

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147496.7 + phase: 0
(110 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
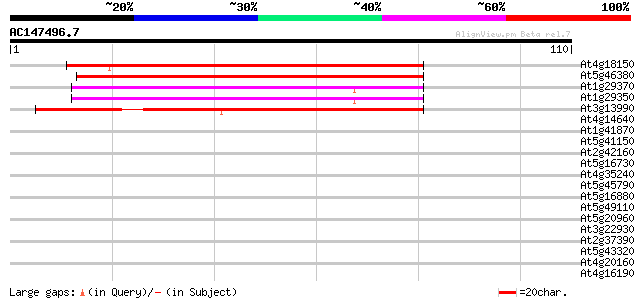
Score E
Sequences producing significant alignments: (bits) Value
At4g18150 hypothetical protein 77 1e-15
At5g46380 putative protein 71 1e-13
At1g29370 unknown protein 64 2e-11
At1g29350 unknown protein 64 2e-11
At3g13990 unknown protein 57 2e-09
At4g14640 calmodulin 30 0.20
At1g41870 hypothetical protein 30 0.20
At5g41150 repair endonuclease (UVH1) 29 0.34
At2g42160 pseudogene 29 0.44
At5g16730 putative protein 28 0.75
At4g35240 putative protein 28 0.98
At5g45790 tyrosine-specific protein phosphatase-like protein 27 1.7
At5g16880 TOM (target of myb1) -like protein 27 1.7
At5g49110 unknown protein 27 2.2
At5g20960 aldehyde oxidase AAO1 27 2.2
At3g22930 calmodulin, putative 27 2.2
At2g37390 unknown protein 27 2.2
At5g43320 casein kinase I 26 3.7
At4g20160 Glu-rich protein 26 3.7
At4g16190 cysteine proteinase like protein 26 3.7
>At4g18150 hypothetical protein
Length = 762
Score = 77.4 bits (189), Expect = 1e-15
Identities = 40/71 (56%), Positives = 52/71 (72%), Gaps = 1/71 (1%)
Query: 12 GGGGGVK-AVKEVVTTAKNVVAAVKEIVNCTEQEIYDVLRECDMDPNLAVEKLLSQDTFR 70
GGGG V A V +++ V+ +KEIV C+E EIY +L ECDM+P+ AV +LLSQDTF
Sbjct: 3 GGGGSVSNANGGVPASSRKVIQDLKEIVECSELEIYAMLVECDMNPDEAVNRLLSQDTFH 62
Query: 71 EVRSKREKRKE 81
E +SKREK+KE
Sbjct: 63 EAKSKREKKKE 73
>At5g46380 putative protein
Length = 607
Score = 70.9 bits (172), Expect = 1e-13
Identities = 33/68 (48%), Positives = 50/68 (73%)
Query: 14 GGGVKAVKEVVTTAKNVVAAVKEIVNCTEQEIYDVLRECDMDPNLAVEKLLSQDTFREVR 73
G V + + +V ++KEIVNC+EQEIY +L EC+M+ + A+ +LLSQD+F+EV+
Sbjct: 2 GSSAIEVNAIPEYTRAIVRSMKEIVNCSEQEIYAMLVECNMNADEAITRLLSQDSFQEVK 61
Query: 74 SKREKRKE 81
SKR+K+KE
Sbjct: 62 SKRDKKKE 69
>At1g29370 unknown protein
Length = 858
Score = 63.5 bits (153), Expect = 2e-11
Identities = 35/97 (36%), Positives = 52/97 (53%), Gaps = 28/97 (28%)
Query: 13 GGGGVKAVKEVVTTAKNVVAAVKEIVNCTEQEIYDVLRECDMDPNLAVEKLLSQ------ 66
GGG K ++++ + ++ +V ++ EIVN E EIY +L+EC+MDPN V +LLSQ
Sbjct: 7 GGGARKGIQDIPSGSRKIVQSLTEIVNSPEAEIYAMLKECNMDPNETVSRLLSQAVAFNM 66
Query: 67 ----------------------DTFREVRSKREKRKE 81
D F EV+SK+EK+KE
Sbjct: 67 VPLVIVQLSSWFGFIEKKDVVNDPFHEVKSKKEKKKE 103
>At1g29350 unknown protein
Length = 858
Score = 63.5 bits (153), Expect = 2e-11
Identities = 35/97 (36%), Positives = 52/97 (53%), Gaps = 28/97 (28%)
Query: 13 GGGGVKAVKEVVTTAKNVVAAVKEIVNCTEQEIYDVLRECDMDPNLAVEKLLSQ------ 66
GGG K ++++ + ++ +V ++ EIVN E EIY +L+EC+MDPN V +LLSQ
Sbjct: 7 GGGARKGIQDIPSGSRKIVQSLTEIVNSPEAEIYAMLKECNMDPNETVSRLLSQAVAFNM 66
Query: 67 ----------------------DTFREVRSKREKRKE 81
D F EV+SK+EK+KE
Sbjct: 67 VPLVIVQLSSWFGFIEKKDVVNDPFHEVKSKKEKKKE 103
>At3g13990 unknown protein
Length = 848
Score = 57.0 bits (136), Expect = 2e-09
Identities = 31/77 (40%), Positives = 48/77 (62%), Gaps = 5/77 (6%)
