

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147712.6 + phase: 0
(80 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
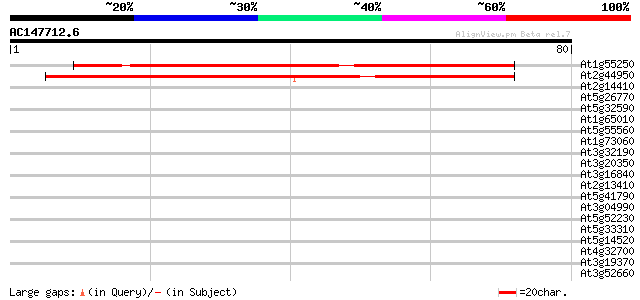
Score E
Sequences producing significant alignments: (bits) Value
At1g55250 unknown protein 62 4e-11
At2g44950 unknown protein 41 9e-05
At2g14410 hypothetical protein 32 0.043
At5g26770 unknown protein 32 0.057
At5g32590 putative protein 32 0.074
At1g65010 hypothetical protein 31 0.13
At5g55560 putative protein 30 0.16
At1g73060 unknown protein 30 0.21
At3g32190 hypothetical protein 30 0.28
At3g20350 unknown protein 30 0.28
At3g16840 ATP-dependent RNA helicase 30 0.28
At2g13410 T26C18.1 30 0.28
At5g41790 myosin heavy chain-like protein 29 0.48
At3g04990 hypothetical protein 29 0.48
At5g52230 unknown protein 28 0.63
At5g33310 putative protein 28 0.63
At5g14520 pescadillo - like protein 28 0.63
At4g32700 putative protein 28 0.63
At3g19370 unknown protein 28 0.63
At3g52660 putative RNA binding protein 28 0.82
>At1g55250 unknown protein
Length = 899
Score = 62.4 bits (150), Expect = 4e-11
Identities = 32/63 (50%), Positives = 46/63 (72%), Gaps = 3/63 (4%)
Query: 10 DLDSKRSSQKKLEEELMNVNNQIAELNSETGETVVQQLEEEEEIRVCKNMIKCTVFSDRP 69
+LD +R +KKLEEELM +N ++ EL SE+ E + +L +EE++ CKN++KC V DRP
Sbjct: 798 ELDDERE-KKKLEEELMELNKELEELGSESVEAAIVRL--QEEVKNCKNILKCGVCFDRP 854
Query: 70 KEV 72
KEV
Sbjct: 855 KEV 857
>At2g44950 unknown protein
Length = 518
Score = 41.2 bits (95), Expect = 9e-05
Identities = 22/68 (32%), Positives = 42/68 (61%), Gaps = 3/68 (4%)
Query: 6 ASEKDLDSKRSSQKKLEEELMNVNNQIAELNSET-GETVVQQLEEEEEIRVCKNMIKCTV 64
A E +L+ +R +++++EEE+ +++ L S G + +Q+L +E + K ++KC
Sbjct: 411 ALELELEIERFNRRRIEEEMEIAKKKVSRLRSLIEGSSAIQKLRQE--LSEFKEILKCKA 468
Query: 65 FSDRPKEV 72
+DRPKEV
Sbjct: 469 CNDRPKEV 476
>At2g14410 hypothetical protein
Length = 679
Score = 32.3 bits (72), Expect = 0.043
Identities = 15/57 (26%), Positives = 34/57 (59%)
Query: 4 ASASEKDLDSKRSSQKKLEEELMNVNNQIAELNSETGETVVQQLEEEEEIRVCKNMI 60
+S+ E DL S + ++ KLE++L N++ ++ + N E E + + +EE+ ++ +
Sbjct: 453 SSSLESDLHSSKDARLKLEDQLDNLSTELMKSNGELQEQYQRYDKIQEELSTSRDRL 509
>At5g26770 unknown protein
Length = 335
Score = 32.0 bits (71), Expect = 0.057
Identities = 16/46 (34%), Positives = 30/46 (64%), Gaps = 3/46 (6%)
Query: 8 EKDLDSKRSSQKKLEEELMNVNNQIAELNSETG---ETVVQQLEEE 50
++DL ++ SQK+L EE+M + +I E +++G E +++L EE
Sbjct: 157 QRDLKTRECSQKQLREEVMRIEREITEAVAKSGKGTECELRKLLEE 202
