

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147496.2 - phase: 0 /pseudo
(550 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
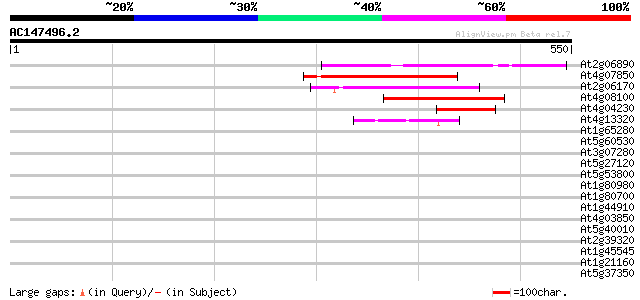
Score E
Sequences producing significant alignments: (bits) Value
At2g06890 putative retroelement integrase 178 8e-45
At4g07850 putative polyprotein 171 1e-42
At2g06170 putative Ty3-gypsy-like retroelement pol polyprotein 118 1e-26
At4g08100 putative polyprotein 117 1e-26
At4g04230 putative transposon protein 64 2e-10
At4g13320 hypothetical protein 45 1e-04
At1g65280 35 0.083
At5g60530 late embryonic abundant protein - like 34 0.18
At3g07280 unknown protein 34 0.18
At5g27120 SAR DNA-binding protein - like 33 0.31
At5g53800 unknown protein 32 0.70
At1g80980 hypothetical protein 32 0.70
At1g80700 hypothetical protein 32 0.70
At1g44910 splicing factor like protein 32 0.70
At4g03850 putative transposon protein 32 0.91
At5g40010 putative protein 32 1.2
At2g39320 hypothetical protein 32 1.2
At1g45545 hypothetical protein 32 1.2
At1g21160 transcription factor, putative 32 1.2
At5g37350 unknown protein 31 1.6
>At2g06890 putative retroelement integrase
Length = 1215
Score = 178 bits (451), Expect = 8e-45
Identities = 109/243 (44%), Positives = 145/243 (58%), Gaps = 19/243 (7%)
Query: 306 EEAEEIPS-GDLFMIRRFLGNQAKEEESNQRETLFHTRCLVQGKVCFLIIDGGSRTNVAS 364
+E EE+P+ G+L + RR L Q K +E QR+ LFHTRC V GKVC LIIDGGS TNVAS
Sbjct: 200 KENEELPAQGELLVARRTLSVQTKTDEQEQRKNLFHTRCHVHGKVCSLIIDGGSCTNVAS 259
Query: 365 TRLVSKMELETKPHPKPYKLQWLNENVEILVDKQVEVCFKIGKYEDDVLCDVVPMEASHL 424
+V K+ L+ WLN++ ++ V QV V IGKYED++LCDV+PMEA H+
Sbjct: 260 ETMVKKLGLK-----------WLNDSGKMRVKNQVVVPIVIGKYEDEILCDVLPMEAGHI 308
Query: 425 LLGRP*QFDRSVLHDGRTNKYSFMHSGQKISLAPLSPSEVRDDQKKRKEKYEKEKRKIKR 484
LLGRP Q DR V+HDG TN++SF G K L ++P EV DQ K+K K ++ +
Sbjct: 309 LLGRPWQSDRKVMHDGFTNRHSFEFKGGKTILVSMTPHEVYQDQIHLKQK----KEQVVK 364
Query: 485 KEREFA*KKKKRRVSRNSHEQTAPIFTTLQKVALLTN-TQNFPSCTKFLLQECEDVFPKK 543
+ FA K S S +Q +F + + LTN PS LLQ+ +DVFP+
Sbjct: 365 QPNFFA--KSGEVKSAYSSKQPMLLFVFKEALTSLTNFAPVLPSEMTSLLQDYKDVFPED 422
Query: 544 VPQ 546
P+
Sbjct: 423 NPK 425
>At4g07850 putative polyprotein
Length = 1138
Score = 171 bits (432), Expect = 1e-42
Identities = 88/151 (58%), Positives = 105/151 (69%), Gaps = 3/151 (1%)
Query: 289 DLGE*NVEVENHEGYVEEEAEEIPSGDLFMIRRFLGNQAKEEESNQRETLFHTRCLVQGK 348
D GE E E E E + EE P G+L + R L K EE QRE LFHTRCL++GK
