

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147484.3 - phase: 0 /pseudo
(526 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
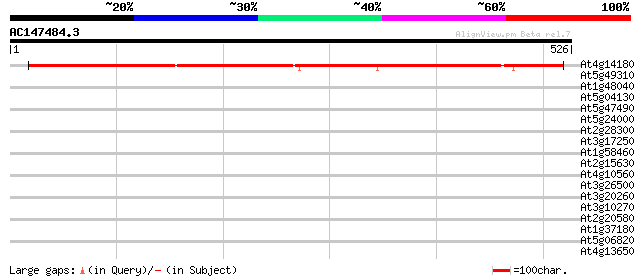
Score E
Sequences producing significant alignments: (bits) Value
At4g14180 hypothetical protein 489 e-138
At5g49310 importin alpha 36 0.046
At1g48040 protein phosphatase-2C, putative 34 0.23
At5g04130 DNA gyrase subunit B - like protein 33 0.51
At5g47490 putative protein 31 1.9
At5g24000 putative protein 31 1.9
At2g28300 unknown protein 31 1.9
At3g17250 protein phosphatase like 30 2.5
At1g58460 unknown protein 30 2.5
At2g15630 putative salt-inducible protein 30 4.3
At4g10560 putative protein 29 5.6
At3g26500 unknown protein 29 5.6
At3g20260 unknown protein 29 5.6
At3g10270 putative DNA gyrase subunit B 29 5.6
At2g20580 putative 26S proteasome regulatory subunit S2 29 7.3
At1g37180 hypothetical protein 29 7.3
At5g06820 SRF2 (LRR receptor protein kinase like) 28 9.6
At4g13650 unknown protein 28 9.6
>At4g14180 hypothetical protein
Length = 1268
Score = 489 bits (1259), Expect = e-138
Identities = 268/533 (50%), Positives = 356/533 (66%), Gaps = 35/533 (6%)
Query: 18 CSQGHRSSLNLQTNQGGTICLVCFSNLISNPSSPTIHVSYALSQLSRSLSIPQFLHSIVT 77
C+ GHRS+++L+ +QGGT CL+CFSNL+S+P PT+HVSYAL QLS ++S P FL ++++
Sbjct: 31 CANGHRSTISLRDDQGGTFCLICFSNLVSDPRIPTVHVSYALHQLSIAISEPIFLRTLLS 90
Query: 78 FHPHFLVSPLVAALSSFDDEPIAEQLVHLILNLSASSDPSVCREFVARVSDRVSSGALGW 137
H HFLVSPLV ALSS DD PIA Q++ +I L + + S+ +FV R+SD++SSGALGW
Sbjct: 91 SHIHFLVSPLVHALSSIDDAPIAIQIMDMISLLCSVEESSIGEDFVERISDQLSSGALGW 150
Query: 138 SSRQLHSLHCLGVLLNCEKEDDLHVHIKDVYSLISVLVTGLQLPSEEIRGEVLFVLYKLF 197
S RQLH LHC GVL++CE +++ HI+D +L+ LV GLQLPSEEIRGE+LF LYK
Sbjct: 151 SRRQLHMLHCFGVLMSCE-NININSHIRDKEALVCQLVEGLQLPSEEIRGEILFALYKFS 209
Query: 198 ALHSTSDEGDGSDMLIPFCPKLLYLLGDVLLKTQNDDVRLNCIALLTMLAQRQLLREEPA 257
AL T DG ++L CPKLL L + L KTQ DDVRLNC+ALLT+LAQ+ LL +
Sbjct: 210 ALQFTEQNVDGIEVLSLLCPKLLCLSLEALAKTQRDDVRLNCVALLTILAQQGLLANSHS 269
Query: 258 YDTGNMSLSGEVN---FKETEDVPEGASLVNLFAEAIKGPLLSSVSQVQIGALDLLFHYL 314
+MSL EV+ + E+V L LFAEAIKGPLLS+ S+VQI LDL+FHY+
Sbjct: 270 NSASSMSLD-EVDDDPMQTAENVAARPCLNVLFAEAIKGPLLSTDSEVQIKTLDLIFHYI 328