Query: 6 VVNRNSGGGGGVKAVKEVVTTAKNVVAAVKEIVNC-TEQEIYDVLRECDMDPNLAVEKLL 64
V + G GV E AK ++ ++KE+V+ ++ +IY L+E +MD N AVEKL+
Sbjct: 2 VTGSRTSGNRGVGLDDE----AKKMIQSIKEVVDSHSDADIYTALKEANMDANEAVEKLI 57
Query: 65 SQDTFREVRSKREKRKE 81
QD F EV+ KR+++KE
Sbjct: 58 HQDPFHEVKRKRDRKKE 74
>At4g14640 calmodulin
Length = 151
Score = 30.0 bits (66), Expect = 0.20
Identities = 16/48 (33%), Positives = 26/48 (53%), Gaps = 6/48 (12%)
Query: 14 GGGVKAVKEVVTTAKNVVAAVKEIVNCTEQEIYDVLRECDMDPNLAVE 61
G G V+E+ T +++ N TEQE++D++ E D D N +E
Sbjct: 25 GDGCITVEELATVIRSLDQ------NPTEQELHDIITEIDSDSNGTIE 66
>At1g41870 hypothetical protein
Length = 355
Score = 30.0 bits (66), Expect = 0.20
Identities = 16/44 (36%), Positives = 22/44 (49%)
Query: 25 TTAKNVVAAVKEIVNCTEQEIYDVLRECDMDPNLAVEKLLSQDT 68
T + + EI+N T EI VL ECD+ L ++KL T
Sbjct: 234 TMKDKLAMPISEIINATYPEIPIVLNECDVVMTLDMDKLKKPPT 277
>At5g41150 repair endonuclease (UVH1)
Length = 956
Score = 29.3 bits (64), Expect = 0.34
Identities = 19/95 (20%), Positives = 46/95 (48%), Gaps = 3/95 (3%)
Query: 3 SESVVNRNSGGGGGVKAVKEVVTTAKNVVAAVKEIVNCTEQEIYDVLRECD---MDPNLA 59
+++ + R +GG ++ +V+ + ++++ +++ +I V E + P++
Sbjct: 706 TQNSLTRKAGGRKELEKETQVIVDMREFMSSLPNVLHQKGMKIIPVTLEVGDYILSPSIC 765
Query: 60 VEKLLSQDTFREVRSKREKRKEVILLRYVNIDMRL 94
VE+ QD F+ S R + ++ RY I + L
Sbjct: 766 VERKSIQDLFQSFTSGRLFHQVEMMSRYYRIPVLL 800
>At2g42160 pseudogene
Length = 506
Score = 28.9 bits (63), Expect = 0.44
Identities = 18/60 (30%), Positives = 36/60 (60%), Gaps = 4/60 (6%)
Query: 29 NVVAAVKEIVNCTEQEIYDVLRECDMDPNLAVE---KLL-SQDTFREVRSKREKRKEVIL 84
++ AV++IV T QE+ + + +C+ + + E KL+ QDT+R+ + E+R+ +L
Sbjct: 382 SIAEAVEQIVVNTMQELQNKIEKCEEEKSGITEVNTKLIKEQDTWRKKAKEIEEREAALL 441
>At5g16730 putative protein
Length = 853
Score = 28.1 bits (61), Expect = 0.75
Identities = 19/63 (30%), Positives = 35/63 (55%), Gaps = 2/63 (3%)
Query: 20 VKEVVTTAKNVVAAVKEIVNCTEQEIYDVLRECDMDPNLAVEKLLSQ-DTFREVRSKREK 78
+KE + T + VA KE + +EQ + V E + VEKL S+ +T +E +++ K
Sbjct: 371 LKERIVTLETTVAKQKEDLEVSEQRLGSVEEEVSKNEK-EVEKLKSELETVKEEKNRALK 429
Query: 79 RKE 81
+++
Sbjct: 430 KEQ 432
>At4g35240 putative protein
Length = 828
Score = 27.7 bits (60), Expect = 0.98
Identities = 12/27 (44%), Positives = 15/27 (55%)
Query: 3 SESVVNRNSGGGGGVKAVKEVVTTAKN 29
S + R GGGGG +AV EV +N
Sbjct: 404 SNATATRGGGGGGGPRAVPEVAKEIEN 430
>At5g45790 tyrosine-specific protein phosphatase-like protein
Length = 423
Score = 26.9 bits (58), Expect = 1.7
Identities = 19/58 (32%), Positives = 30/58 (50%), Gaps = 2/58 (3%)
Query: 27 AKNVVAAVKEIVNCTEQEIYDVLRECDMDPNLAVEK--LLSQDTFREVRSKREKRKEV 82
+KN+ + E E +V+REC++D V+K LLS+ V + R RKE+
Sbjct: 124 SKNLCCKITEKFKVLIDEEENVVRECEVDAVKRVKKLLLLSKHGVLRVHALRLIRKEL 181
>At5g16880 TOM (target of myb1) -like protein
Length = 407
Score = 26.9 bits (58), Expect = 1.7
Identities = 19/77 (24%), Positives = 37/77 (47%), Gaps = 7/77 (9%)
Query: 15 GGVKAVKEVVTTAKNVVAAVKEIV---NCTEQEIYDV----LRECDMDPNLAVEKLLSQD 67
GG + ++ ++ VKE+ N T++ + D L E D D NL + +++Q+