Sbjct: 303 DSGEVESEDEKPE---ESDVEEAPKGELLVTMRVLSVLNKAEEQAQRENLFHTRCLIKGK 359
Query: 349 VCFLIIDGGSRTNVASTRLVSKMELETKPHPKPYKLQWLNENVEILVDKQVEVCFKIGKY 408
VC LIIDGGS TNVAS +V K+ LE PHPKPYKLQWLNE+ E+ V +QV+V IGKY
Sbjct: 360 VCSLIIDGGSCTNVASETMVQKLGLEEFPHPKPYKLQWLNESGEMAVTRQVQVPLAIGKY 419
Query: 409 EDDVLCDVVPMEASHLLLGRP*QFDRSVLHD 439
ED++LCD++P+EASH+LLGRP Q DR D
Sbjct: 420 EDEILCDILPLEASHVLLGRPWQSDRKDYQD 450
>At2g06170 putative Ty3-gypsy-like retroelement pol polyprotein
Length = 587
Score = 118 bits (295), Expect = 1e-26
Identities = 64/168 (38%), Positives = 96/168 (57%), Gaps = 6/168 (3%)
Query: 296 EVENHEGYVEEEAEEIPSGDLF--MIRRFLGNQAKEEESNQRETLFHTRCLVQGKVCFLI 353
E+E + Y E E E S ++ +++R L +E QR LF TRC + KVC LI
Sbjct: 183 ELEEEDEYAEVEFAEEESNEMINLVLQRIL---LSSKEEGQRRNLFRTRCSINDKVCNLI 239
Query: 354 IDGGSRTNVASTRLVSKMELETKPHPKPYKLQWLNENVEILVDKQVEVCFKIGK-YEDDV 412
+D GS N+ S +LV ++L T H KPY L W+++ + V V IGK Y+++V
Sbjct: 240 VDIGSSENLVSQKLVEYLKLPTTLHQKPYSLGWVSKGSQFCVSLSCRVPISIGKHYKEEV 299
Query: 413 LCDVVPMEASHLLLGRP*QFDRSVLHDGRTNKYSFMHSGQKISLAPLS 460
LCDV+ M+ H++LGR Q+D + + G+ N F +G KI +AP+S
Sbjct: 300 LCDVLNMDVCHIILGRSWQYDNDITYRGKDNVLMFTWNGHKIVMAPVS 347
>At4g08100 putative polyprotein
Length = 1054
Score = 117 bits (294), Expect = 1e-26
Identities = 57/119 (47%), Positives = 79/119 (65%)
Query: 367 LVSKMELETKPHPKPYKLQWLNENVEILVDKQVEVCFKIGKYEDDVLCDVVPMEASHLLL 426
+V K+ LE HP+PY LQW NE E+ V +QV+V IGKY D+++CD++ M+ASH+LL
Sbjct: 388 MVEKLGLEVLKHPRPYSLQWRNETGEMSVKEQVKVPLSIGKYHDEIMCDILHMDASHILL 447
Query: 427 GRP*QFDRSVLHDGRTNKYSFMHSGQKISLAPLSPSEVRDDQKKRKEKYEKEKRKIKRK 485
GRP Q DR VL DG TN+ +F H+G+K +L P++ EV DQ K++ K I K
Sbjct: 448 GRPWQSDRKVLQDGFTNRQTFEHNGRKTTLIPMTLHEVYLDQLSMKQRAIKPTEPIDTK 506
>At4g04230 putative transposon protein
Length = 315
Score = 64.3 bits (155), Expect = 2e-10
Identities = 31/58 (53%), Positives = 40/58 (68%)
Query: 419 MEASHLLLGRP*QFDRSVLHDGRTNKYSFMHSGQKISLAPLSPSEVRDDQKKRKEKYE 476
M HLLLGRP QFDR+ H+ RTN YSF ++ +K +LAPLSP EV D Q +++E
Sbjct: 1 MRVGHLLLGRPWQFDRATCHNRRTNHYSFTYNDRKYNLAPLSPLEVHDLQIHMNKEHE 58
>At4g13320 hypothetical protein
Length = 216
Score = 45.1 bits (105), Expect = 1e-04
Identities = 34/108 (31%), Positives = 56/108 (51%), Gaps = 8/108 (7%)
Query: 338 LFHTRCLVQGKVCFLIIDGGSRTNVASTRLVSKMELET-KPHPKPYKLQWLNENVEILVD 396
+F T+C++ + C L++ GG+ N+ S LV +++L+T K +P + E + + +
Sbjct: 98 VFRTQCVINDEACRLVLYGGN--NIISKGLVKQLKLKTLKKYPSVRVMATRRE--DKVAE 153
Query: 397 KQVEVCFKIGK-YEDDVLCDVVPM--EASHLLLGRP*QFDRSVLHDGR 441