Query: 315 SSVGTSGNQIQVLVEENIADYLFEILRLS-------------------------VPFHPV 349
S T QIQV+VEEN+ADY+FEILRLS VP HP
Sbjct: 329 SQESTPSKQIQVMVEENVADYIFEILRLSAEHSFRKRLVIGFPSVIRVLHYVGEVPCHPF 388
Query: 350 QYETLKLIYECISECPGAVSTSQMEELVLVLIKMLTKHSDGEMGMIPETFIMACSVFVAL 409
Q +TLKLI CIS+ PG S+SQ++E+ LVL KML ++ EMG+ P+ F + CSVFV+L
Sbjct: 389 QIQTLKLISSCISDFPGIASSSQVQEIALVLKKMLERYYSQEMGLFPDAFAIICSVFVSL 448
Query: 410 MRSPSCNGALDLSKSIEEAVKQAILACLYVSERNINQILQCFYLLKEAYAYSHDENSTHI 469
M++PS D+ S++E+++ +ILA L + E++ QIL YLL E Y Y ST I
Sbjct: 449 MKTPSFGETADVLTSLQESLRHSILASLSLPEKDSTQILHAVYLLNEVYVYC--TASTSI 506
Query: 470 SK---LELRSGILDICRTHLLPWLATGIMNGEEELFLGLLEIFHSILLLHCSI 519
+K +ELR ++D+C +HLLPW + + EE LG++E FHSILL + I
Sbjct: 507 NKTICIELRHCVIDVCTSHLLPWFLSDVNEVNEEATLGIMETFHSILLQNSDI 559
>At5g49310 importin alpha
Length = 519
Score = 36.2 bits (82), Expect = 0.046
Identities = 71/290 (24%), Positives = 115/290 (39%), Gaps = 60/290 (20%)
Query: 84 VSPLVAALSSFDDEPIAEQLVHLILNLSASSDPSVCREFVARVSDRVSSGALGWSSRQLH 143
V PL L + D+ + EQ + + N++ D CR+FV ++SGA QL+
Sbjct: 157 VVPLFVQLLASPDDDVREQAIWGLGNVAG--DSIQCRDFV------LNSGAFIPLLHQLN 208
Query: 144 SLHCLGVLLNCE---------KEDDLHVHIKDVYSLISVLVTGLQLPSEEIRGEVLFVLY 194
+ L +L N K +K V ++ LV E++ ++ +
Sbjct: 209 NHATLSILRNATWTLSNFFRGKPSPPFDLVKHVLPVLKRLVYS---DDEQV---LIDACW 262
Query: 195 KLFALHSTSDEGDGSDMLIPFCPKLLYLL---------------GDVLL-KTQNDDVRLN 238
L L S+E S + P+L+ LL G+++ +Q +N
Sbjct: 263 ALSNLSDASNENIQSVIEAGVVPRLVELLQHASPVVLVPALRCIGNIVSGNSQQTHCVIN 322
Query: 239 C---IALLTMLAQRQL--LREEPAYDTGNMSLSGEVNFKETEDVPEGASLVNLFAEA--- 290
C L +L Q + +R E + N++ E + D SLVNL A
Sbjct: 323 CGVLPVLADLLTQNHMRGIRREACWTISNITAGLEEQIQSVIDANLIPSLVNLAQHAEFD 382
Query: 291 IKGPLLSSVSQVQIGALDLLFHYLSSVGTSGNQIQVLVEENIADYLFEIL 340
IK + ++S +SVG S NQI+ LVE+N L +IL
Sbjct: 383 IKKEAIWAISN-------------ASVGGSPNQIKYLVEQNCIKALCDIL 419
>At1g48040 protein phosphatase-2C, putative
Length = 377
Score = 33.9 bits (76), Expect = 0.23
Identities = 13/40 (32%), Positives = 24/40 (59%)
Query: 12 GDSRWCCSQGHRSSLNLQTNQGGTICLVCFSNLISNPSSP 51
GD R C + + + LQ++ T+ ++CFS++ S+P P
Sbjct: 314 GDPRQCAMELGKEAARLQSSDNMTVIVICFSSVPSSPKQP 353