Sbjct: 19 GGSEVSNKISAGVSSMSFKVKELFQGPNPTDKIVEDATTENLEEPDWDMNLEICDMINQE 78
Query: 68 TFREVRSKREKRKEVIL 84
T V R +K +++
Sbjct: 79 TINSVELIRGIKKRIMM 95
>At5g49110 unknown protein
Length = 1487
Score = 26.6 bits (57), Expect = 2.2
Identities = 9/33 (27%), Positives = 20/33 (60%)
Query: 20 VKEVVTTAKNVVAAVKEIVNCTEQEIYDVLREC 52
VKEV + ++++ V ++ E +++D+L C
Sbjct: 127 VKEVKSVVDSILSGVTRVITVDEAQVFDLLPVC 159
>At5g20960 aldehyde oxidase AAO1
Length = 1368
Score = 26.6 bits (57), Expect = 2.2
Identities = 14/30 (46%), Positives = 17/30 (56%)
Query: 6 VVNRNSGGGGGVKAVKEVVTTAKNVVAAVK 35
V+ R GGG G KAVK + A +AA K
Sbjct: 823 VITRRVGGGFGGKAVKSMPVAAACALAASK 852
>At3g22930 calmodulin, putative
Length = 173
Score = 26.6 bits (57), Expect = 2.2
Identities = 13/37 (35%), Positives = 21/37 (56%), Gaps = 1/37 (2%)
Query: 26 TAKNVVAAVKEI-VNCTEQEIYDVLRECDMDPNLAVE 61
TA + ++ + N TEQE+ D++ E D D N +E
Sbjct: 52 TADELATVIRSLDQNPTEQELQDMITEIDSDGNGTIE 88
>At2g37390 unknown protein
Length = 259
Score = 26.6 bits (57), Expect = 2.2
Identities = 14/52 (26%), Positives = 27/52 (51%), Gaps = 6/52 (11%)
Query: 2 GSESVVNRNS---GGGGGVKAVKEVVTTAKNVVAAVKEIVNCTEQEIYDVLR 50
G S+ R S GGGGG A+ +++T N + ++ +C + D+++
Sbjct: 79 GRRSITGRRSTGGGGGGGAAALLKLIT---NDIGLARKSFSCVARPACDLIK 127
>At5g43320 casein kinase I
Length = 480
Score = 25.8 bits (55), Expect = 3.7
Identities = 12/33 (36%), Positives = 20/33 (60%)
Query: 4 ESVVNRNSGGGGGVKAVKEVVTTAKNVVAAVKE 36
E+ RN G G V+A + T++NV+A+ K+
Sbjct: 346 EAYARRNGSGSGVVQADRSRPRTSENVLASSKD 378
>At4g20160 Glu-rich protein
Length = 1188
Score = 25.8 bits (55), Expect = 3.7
Identities = 20/74 (27%), Positives = 37/74 (49%), Gaps = 8/74 (10%)
Query: 16 GVKAVKEVVTTAKN----VVAAVKEIVNCTEQEIYDVLR----ECDMDPNLAVEKLLSQD 67
GVK +KEVV+ +N V++ + E N ++ + ++ E + EK++S++
Sbjct: 395 GVKEIKEVVSAVENAKKGVLSEISENRNGLKKLAGETIQKETVEGKGEKRETKEKVISKE 454
Query: 68 TFREVRSKREKRKE 81
+ KRE KE
Sbjct: 455 SLEGKGEKRESTKE 468
>At4g16190 cysteine proteinase like protein
Length = 373
Score = 25.8 bits (55), Expect = 3.7
Identities = 10/21 (47%), Positives = 15/21 (70%)
Query: 33 AVKEIVNCTEQEIYDVLRECD 53
A KE+V+ +EQ++ D ECD
Sbjct: 180 ATKELVSLSEQQLVDCDHECD 200
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.135 0.369
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,198,096
Number of Sequences: 26719
Number of extensions: 79276
Number of successful extensions: 417
Number of sequences better than 10.0: 44
Number of HSP's better than 10.0 without gapping: 25
Number of HSP's successfully gapped in prelim test: 19
Number of HSP's that attempted gapping in prelim test: 388
Number of HSP's gapped (non-prelim): 48
length of query: 110
length of database: 11,318,596
effective HSP length: 86
effective length of query: 24
effective length of database: 9,020,762
effective search space: 216498288
effective search space used: 216498288
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 52 (24.6 bits)
Medicago: description of AC147496.7