>At5g32590 putative protein
Length = 871
Score = 31.6 bits (70), Expect = 0.074
Identities = 14/56 (25%), Positives = 34/56 (60%)
Query: 5 SASEKDLDSKRSSQKKLEEELMNVNNQIAELNSETGETVVQQLEEEEEIRVCKNMI 60
S+ E DL S +++KLE++L N+++++ + N E + + + +EE+ ++ +
Sbjct: 599 SSLESDLRSSNDARQKLEDQLDNLSSELMKSNGELQDQYQRYDKIQEELSTARDTL 654
>At1g65010 hypothetical protein
Length = 1318
Score = 30.8 bits (68), Expect = 0.13
Identities = 15/54 (27%), Positives = 28/54 (51%)
Query: 8 EKDLDSKRSSQKKLEEELMNVNNQIAELNSETGETVVQQLEEEEEIRVCKNMIK 61
EK+++ S+ E + +V Q+AELN ET ++E+I + + I+
Sbjct: 293 EKEVEESNRSKSSASESMESVMKQLAELNHVLHETKSDNAAQKEKIELLEKTIE 346
Score = 26.6 bits (57), Expect = 2.4
Identities = 16/48 (33%), Positives = 26/48 (53%), Gaps = 1/48 (2%)
Query: 5 SASEKDLDSKRSSQKKLEEELMNVNNQIAELNSETGETVVQQLEEEEE 52
S K+L + ++ K EEL +N + + S+ +TVVQ+ EE E
Sbjct: 830 SEENKELRERETTLLKKAEELSELNESLVDKASKL-QTVVQENEELRE 876
Score = 24.6 bits (52), Expect = 9.0
Identities = 10/70 (14%), Positives = 37/70 (52%)
Query: 10 DLDSKRSSQKKLEEELMNVNNQIAELNSETGETVVQQLEEEEEIRVCKNMIKCTVFSDRP 69
+L+ + ++K ++++ ++ + E ++E+ E L +EE++ C++ + + +
Sbjct: 414 ELERCKVEEEKSKKDMESLTLALQEASTESSEAKATLLVCQEELKNCESQVDSLKLASKE 473
Query: 70 KEVKTGQLIQ 79
K ++++
Sbjct: 474 TNEKYEKMLE 483
>At5g55560 putative protein
Length = 313
Score = 30.4 bits (67), Expect = 0.16
Identities = 19/70 (27%), Positives = 34/70 (48%), Gaps = 1/70 (1%)
Query: 1 MSVASASEKDLDSKRSSQKKLEEELMNVNNQIAEL-NSETGETVVQQLEEEEEIRVCKNM 59
M+ AS+ + + + + S+ +E + + EL S + V + ++EE I V N
Sbjct: 1 MTCASSDDNESEKDKDSESFVEVDPTGRYGRYGELLGSGAVKKVYRAFDQEEGIEVAWNQ 60
Query: 60 IKCTVFSDRP 69
+K FSD P
Sbjct: 61 VKLRCFSDDP 70
>At1g73060 unknown protein
Length = 358
Score = 30.0 bits (66), Expect = 0.21
Identities = 21/66 (31%), Positives = 34/66 (50%), Gaps = 13/66 (19%)
Query: 11 LDSKRSSQKKLEEELMNVNNQIAELNSETGETVVQQLEEEEEIRVCKNMIKCTVFSDRPK 70
L S SS K ++ ++ N Q+A TVV++L+E+ E R C VF D+P+
Sbjct: 84 LPSSISSYKGSSDDFIDANIQLAV-------TVVRKLQEKIETRA------CIVFPDKPE 130
Query: 71 EVKTGQ 76
+ + Q
Sbjct: 131 KRRASQ 136
>At3g32190 hypothetical protein
Length = 358
Score = 29.6 bits (65), Expect = 0.28
Identities = 14/50 (28%), Positives = 31/50 (62%)
Query: 4 ASASEKDLDSKRSSQKKLEEELMNVNNQIAELNSETGETVVQQLEEEEEI 53
+S+ E DL S ++KLE++L N+++++ + N E + + + +EE+
Sbjct: 80 SSSLESDLRSSTEVKQKLEDQLENLSSKLMQSNGELQDQYQRYDKIQEEL 129
>At3g20350 unknown protein
Length = 673