>At5g04130 DNA gyrase subunit B - like protein
Length = 732
Score = 32.7 bits (73), Expect = 0.51
Identities = 30/138 (21%), Positives = 58/138 (41%), Gaps = 15/138 (10%)
Query: 67 SIPQFLHSIVTFHPHFLVSPLVAALSSFDDEPIAEQLVHLILNLS--------------A 112
S+ ++L + HP L S + +L+++ A++ L+ + S +
Sbjct: 447 SVQEYLTEFLELHPDILESIISKSLNAYKAALAAKRARELVRSKSVLKSSSLPGKLADCS 506
Query: 113 SSDPSVCREFVARVSDRVSSGALGWSSRQLHSLHCLGVLLNCEKEDDLHVHIKDVYSLIS 172
S+DP V F+ S G R L G +LN E++D+ ++ + +
Sbjct: 507 STDPEVSEIFIVEGDSAGGSAKQGRDRRFQAILPLRGKILNIERKDEAAMYKNEEIQNL- 565
Query: 173 VLVTGLQLPSEEIRGEVL 190
+L GL + E+ + E L
Sbjct: 566 ILGLGLGVKGEDFKKENL 583
>At5g47490 putative protein
Length = 1361
Score = 30.8 bits (68), Expect = 1.9
Identities = 25/106 (23%), Positives = 49/106 (45%), Gaps = 3/106 (2%)
Query: 49 SSPTIHVSYALSQLSRSLSIPQFLHSIVTFHPH-FLVSPLVAALSSFDDEPIAEQLVHLI 107
+SP L + S++L Q+ +++TF P+ + + ++A + Q V
Sbjct: 872 ASPEAIQRTELYEYSKTLGNSQY--TLLTFQPYKVMYAHMLAEVGKLSTAQKYCQAVLKC 929
Query: 108 LNLSASSDPSVCREFVARVSDRVSSGALGWSSRQLHSLHCLGVLLN 153
L S + + ++FV+ + +R+ G + LH +GVLLN
Sbjct: 930 LKTGRSPEVEMWKQFVSSLEERIRIHQQGGYTANLHPEKLVGVLLN 975
>At5g24000 putative protein
Length = 443
Score = 30.8 bits (68), Expect = 1.9
Identities = 21/48 (43%), Positives = 27/48 (55%), Gaps = 3/48 (6%)
Query: 41 FSNLISNPSSPTI---HVSYALSQLSRSLSIPQFLHSIVTFHPHFLVS 85
F+ + NP SPTI HVS + LS S++I + VT PH LVS
Sbjct: 5 FTVVSPNPLSPTISSLHVSPSRHHLSSSITIRRNKSPPVTLTPHRLVS 52
>At2g28300 unknown protein
Length = 2218
Score = 30.8 bits (68), Expect = 1.9
Identities = 25/91 (27%), Positives = 43/91 (46%), Gaps = 5/91 (5%)
Query: 88 VAALSSFDDEPIAEQLVHLILNLSASSDPSVCREFV-ARVSDRVSSGALGWSSRQLHSLH 146
+A L ++E QL N+ SS PSV + + A+ D+V + S S
Sbjct: 1823 LATLPLTEEENADSQLA----NIEPSSSPSVVEKNIEAQDQDQVKTAGCELVSTGCSSEP 1878
Query: 147 CLGVLLNCEKEDDLHVHIKDVYSLISVLVTG 177
+ + + E + D+HVH+K+ S++V G
Sbjct: 1879 QVHLPPSAEPDGDIHVHLKETEKSESMVVVG 1909
>At3g17250 protein phosphatase like
Length = 384
Score = 30.4 bits (67), Expect = 2.5
Identities = 15/54 (27%), Positives = 29/54 (52%), Gaps = 3/54 (5%)
Query: 12 GDSRWCCSQGHRSSLNLQTNQGGTICLVCFSNLISNPSSPTIHVSYALSQLSRS 65