+ V IG Y+D V C VV M E LL G P + H+GR
Sbjct: 154 ETCRVPVSIGDFYKDKVTCYVVNMEEEEDQLLFGGPWLYRVQATHNGR 201
>At1g65280
Length = 605
Score = 35.4 bits (80), Expect = 0.083
Identities = 14/37 (37%), Positives = 27/37 (72%)
Query: 468 QKKRKEKYEKEKRKIKRKEREFA*KKKKRRVSRNSHE 504
+K+RK + E++++KI+RKER+ KKK++ + +E
Sbjct: 56 EKRRKREKERKRKKIERKERKRRDMKKKKKTKKREYE 92
>At5g60530 late embryonic abundant protein - like
Length = 439
Score = 34.3 bits (77), Expect = 0.18
Identities = 15/40 (37%), Positives = 27/40 (67%)
Query: 468 QKKRKEKYEKEKRKIKRKEREFA*KKKKRRVSRNSHEQTA 507
+K++K+K EKEK+ +RKE+E K++K + ++ E A
Sbjct: 95 EKEKKDKLEKEKKDKERKEKERKEKERKAKEKKDKEESEA 134
Score = 30.8 bits (68), Expect = 2.0
Identities = 13/41 (31%), Positives = 26/41 (62%)
Query: 465 RDDQKKRKEKYEKEKRKIKRKEREFA*KKKKRRVSRNSHEQ 505
++ +KK KEK K+K++ ++K++E KK K R + ++
Sbjct: 61 KEQEKKDKEKAAKDKKEKEKKDKEEKEKKDKERKEKEKKDK 101
Score = 30.8 bits (68), Expect = 2.0
Identities = 14/38 (36%), Positives = 25/38 (64%)
Query: 468 QKKRKEKYEKEKRKIKRKEREFA*KKKKRRVSRNSHEQ 505
+K++K+K EKEK+ +RKE+E K +K + + E+
Sbjct: 77 EKEKKDKEEKEKKDKERKEKEKKDKLEKEKKDKERKEK 114
Score = 30.4 bits (67), Expect = 2.7
Identities = 19/79 (24%), Positives = 39/79 (49%)
Query: 427 GRP*QFDRSVLHDGRTNKYSFMHSGQKISLAPLSPSEVRDDQKKRKEKYEKEKRKIKRKE 486
G Q D+ +G++N Q+ + + ++ +KK KE+ EK+ ++ K KE
Sbjct: 38 GNEVQVDKGKGDNGKSNGNGPKDKEQEKKDKEKAAKDKKEKEKKDKEEKEKKDKERKEKE 97
Query: 487 REFA*KKKKRRVSRNSHEQ 505
++ +K+K+ R E+
Sbjct: 98 KKDKLEKEKKDKERKEKER 116
>At3g07280 unknown protein
Length = 481
Score = 34.3 bits (77), Expect = 0.18
Identities = 19/56 (33%), Positives = 32/56 (56%)
Query: 467 DQKKRKEKYEKEKRKIKRKEREFA*KKKKRRVSRNSHEQTAPIFTTLQKVALLTNT 522
++K+ KEK EKEK K K KER+ +K K R ++ ++ + + +L+NT
Sbjct: 39 EKKEGKEKREKEKSKDKHKERKERKEKHKDRKDKDRDKEKSRTSEDRKAAGVLSNT 94
Score = 29.6 bits (65), Expect = 4.5
Identities = 15/60 (25%), Positives = 32/60 (53%)
Query: 463 EVRDDQKKRKEKYEKEKRKIKRKEREFA*KKKKRRVSRNSHEQTAPIFTTLQKVALLTNT 522
E R+ +K + + E+++RK K K+R+ + K++ + + + T + L+TNT
Sbjct: 45 EKREKEKSKDKHKERKERKEKHKDRKDKDRDKEKSRTSEDRKAAGVLSNTGDREKLVTNT 104
>At5g27120 SAR DNA-binding protein - like
Length = 533
Score = 33.5 bits (75), Expect = 0.31
Identities = 15/46 (32%), Positives = 29/46 (62%)
Query: 452 QKISLAPLSPSEVRDDQKKRKEKYEKEKRKIKRKEREFA*KKKKRR 497
+K P + E +KK+K K+E+E+ ++ K++E + KKKK++
Sbjct: 485 KKTEAEPETAEEPAKKEKKKKRKHEEEETEMPAKKKEKSEKKKKKK 530
>At5g53800 unknown protein