Score = 29.6 bits (65), Expect = 0.28
Identities = 26/78 (33%), Positives = 40/78 (50%), Gaps = 15/78 (19%)
Query: 4 ASASEKDLDS-KRSSQKKLEEELMNVNNQIAELNSETGETVVQQLEEEEEIRVCKNMIKC 62
A A KDL+S KRS +KKLE+ L V+ + A S E E++R + +K
Sbjct: 226 ARACIKDLESEKRSQKKKLEQFLKKVSEERAAWRS----------REHEKVRAIIDDMK- 274
Query: 63 TVFSDRPKEVKTGQLIQL 80
+D +E KT Q +++
Sbjct: 275 ---ADMNQEKKTRQRLEI 289
>At3g16840 ATP-dependent RNA helicase
Length = 832
Score = 29.6 bits (65), Expect = 0.28
Identities = 18/46 (39%), Positives = 24/46 (52%), Gaps = 3/46 (6%)
Query: 8 EKDLDSKRSSQKKLEEELMNVNNQIAELNSETGETVVQQLEEEEEI 53
EK + K+ QKK+ E NQ A S G+ V++ EEEEI
Sbjct: 143 EKKKEKKKKKQKKINEA---AKNQDASAVSCDGDDTVEEQVEEEEI 185
>At2g13410 T26C18.1
Length = 862
Score = 29.6 bits (65), Expect = 0.28
Identities = 14/45 (31%), Positives = 28/45 (62%)
Query: 4 ASASEKDLDSKRSSQKKLEEELMNVNNQIAELNSETGETVVQQLE 48
+S+ E DL S +++KLE++L N+++++ + N E QL+
Sbjct: 360 SSSLESDLRSSNDARQKLEDQLDNLSSELMKSNGAVPELSELQLK 404
>At5g41790 myosin heavy chain-like protein
Length = 1305
Score = 28.9 bits (63), Expect = 0.48
Identities = 12/35 (34%), Positives = 23/35 (65%)
Query: 19 KKLEEELMNVNNQIAELNSETGETVVQQLEEEEEI 53
K+L++E+ + Q+A L+S+ E +Q ++ EEI
Sbjct: 868 KRLDDEVNGLRQQVASLDSQRAELEIQLEKKSEEI 902
Score = 27.3 bits (59), Expect = 1.4
Identities = 10/47 (21%), Positives = 28/47 (59%), Gaps = 4/47 (8%)
Query: 8 EKDLDSKRSSQKKLEEELMNVNNQIAELNSETGETVVQQLEEEEEIR 54
++ LD+ +K L + +++++N+I E +T+ + + E E+++
Sbjct: 417 KQSLDNAEEEKKMLSQRILDISNEI----QEAQKTIQEHMSESEQLK 459
>At3g04990 hypothetical protein
Length = 227
Score = 28.9 bits (63), Expect = 0.48
Identities = 23/80 (28%), Positives = 41/80 (50%), Gaps = 8/80 (10%)
Query: 7 SEKDLDSKRSSQKKLEEELMNVNNQIAELNSETGETVVQQLEEE--EEIRVCKNMIKCTV 64
S ++LD K + L +L ++ SE G+ +++L EE EE+R +N++ +
Sbjct: 35 SSRELDLKEKELQILSSDLEQKSHAFEAEKSEVGD--LKKLVEECTEELRSKRNLLTVKL 92
Query: 65 FS----DRPKEVKTGQLIQL 80
S R E+K QL+Q+
Sbjct: 93 DSLIRVQRELELKDNQLVQV 112
>At5g52230 unknown protein
Length = 746
Score = 28.5 bits (62), Expect = 0.63
Identities = 19/73 (26%), Positives = 36/73 (49%), Gaps = 3/73 (4%)
Query: 8 EKDLDSKRSSQKKLEEELMNVNNQIAELNSETGETVVQQLEEEEEIRVCKNMIKCTVFSD 67
E+D KR ++ K+EE+ ++N +A S+ + +LE E++ + + D
Sbjct: 241 EQDSSEKRITRSKVEEKKNELSNSVARRTSKRLAGI--ELEPTPELKTRAKVQRIVPLDD 298
Query: 68 RP-KEVKTGQLIQ 79
P E+KT +Q
Sbjct: 299 EPTPELKTRTKVQ 311
>At5g33310 putative protein
Length = 1137
Score = 28.5 bits (62), Expect = 0.63