GD R C + R +L L ++ T+ ++CFS S+P+ + + +S +R+
Sbjct: 326 GDPRRCAMELGREALRLDSSDNVTVVVICFS---SSPAPQRRRIRFCVSDEARA 376
>At1g58460 unknown protein
Length = 134
Score = 30.4 bits (67), Expect = 2.5
Identities = 13/28 (46%), Positives = 16/28 (56%)
Query: 1 MDSESIDIEDEGDSRWCCSQGHRSSLNL 28
MD +D D GDS W GH SS++L
Sbjct: 1 MDFSDLDYSDAGDSGWTMYLGHSSSVSL 28
>At2g15630 putative salt-inducible protein
Length = 627
Score = 29.6 bits (65), Expect = 4.3
Identities = 12/31 (38%), Positives = 18/31 (57%)
Query: 442 RNINQILQCFYLLKEAYAYSHDENSTHISKL 472
R +++ ++CFYL+KE Y E HI L
Sbjct: 169 RMVDEAIECFYLMKEKGFYPKTETCNHILTL 199
>At4g10560 putative protein
Length = 703
Score = 29.3 bits (64), Expect = 5.6
Identities = 37/167 (22%), Positives = 74/167 (44%), Gaps = 25/167 (14%)
Query: 59 LSQLSRSLSIPQFLHSIVTFHPHFLVSPLVAALSSFDDEPIAEQLVHLILNLSASSDPSV 118
+S +S+ +S+P+ L S H + L+A Q++ L+ ++ S P
Sbjct: 8 ISLISQLISLPKILDSKWNSVTHSEIISLIA------------QIISLVSSMDLHSQPKP 55
Query: 119 CREFVARVSDRVS---SGALGWSSRQLHSL-HCLGVLLNCEKEDDLHVHIK-DVYSLISV 173
EF++ V+ +S S L L L L +++ + + D + ++ D+ +
Sbjct: 56 ESEFMSLVTQAISLFDSMDLNPELSPLSELISLLSQIISTDSDSDSNRKVEWDLDRPLCE 115
Query: 174 LVTGLQLPSEEIRGEVLFVLYKLFALHSTSDEGDGSDMLIPFCPKLL 220
+ G P E++ ++Y++F+L S S+ I CP+LL
Sbjct: 116 TLFGQPEP------ELVSLIYQIFSL--VSSINSKSEKFISLCPQLL 154
>At3g26500 unknown protein
Length = 471
Score = 29.3 bits (64), Expect = 5.6
Identities = 92/445 (20%), Positives = 173/445 (38%), Gaps = 73/445 (16%)
Query: 55 VSYALSQLSRSLSIPQFLHSIVTFHPHF--LVSPLVAALSSFDDEPIAEQLVHLILNLSA 112
+SY L Q +L P + + T P F L +P + ++ + Q + + +L +
Sbjct: 11 LSYVLHQHDSNLHAPPSMAAQETLLPSFPLLSNPEIMSMLTQSIPTTITQTLFVFNSLGS 70
Query: 113 SSDP---SVCREFVARVSDRVSSGALGWSSRQLHSLHCLGVLLNCEKEDDLHVHIKDVYS 169
DP S R +A++ D +S S ++++ GV+ E D +KD
Sbjct: 71 RPDPLAVSSARFKIAQIMDSLSPEEAAKES-EIYA----GVVRLDEVHDSYEKKLKDTEE 125
Query: 170 LISVL----VTGLQLPSEEIRGEVLFVLYKLFALHSTSDEGDGSDMLIPFCPKLLY-LLG 224
+S + V + EE+ +VL VL K T + D S + P+ + ++G
Sbjct: 126 ELSRVYSTEVESMLRSGEEVNEKVLAVL-KEAESGGTVERIDLSSQELKLIPEAFWKVVG 184
Query: 225 DVLLKTQNDDVRLNCIALLTMLAQRQLLREEPAYDTGNMSLSGEVNFKETEDVPEGAS-L 283
V L +D+ A+ + +L +V+ E +P+ L
