

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147484.9 - phase: 0
(125 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
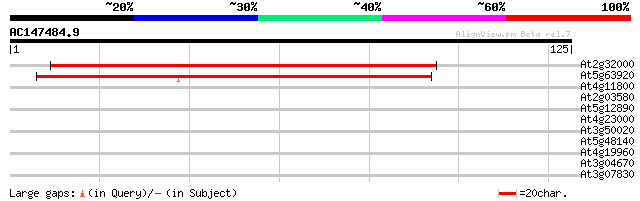
Score E
Sequences producing significant alignments: (bits) Value
At2g32000 putative DNA topoisomerase III beta 136 2e-33
At5g63920 DNA topoisomerase III 79 6e-16
At4g11800 putative protein 31 0.14
At2g03580 hypothetical protein 30 0.41
At5g12890 glucosyltransferase -like protein 28 1.2
At4g23000 hypothetical protein 27 2.6
At3g50020 putative protein 27 3.4
At5g48140 polygalacturonase 26 4.5
At4g19960 potassium transporter-like protein 26 4.5
At3g04670 unknown protein 26 4.5
At3g07830 polygalacturonase like protein 25 7.7
>At2g32000 putative DNA topoisomerase III beta
Length = 836
Score = 136 bits (343), Expect = 2e-33
Identities = 66/86 (76%), Positives = 74/86 (85%)
Query: 10 VAEKPSIALSIATVLSHGKMNTRRGSTEVHEFDGKFRKAPARFKVTSVIGHVFSVDFPGK 69
VAEKPSIALSIA+VLSHG+M+TRRGSTEVHEFDG FR A ++VTSVIGHVFSVDFP K
Sbjct: 40 VAEKPSIALSIASVLSHGQMSTRRGSTEVHEFDGMFRGFKAHYRVTSVIGHVFSVDFPEK 99
Query: 70 YQDWAATDPLDLFQAQVIKNESNPKL 95
YQ+WA DP DLF A +IK ESNPK+
Sbjct: 100 YQNWATIDPQDLFDAPIIKKESNPKV 125
>At5g63920 DNA topoisomerase III
Length = 926
Score = 79.0 bits (193), Expect = 6e-16
Identities = 40/91 (43%), Positives = 56/91 (60%), Gaps = 3/91 (3%)
Query: 7 VLMVAEKPSIALSIATVLSHGKMNTRRGST---EVHEFDGKFRKAPARFKVTSVIGHVFS 63
VL VAEKPS+A S+A +LS G TR G + ++ EFD P R +TSVIGH+
Sbjct: 11 VLNVAEKPSVAKSVAGILSRGTFRTREGRSRYNKIFEFDYAINGQPCRMLMTSVIGHLME 70
Query: 64 VDFPGKYQDWAATDPLDLFQAQVIKNESNPK 94
++F +Y+ W + DP DL+QA V+K+ K
Sbjct: 71 LEFADRYRKWHSCDPADLYQAPVMKHVPEDK 101
>At4g11800 putative protein
Length = 1012
Score = 31.2 bits (69), Expect = 0.14
Identities = 15/47 (31%), Positives = 26/47 (54%), Gaps = 1/47 (2%)
Query: 73 WAATDPLDLFQAQVIKNES-NPKLVDVVIWYCGWIVTVKEKIYVLRV 118
WA T PL + + + +KN+ P +D+V WY G + + ++ L V
Sbjct: 281 WALTHPLSVDKYEKLKNQQLKPDFLDMVPWYSGTSADLFKTVFDLLV 327
>At2g03580 hypothetical protein
Length = 144
Score = 29.6 bits (65), Expect = 0.41
Identities = 12/34 (35%), Positives = 18/34 (52%)
Query: 87 IKNESNPKLVDVVIWYCGWIVTVKEKIYVLRVIF 120
++ E PK+ D WY W VK +Y R+I+
Sbjct: 5 MREEKRPKVFDNHDWYFVWKTCVKRPLYPWRIIY 38
>At5g12890 glucosyltransferase -like protein
Length = 488
Score = 28.1 bits (61), Expect = 1.2
Identities = 12/44 (27%), Positives = 24/44 (54%)
Query: 77 DPLDLFQAQVIKNESNPKLVDVVIWYCGWIVTVKEKIYVLRVIF 120
+P F +++K E ++ + ++ GWI V +++ V VIF
Sbjct: 109 EPFRDFMTKILKEEGQSSVIVIGDFFLGWIGKVCKEVGVYSVIF 152