Identities = 13/49 (26%), Positives = 31/49 (62%)
Query: 5 SASEKDLDSKRSSQKKLEEELMNVNNQIAELNSETGETVVQQLEEEEEI 53
S+ E +L S +++KLE++L N+++++ + N E + + + +EE+
Sbjct: 526 SSLESELRSSTDARQKLEDQLDNLSSELMKSNDELQDQYQRYDKIQEEL 574
>At5g14520 pescadillo - like protein
Length = 590
Score = 28.5 bits (62), Expect = 0.63
Identities = 22/66 (33%), Positives = 35/66 (52%), Gaps = 12/66 (18%)
Query: 6 ASEKDLDSKRSSQKKLEEE----LMNVNNQIAELNSETGETVV------QQLEEEEEIRV 55
A E +D+ SSQ EE L + +Q+ +SE G + +++EE+EE RV
Sbjct: 275 AVEPKVDASFSSQSNDREESELRLAQLQHQLP--SSEPGALMHLVADNNKEVEEDEETRV 332
Query: 56 CKNMIK 61
CK++ K
Sbjct: 333 CKSLFK 338
>At4g32700 putative protein
Length = 1548
Score = 28.5 bits (62), Expect = 0.63
Identities = 13/50 (26%), Positives = 24/50 (48%)
Query: 30 NQIAELNSETGETVVQQLEEEEEIRVCKNMIKCTVFSDRPKEVKTGQLIQ 79
N E +S G T ++ L +E+ +RVCK + S +E ++ +
Sbjct: 550 NLRGEYDSRHGSTSMKMLSDEQMLRVCKRFFVALILSKLVQEASVTEVCE 599
>At3g19370 unknown protein
Length = 704
Score = 28.5 bits (62), Expect = 0.63
Identities = 15/45 (33%), Positives = 26/45 (57%), Gaps = 2/45 (4%)
Query: 12 DSKRSSQKKLEEELMNVNNQIAELNS--ETGETVVQQLEEEEEIR 54
D K+ +KKLEE + + N AE+ + E E V ++E E+ ++
Sbjct: 481 DEKQELRKKLEESVEKIRNLEAEMKTLRENKEKVEAEMETEKSMK 525
>At3g52660 putative RNA binding protein
Length = 471
Score = 28.1 bits (61), Expect = 0.82
Identities = 19/54 (35%), Positives = 26/54 (47%), Gaps = 3/54 (5%)
Query: 2 SVASASEKDLDSKRSSQKKLEEELMN---VNNQIAELNSETGETVVQQLEEEEE 52
S+ S DLD ++ LEEE+ +I E+ E E V + EEEEE
Sbjct: 14 SMESEERVDLDGDNDPEEILEEEVEYEEVEEEEIEEIEEEIEEEVEVEEEEEEE 67
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.304 0.122 0.303
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,627,946
Number of Sequences: 26719
Number of extensions: 58075
Number of successful extensions: 524
Number of sequences better than 10.0: 112
Number of HSP's better than 10.0 without gapping: 68
Number of HSP's successfully gapped in prelim test: 45
Number of HSP's that attempted gapping in prelim test: 347
Number of HSP's gapped (non-prelim): 217
length of query: 80
length of database: 11,318,596
effective HSP length: 56
effective length of query: 24
effective length of database: 9,822,332
effective search space: 235735968
effective search space used: 235735968
T: 11
A: 40
X1: 16 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 43 (21.9 bits)
S2: 52 (24.6 bits)
Medicago: description of AC147712.6


