

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147712.16 + phase: 0
(236 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
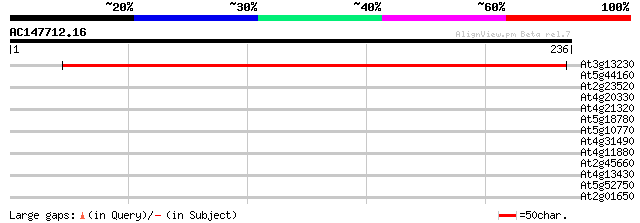
Score E
Sequences producing significant alignments: (bits) Value
At3g13230 unknown protein 333 6e-92
At5g44160 unknown protein 30 1.5
At2g23520 hypothetical protein 29 1.9
At4g20330 putative protein 28 3.2
At4g21320 putative protein 28 4.2
At5g18780 putative protein 28 5.5
At5g10770 nucleoid DNA-binding protein cnd41 - like protein 28 5.5
At4g31490 Beta-COP-like protein 28 5.5
At4g11880 MADS-box protein AGL14 28 5.5
At2g45660 MADS-box protein (AGL20) 28 5.5
At4g13430 aconitate hydratase like protein 27 7.2
At5g52750 unknown protein 27 9.5
At2g01650 unknown protein 27 9.5
>At3g13230 unknown protein
Length = 215
Score = 333 bits (853), Expect = 6e-92
Identities = 160/212 (75%), Positives = 188/212 (88%)
Query: 23 SSSMEVDNVLSEQKVLPPKPKFEPLKPHEMPGAAVQFRKVSVPPHRYTPLKKVWMDIYTP 82
S+ MEV+ LPPKP F+PLK HEM VQFRK++VPP+RY+PLKK W+DIYTP
Sbjct: 4 STQMEVETATEGTVPLPPKPTFKPLKAHEMSDGKVQFRKIAVPPNRYSPLKKAWLDIYTP 63
Query: 83 VFEQMKIDIRMNLKGRKVELKTRHDTPDISNLQKCADFVHAFMLGFDVIDAIAILRLDEL 142
+++QMK+DIRMNLK RKVELKTR DTPDISNLQK ADFVHAFMLGFD+ DAI++LR+DEL
Sbjct: 64 IYDQMKVDIRMNLKARKVELKTRADTPDISNLQKSADFVHAFMLGFDIPDAISLLRMDEL 123
Query: 143 YVESFEIKDVKTLRGDHLSRAIGRLSGKGGKTKFTIENASKTRIVIADTKIHILGSFANI 202
YVESFEIKDVKTL+G+HLSRAIGRLSGKGGKTKF IEN++KTRIVIADT+IHILG+F+NI
Sbjct: 124 YVESFEIKDVKTLKGEHLSRAIGRLSGKGGKTKFAIENSTKTRIVIADTRIHILGAFSNI 183
Query: 203 KIARDSLCYLIMGSPAGKVYSKLRAVTSRMAE 234
K+AR SLC LIMGSPAGKVYSKLR+V++R+ E
Sbjct: 184 KVARSSLCSLIMGSPAGKVYSKLRSVSARLNE 215
>At5g44160 unknown protein
Length = 466
Score = 29.6 bits (65), Expect = 1.5
Identities = 19/49 (38%), Positives = 25/49 (50%), Gaps = 7/49 (14%)
Query: 153 KTLRGDHLSRAIGRLSG-------KGGKTKFTIENASKTRIVIADTKIH 194
KT H SRA+G L+G K G+ K+T E +K V +D K H
Sbjct: 112 KTCVHHHSSRALGDLTGIKKHFCRKHGEKKWTCEKCAKRYAVQSDWKAH 160
>At2g23520 hypothetical protein
Length = 862
Score = 29.3 bits (64), Expect = 1.9
Identities = 18/57 (31%), Positives = 27/57 (46%), Gaps = 3/57 (5%)
Query: 126 LGFDVIDAIAILRLDELYVESFEIKDVKTLRGDHLSRAIGRLSGKGGKTKFTIENAS 182
LG ++ I I+ L + IK+ +L HL R G+ GK G +F + AS
Sbjct: 765 LGIGILSHIRIMDLPRNHRGGARIKEDSSL---HLQREAGKRGGKNGFVRFEVVTAS 818
>At4g20330 putative protein
Length = 286
Score = 28.5 bits (62), Expect = 3.2
Identities = 23/96 (23%), Positives = 41/96 (41%), Gaps = 11/96 (11%)
Query: 60 RKVSVPPHRYTPLKKVWMDIYTPVFEQMKIDIRMNLKGRKVELKTRHDTPDISNLQKCAD 119
R++++ P + V M VF+ ++ + + + GR+ K HD D + L
Sbjct: 88 RRLALTPEQINEWCHVDMHANKAVFDSLRKNPKAHYDGRRFSYKATHDVNDKNQL----- 142
Query: 120 FVHAFMLGFDVIDAIAILRLDELYVESFEIKDVKTL 155
L +D IA++ L + Y E D+K L
Sbjct: 143 ----LSLVRKYLDGIAVVDLKDAYPNVME--DLKAL 172
>At4g21320 putative protein
Length = 1736
Score = 28.1 bits (61), Expect = 4.2
Identities = 14/32 (43%), Positives = 20/32 (61%)
Query: 142 LYVESFEIKDVKTLRGDHLSRAIGRLSGKGGK 173
LYV+ +I D++ LRG HL+ A G + GK
Sbjct: 285 LYVDHSQIMDLECLRGRHLAIAFGFVFIMNGK 316
>At5g18780 putative protein
Length = 441
Score = 27.7 bits (60), Expect = 5.5
Identities = 19/67 (28%), Positives = 30/67 (44%), Gaps = 22/67 (32%)
Query: 89 IDIRMNLKG----RKVELKTRHDTPDISNLQKCADFVHAFMLGFDVIDAIAILRLDELYV 144
+D +N +G R +L T HD DIS L C ++RLD+ +