Sbjct: 185 LVYLNLSGNDLTFIPDAISKLKKLEEL----------------DVSSNSLESLPDSIGML 228
Query: 284 VNLFAEAIKGPLLSSVSQV-----QIGALDLLFHYLSSVGTSGNQIQVLVEENIADYLFE 338
+NL + L+++ + + LD ++ L+S+ T NI L
Sbjct: 229 LNLRILNVNANNLTALPESIAHCRSLVELDASYNNLTSLPT-----------NIGYGLQN 277
Query: 339 ILRLSVPFHPVQYETLKLIYECISECPGAVSTSQMEELVLVLIKMLTKHSDGEMGMIPET 398
+ RLS+ + ++Y PG++S + +K L H + E+ IP +
Sbjct: 278 LERLSIQLNKLRY------------FPGSISE-------MYNLKYLDAHMN-EIHGIPNS 317
Query: 399 FIMACSVFVALMRSPSCNGALDLSKSIEEAVKQAILACLYVSERNINQILQCFYLLKEAY 458
I + L S + N + + +I + L L +S I I FY L++
Sbjct: 318 -IGRLTKLEVLNLSSNFNNLMGVPDTITDLTN---LRELDLSNNQIQAIPDSFYRLRKLE 373
Query: 459 AYSHDENSTHISKLELRSGILDICR 483
+ D+N I E+ + ++ R
Sbjct: 374 KLNLDQNPLEIPSQEVATQGAEVVR 398
>At3g20260 unknown protein
Length = 437
Score = 29.3 bits (64), Expect = 5.6
Identities = 18/70 (25%), Positives = 33/70 (46%), Gaps = 1/70 (1%)
Query: 350 QYETLKLIYECISECPGAVS-TSQMEELVLVLIKMLTKHSDGEMGMIPETFIMACSVFVA 408
QY L + C E P + T+Q+ + LVL++ ++ E G E + A +
Sbjct: 217 QYTQLSHLISCQPETPTCYNHTAQLFQQFLVLLQRYIENEPFEQGSRSELYARARNAMPK 276
Query: 409 LMRSPSCNGA 418
L+++P G+
Sbjct: 277 LLQAPKIQGS 286
>At3g10270 putative DNA gyrase subunit B
Length = 657
Score = 29.3 bits (64), Expect = 5.6
Identities = 29/138 (21%), Positives = 57/138 (41%), Gaps = 15/138 (10%)
Query: 67 SIPQFLHSIVTFHPHFLVSPLVAALSSFDDEPIAEQLVHLILNLS--------------A 112
S+ ++L + HP L S + +L+++ A++ L+ + S +
Sbjct: 372 SVQEYLTEYLELHPDVLESIISKSLNAYKAALAAKRARELVRSKSVLKSSSLPGKLADCS 431
Query: 113 SSDPSVCREFVARVSDRVSSGALGWSSRQLHSLHCLGVLLNCEKEDDLHVHIKDVYSLIS 172
S+DP+ F+ S G R L G +LN E++D+ ++ + +
Sbjct: 432 STDPAESEIFIVEGDSAGGSAKQGRDRRFQAILPLRGKILNIERKDEAAMYKNEEIQNL- 490
Query: 173 VLVTGLQLPSEEIRGEVL 190
+L GL + E+ E L
Sbjct: 491 ILGLGLGVKGEDFNKENL 508
>At2g20580 putative 26S proteasome regulatory subunit S2
Length = 891
Score = 28.9 bits (63), Expect = 7.3
Identities = 32/152 (21%), Positives = 62/152 (40%), Gaps = 21/152 (13%)
Query: 300 SQVQIGALDLLFHYLSSVGTSGNQIQVLVEENIADYLFEILRLSVPFHPVQYETLKLIYE 359
S V+IGA+ L +S G+ +QI+ + + D P + + +L L
Sbjct: 480 SSVRIGAIMGLG--ISYAGSQNDQIRNKLSPILND-------AKAPLDVIAFASLSLGMI 530