Length = 339
Score = 32.3 bits (72), Expect = 0.70
Identities = 18/38 (47%), Positives = 24/38 (62%), Gaps = 1/38 (2%)
Query: 460 SPSEVRDDQKKRKEKYEKEKRKIKRKEREFA*KKKKRR 497
S SE D ++ E E+ +RK KRKERE K++KRR
Sbjct: 114 SESEYSDSEESESED-ERRRRKRKRKEREEEEKERKRR 150
>At1g80980 hypothetical protein
Length = 214
Score = 32.3 bits (72), Expect = 0.70
Identities = 20/79 (25%), Positives = 38/79 (47%)
Query: 468 QKKRKEKYEKEKRKIKRKEREFA*KKKKRRVSRNSHEQTAPIFTTLQKVALLTNTQNFPS 527
+KKRKE+ +KEK + ++K + K ++ I T++++ L T + +
Sbjct: 119 EKKRKEEEKKEKEEAEQKALQVEAATKSHEELMEMRQRLGKIEETIKEIVLETKKPSGNA 178
Query: 528 CTKFLLQECEDVFPKKVPQ 546
TK + + PK+V Q
Sbjct: 179 PTKTQEDQSTKLSPKEVSQ 197
>At1g80700 hypothetical protein
Length = 214
Score = 32.3 bits (72), Expect = 0.70
Identities = 20/79 (25%), Positives = 38/79 (47%)
Query: 468 QKKRKEKYEKEKRKIKRKEREFA*KKKKRRVSRNSHEQTAPIFTTLQKVALLTNTQNFPS 527
+KKRKE+ +KEK + ++K + K ++ I T++++ L T + +
Sbjct: 119 EKKRKEEEKKEKEEAEQKALQVEAATKSHEELMEMRQRLGKIEETIKEIVLETKKPSGNA 178
Query: 528 CTKFLLQECEDVFPKKVPQ 546
TK + + PK+V Q
Sbjct: 179 PTKTQEDQSTKLSPKEVSQ 197
>At1g44910 splicing factor like protein
Length = 958
Score = 32.3 bits (72), Expect = 0.70
Identities = 20/44 (45%), Positives = 27/44 (60%), Gaps = 5/44 (11%)
Query: 465 RDDQKKRKEKY--EKEKRKIKRKEREFA*KKKKRRVSRNSHEQT 506
RD++K RKEK EKEKRK K KER +K++ R E++
Sbjct: 813 RDEEKVRKEKERDEKEKRKDKDKERR---EKEREREKEKGKERS 853
Score = 29.3 bits (64), Expect = 5.9
Identities = 12/35 (34%), Positives = 24/35 (68%)
Query: 465 RDDQKKRKEKYEKEKRKIKRKEREFA*KKKKRRVS 499
RD+++KRK+K ++ + K + +E+E ++ KR S
Sbjct: 824 RDEKEKRKDKDKERREKEREREKEKGKERSKREES 858
>At4g03850 putative transposon protein
Length = 334
Score = 32.0 bits (71), Expect = 0.91
Identities = 36/117 (30%), Positives = 53/117 (44%), Gaps = 20/117 (17%)
Query: 416 VVPMEASH---LLLGRP*QFDRSVLHDGRTNKYSFMHSGQKISLA---PLSPSEVRDDQK 469
V+ ME H L+LGRP + D R K S ++ G+ I L +P E ++D+K
Sbjct: 153 VLDMEVEHKDPLILGRPFLASVGAVIDVREGKIS-LNLGKHIMLQFDINKTPQESKEDEK 211
Query: 470 KRK-------EKYEKEKRKIKRKEREFA*KKKKRRVSRNSH--EQTAPIFTTLQKVA 517
EKYE EK K +K + K++ + R +H E+ LQK A
Sbjct: 212 TSGDDRVIPGEKYETEKVKELKKRSD----KQEETIERLAHSVEELRSKLNQLQKEA 264
>At5g40010 putative protein
Length = 514
Score = 31.6 bits (70), Expect = 1.2
Identities = 17/47 (36%), Positives = 30/47 (63%), Gaps = 3/47 (6%)
Query: 463 EVRDDQKKR---KEKYEKEKRKIKRKEREFA*KKKKRRVSRNSHEQT 506
E +++ K+R +EK +KE+ +IKRK+RE KK+ + + +E T