Sbjct: 70 VDKFLNFRGESYLRGFKLNTDHDVYDISTLDAC------------------LMRLDKCKI 111
Query: 145 ESFEIKD 151
+ FEI++
Sbjct: 112 QHFEIEN 118
>At5g10770 nucleoid DNA-binding protein cnd41 - like protein
Length = 474
Score = 27.7 bits (60), Expect = 5.5
Identities = 18/44 (40%), Positives = 22/44 (49%), Gaps = 1/44 (2%)
Query: 132 DAIAILRLDELYVESFEIKDVKTLRGDHLSRAIGR-LSGKGGKT 174
D + ILRLD+ V S K K L DH+S + L K G T
Sbjct: 83 DHVEILRLDQARVNSIHSKLSKKLATDHVSESKSTDLPAKDGST 126
>At4g31490 Beta-COP-like protein
Length = 958
Score = 27.7 bits (60), Expect = 5.5
Identities = 31/163 (19%), Positives = 66/163 (40%), Gaps = 16/163 (9%)
Query: 7 FFNQLFSCSLNSCTMA-----SSSMEVDNVLSEQKVLPP---KPKFEPLKPHEMPGAAVQ 58
F + +C+L + SS EV+ +S+ ++ + P+ PH + + +
Sbjct: 556 FLGAVVACTLTKLVLRLEEVQSSKTEVNKTVSQALLIMVSILQLGQSPVSPHPIDNDSYE 615
Query: 59 --FRKVSVPPHRYTPLKKVWMDIYTPVFEQMKIDIRM-NLKGRKVELKTRHDTPDISNLQ 115
+ + HR +KK+W++ F +M + ++ ++ K + +T H PD
Sbjct: 616 RIMLCIKLLCHRNVEMKKIWLESCRQSFVKMISEKQLREMEELKAKTQTTHAQPD----- 670
Query: 116 KCADFVHAFMLGFDVIDAIAILRLDELYVESFEIKDVKTLRGD 158
DF H ++ + +L +E D+K G+
Sbjct: 671 DLIDFFHLKSRKMSLLMINFFQGMSQLELEDQVQDDLKRATGE 713
>At4g11880 MADS-box protein AGL14
Length = 221
Score = 27.7 bits (60), Expect = 5.5
Identities = 16/52 (30%), Positives = 28/52 (53%), Gaps = 1/52 (1%)
Query: 172 GKTKFT-IENASKTRIVIADTKIHILGSFANIKIARDSLCYLIMGSPAGKVY 222
GKT+ IENA+ ++ + + +L + + D+ LI+ SP GK+Y
Sbjct: 4 GKTEMKRIENATSRQVTFSKRRNGLLKKAFELSVLCDAEVALIIFSPRGKLY 55
>At2g45660 MADS-box protein (AGL20)
Length = 214
Score = 27.7 bits (60), Expect = 5.5
Identities = 16/52 (30%), Positives = 28/52 (53%), Gaps = 1/52 (1%)
Query: 172 GKTKFT-IENASKTRIVIADTKIHILGSFANIKIARDSLCYLIMGSPAGKVY 222
GKT+ IENA+ ++ + + +L + + D+ LI+ SP GK+Y
Sbjct: 4 GKTQMKRIENATSRQVTFSKRRNGLLKKAFELSVLCDAEVSLIIFSPKGKLY 55
>At4g13430 aconitate hydratase like protein
Length = 509
Score = 27.3 bits (59), Expect = 7.2
Identities = 11/21 (52%), Positives = 13/21 (61%)
Query: 60 RKVSVPPHRYTPLKKVWMDIY 80
RKV VP +KVWMD+Y
Sbjct: 396 RKVKVPTFLVPATQKVWMDVY 416
>At5g52750 unknown protein
Length = 139
Score = 26.9 bits (58), Expect = 9.5
Identities = 14/37 (37%), Positives = 18/37 (47%), Gaps = 4/37 (10%)
Query: 35 QKVLPPKPKFEPLKPHEMPGAAVQFRKVSVPPHRYTP 71
+K PPKP P KP E+ VQ P++Y P
Sbjct: 82 EKPAPPKPAPAPAKPAEIVAWPVQMNN----PYQYNP 114
>At2g01650 unknown protein
Length = 458
Score = 26.9 bits (58), Expect = 9.5
Identities = 16/61 (26%), Positives = 33/61 (53%), Gaps = 5/61 (8%)
Query: 28 VDNVLSEQKVLP--PKPKFEPLKPHE---MPGAAVQFRKVSVPPHRYTPLKKVWMDIYTP 82
+D VL +++V+P P P +P+ + +P A ++FR + +T L+ ++I P
Sbjct: 397 LDPVLVKRRVIPHTPAPGQKPITLEDEELVPSALIRFRPIETDSLVFTGLRNELLEISEP 456
Query: 83 V 83
+
Sbjct: 457 L 457
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.324 0.138 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,068,065
Number of Sequences: 26719
Number of extensions: 208270
Number of successful extensions: 625
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 621
Number of HSP's gapped (non-prelim): 13
length of query: 236
length of database: 11,318,596
effective HSP length: 96
effective length of query: 140
effective length of database: 8,753,572
effective search space: 1225500080
effective search space used: 1225500080
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 58 (26.9 bits)
Medicago: description of AC147712.16


