCNLMVHASCYGNPLVKQI-------------PDGDWFC 174
+ C VC D+++ PI C C+ +VH C G P V +I +GDW C
Sbjct: 484 LTCNVCAVADSSS--PIYMCPPCDFVVHQRCTGLPRVIRISRHRHRISFTTSFDEGDWSC 541
Query: 175 DQCRFKNDIDTDTGPIRC 192
CR K ID D G C
Sbjct: 542 GVCRRK--IDNDYGGFSC 557
>At5g09790 unknown protein
Length = 352
Score = 43.9 bits (102), Expect = 1e-04
Identities = 26/97 (26%), Positives = 45/97 (45%), Gaps = 8/97 (8%)
Query: 104 LPSATPIEKQSQKQDIEFEVDDDEILCCVCHSTDANAEDPIVFCDGCNLMVHASCYGNPL 163
+ + P+ +Q +++D E + C C S + +D ++ CD C+ H C P+
Sbjct: 44 MAKSVPVVEQEEEED---EDSYSNVTCEKCGSGEG--DDELLLCDKCDRGFHMKCL-RPI 97
Query: 164 VKQIPDGDWFCDQCRFKNDIDTDTGPIR--CSLCPTK 198
V ++P G W C C + + +T R CSL K
Sbjct: 98 VVRVPIGTWLCVDCSDQRPVRKETRKRRRSCSLTVKK 134
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.137 0.445
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,047,326
Number of Sequences: 26719
Number of extensions: 455962
Number of successful extensions: 2845
Number of sequences better than 10.0: 207
Number of HSP's better than 10.0 without gapping: 55
Number of HSP's successfully gapped in prelim test: 153
Number of HSP's that attempted gapping in prelim test: 2512
Number of HSP's gapped (non-prelim): 410
length of query: 322
length of database: 11,318,596
effective HSP length: 99
effective length of query: 223
effective length of database: 8,673,415
effective search space: 1934171545
effective search space used: 1934171545
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 60 (27.7 bits)
Medicago: description of AC147714.16