Sbjct: 465 EEKEEAKRRIEDEEKKKKEEEEIKRKKREEKKIKKEEKEEKEENETT 511
>At2g39320 hypothetical protein
Length = 189
Score = 31.6 bits (70), Expect = 1.2
Identities = 13/39 (33%), Positives = 27/39 (68%), Gaps = 5/39 (12%)
Query: 465 RDDQKKRKEKYEKEKRKIKRKEREFA*KKKKRRVSRNSH 503
+D + K+K+K +K+K K++++++E KK + +RN H
Sbjct: 151 KDKEDKKKDKEDKKKAKVQKEKKE-----KKEKKNRNHH 184
Score = 28.9 bits (63), Expect = 7.7
Identities = 12/40 (30%), Positives = 26/40 (65%)
Query: 466 DDQKKRKEKYEKEKRKIKRKEREFA*KKKKRRVSRNSHEQ 505
+++K+RK+ ++EK+K K +++ KKK +V + E+
Sbjct: 136 EEEKERKDMEKEEKKKDKEDKKKDKEDKKKAKVQKEKKEK 175
>At1g45545 hypothetical protein
Length = 752
Score = 31.6 bits (70), Expect = 1.2
Identities = 17/76 (22%), Positives = 34/76 (44%)
Query: 465 RDDQKKRKEKYEKEKRKIKRKEREFA*KKKKRRVSRNSHEQTAPIFTTLQKVALLTNTQN 524
R++ +K KE+ ++ K + ER+ KK+ SR S + LQ+ ++
Sbjct: 520 REELRKAKEESDEAKTGLSAVERQLMESKKEMEASRASEKLALAAIKALQETEYANKIED 579
Query: 525 FPSCTKFLLQECEDVF 540
S K ++ E+ +
Sbjct: 580 ISSSPKSIIISVEEYY 595
>At1g21160 transcription factor, putative
Length = 1088
Score = 31.6 bits (70), Expect = 1.2
Identities = 19/57 (33%), Positives = 31/57 (54%), Gaps = 4/57 (7%)
Query: 463 EVRDDQKKRKE----KYEKEKRKIKRKEREFA*KKKKRRVSRNSHEQTAPIFTTLQK 515
E D +KK +E K E+E+R + +ERE ++KR++ + +Q I T QK
Sbjct: 234 EAEDGKKKEEEERLRKEEEERRIEEEREREAEEIRQKRKIRKMEKKQEGLILTAKQK 290
>At5g37350 unknown protein
Length = 531
Score = 31.2 bits (69), Expect = 1.6
Identities = 13/42 (30%), Positives = 29/42 (68%)
Query: 459 LSPSEVRDDQKKRKEKYEKEKRKIKRKEREFA*KKKKRRVSR 500
L P + + +K+ K+K ++EKR+ ++ + + KK+K++VS+
Sbjct: 485 LGPEDKKAARKEHKKKVKEEKRESRKTKTPKSVKKRKKKVSK 526
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.358 0.160 0.569
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,770,819
Number of Sequences: 26719
Number of extensions: 435270
Number of successful extensions: 5060
Number of sequences better than 10.0: 43
Number of HSP's better than 10.0 without gapping: 28
Number of HSP's successfully gapped in prelim test: 16
Number of HSP's that attempted gapping in prelim test: 4605
Number of HSP's gapped (non-prelim): 256
length of query: 550
length of database: 11,318,596
effective HSP length: 104
effective length of query: 446
effective length of database: 8,539,820
effective search space: 3808759720
effective search space used: 3808759720
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 63 (28.9 bits)
Medicago: description of AC147496.2