>At4g23000 hypothetical protein
Length = 932
Score = 26.9 bits (58), Expect = 2.6
Identities = 13/47 (27%), Positives = 23/47 (48%), Gaps = 1/47 (2%)
Query: 73 WAATDPLDLFQAQVIKNES-NPKLVDVVIWYCGWIVTVKEKIYVLRV 118
WA PL + + +K + P +D+V WY G + + ++ L V
Sbjct: 324 WALAHPLSVENYEKLKRQQMKPNFLDMVPWYSGTSADLFKTVFDLLV 370
>At3g50020 putative protein
Length = 96
Score = 26.6 bits (57), Expect = 3.4
Identities = 13/35 (37%), Positives = 19/35 (54%)
Query: 66 FPGKYQDWAATDPLDLFQAQVIKNESNPKLVDVVI 100
FPG DW + + +A+ I NPK+V V+I
Sbjct: 26 FPGIKVDWPELNGVKGLEAKRIIEHDNPKVVVVII 60
>At5g48140 polygalacturonase
Length = 395
Score = 26.2 bits (56), Expect = 4.5
Identities = 14/41 (34%), Positives = 23/41 (55%), Gaps = 2/41 (4%)
Query: 82 FQAQVIKNESNPKLVDVVIWYCGWIVTVKEKIYVLRVIFVS 122
F+ ++KN SNP L+D YC W K+K +++ +S
Sbjct: 285 FENILLKNVSNPILIDQE--YCPWNQCNKQKPSTIKLANIS 323
>At4g19960 potassium transporter-like protein
Length = 842
Score = 26.2 bits (56), Expect = 4.5
Identities = 10/31 (32%), Positives = 19/31 (61%), Gaps = 3/31 (9%)
Query: 95 LVDVVIWYCGWIVTVKEKIYVLRVIFVSLSF 125
L+ +++W+C WI+ + I+ FV LS+
Sbjct: 504 LIMLLVWHCHWILVL---IFTFLSFFVELSY 531
>At3g04670 unknown protein
Length = 330
Score = 26.2 bits (56), Expect = 4.5
Identities = 19/59 (32%), Positives = 26/59 (43%), Gaps = 9/59 (15%)
Query: 43 GKFRKAPARFKVTSVIGHVF-------SVDFPGKYQDWAATDPLDLFQAQVIKNESNPK 94
GKFR +F+ +S H+F D G Y A +PL + A + NE PK
Sbjct: 57 GKFRSTN-KFR-SSFPQHIFLESPICCGNDLSGDYTQVLAPEPLQMVPASAVYNEMEPK 113
>At3g07830 polygalacturonase like protein
Length = 397
Score = 25.4 bits (54), Expect = 7.7
Identities = 13/41 (31%), Positives = 23/41 (55%), Gaps = 2/41 (4%)
Query: 82 FQAQVIKNESNPKLVDVVIWYCGWIVTVKEKIYVLRVIFVS 122
F+ +++N SNP L+D YC W K+K +++ +S
Sbjct: 286 FENIILRNVSNPILIDQE--YCPWNQCNKQKSSSIKLANIS 324
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.323 0.137 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,612,986
Number of Sequences: 26719
Number of extensions: 95958
Number of successful extensions: 236
Number of sequences better than 10.0: 11
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 228
Number of HSP's gapped (non-prelim): 12
length of query: 125
length of database: 11,318,596
effective HSP length: 87
effective length of query: 38
effective length of database: 8,994,043
effective search space: 341773634
effective search space used: 341773634
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 54 (25.4 bits)
Medicago: description of AC147484.9