Query: 360 CISECPGAVSTSQMEELVLVLIKMLTKHSDGEMGMIPETFIMACSVFVALMRSPSCNGAL 419
+ C EE+ +I L S+ E+G F+ + L + S
Sbjct: 531 YVGSCN--------EEVAQSIIFALMDRSEAELGDALTRFLPLGLGLLYLGKQESVEATA 582
Query: 420 DLSKSIEEAVKQ----AILACLYVSERNINQI 447
++SK+ E +++ +L+C Y N+ ++
Sbjct: 583 EVSKTFNEKIRKYCDMTLLSCAYAGTGNVLKV 614
>At1g37180 hypothetical protein
Length = 661
Score = 28.9 bits (63), Expect = 7.3
Identities = 13/34 (38%), Positives = 21/34 (61%)
Query: 275 EDVPEGASLVNLFAEAIKGPLLSSVSQVQIGALD 308
E E A L LF+E + GP L+ +Q++ G++D
Sbjct: 342 EPYEEDAGLCKLFSENLSGPALTWFTQLEEGSID 375
>At5g06820 SRF2 (LRR receptor protein kinase like)
Length = 735
Score = 28.5 bits (62), Expect = 9.6
Identities = 34/124 (27%), Positives = 51/124 (40%), Gaps = 26/124 (20%)
Query: 234 DVRLNCIALLTMLA-QRQLLREEPAYDTGNMSLSGEVNF--------------KETEDVP 278
D++L + LL L Q Q L D +L GE+ F T+ +P
Sbjct: 75 DLQLRELKLLGSLGNQLQHLHNLKILDVSFNNLEGEIPFGLPPNATHINMAYNNLTQSIP 134
Query: 279 EGASLV------NLFAEAIKGPLLSSVSQVQIGALDLLFHYL-----SSVGTSGNQIQVL 327
L+ NL ++ GPL + S +QI +DL F+ L SS GT N +
Sbjct: 135 FSLPLMTSLQSLNLSHNSLSGPLGNVFSGLQIKEMDLSFNNLTGDLPSSFGTLMNLTSLY 194
Query: 328 VEEN 331
++ N
Sbjct: 195 LQNN 198
>At4g13650 unknown protein
Length = 1024
Score = 28.5 bits (62), Expect = 9.6
Identities = 15/43 (34%), Positives = 22/43 (50%)
Query: 82 FLVSPLVAALSSFDDEPIAEQLVHLILNLSASSDPSVCREFVA 124
+ S +++A + I EQL L+L L SSD VC V+
Sbjct: 249 YAFSSVLSACKKIESLEIGEQLHGLVLKLGFSSDTYVCNALVS 291
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.321 0.137 0.397
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,664,337
Number of Sequences: 26719
Number of extensions: 491267
Number of successful extensions: 1219
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 12
Number of HSP's that attempted gapping in prelim test: 1207
Number of HSP's gapped (non-prelim): 22
length of query: 526
length of database: 11,318,596
effective HSP length: 104
effective length of query: 422
effective length of database: 8,539,820
effective search space: 3603804040
effective search space used: 3603804040
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 62 (28.5 bits)
Medicago: description of AC147484.3


