

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147471.10 + phase: 0
(532 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
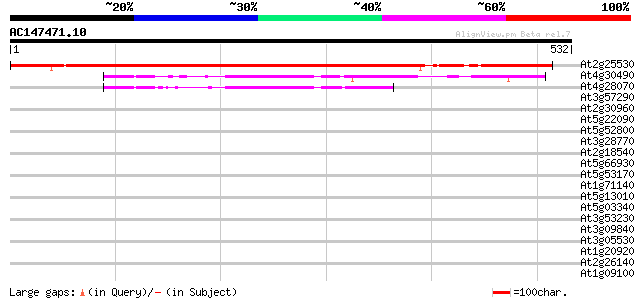
Score E
Sequences producing significant alignments: (bits) Value
At2g25530 hypothetical protein 612 e-175
At4g30490 unknown protein 132 4e-31
At4g28070 unknown protein 119 5e-27
At3g57290 p48 protein 33 0.39
At2g30960 hypothetical protein 33 0.39
At5g22090 unknown protein 31 1.5
At5g52800 unknown protein 30 2.6
At3g28770 hypothetical protein 30 2.6
At2g18540 putative vicilin storage protein (globulin-like) 30 2.6
At5g66930 unknown protein 30 3.3
At5g53170 cell division protein FtsH protease-like 30 3.3
At1g71140 hypothetical protein 30 4.4
At5g13010 pre-mRNA splicing factor ATP-dependent RNA helicase -l... 29 7.4
At5g03340 transitional endoplasmic reticulum ATPase 29 7.4
At3g53230 CDC48 - like protein 29 7.4
At3g09840 putative transitional endoplasmic reticulum ATPase 29 7.4
At3g05530 26S proteasome AAA-ATPase subunit RPT5a 29 7.4
At1g20920 putative RNA helicase 29 7.4
At2g26140 FtsH like protease 28 9.7
At1g09100 putative 26S protease regulatory subunit 6A 28 9.7
>At2g25530 hypothetical protein
Length = 655
Score = 612 bits (1577), Expect = e-175
Identities = 305/518 (58%), Positives = 385/518 (73%), Gaps = 19/518 (3%)
Query: 1 MEEYHVNLSNWEKKRENERRRILMDEVEKQQNDKDWWK--RLNNKITERWTNSRKRPENV 58
ME+YHV L+ WEKKRE ERR+++++E EK+++D W + K+ RW R R NV
Sbjct: 152 MEDYHVRLAEWEKKREEERRKLMVEEAEKKEDDGMWASVNKQGQKLLGRWVLGR-RQMNV 210
Query: 59 DPGVGKWVSYLKREKKLDSLVGRRPTAPPAPKGLYIYGNVGSGKTMLMDMFYSATEGIVK 118
+PGVGKWVSYL RE+KLDS+VG RP PPAPKGLYIYGNVG GKTMLMDMFY AT+GI++
Sbjct: 211 EPGVGKWVSYLNRERKLDSIVGSRPAVPPAPKGLYIYGNVGCGKTMLMDMFYGATDGIIR 270
Query: 119 HRRRYHFHEAMLRINEHMHKTWKKQMEEKPLQSGISSWIMNLPFDTKAKEWLAAEERYKK 178
HR+R+HFHEAML+INE MHK WK+ EKP+Q ISSWIMNLP D K KEWLA EE YK+
Sbjct: 271 HRQRFHFHEAMLKINEQMHKYWKENGAEKPMQYSISSWIMNLPVDEKVKEWLAGEEFYKQ 330
Query: 179 EVQMKHILPDVADKFFLDREGEEKGANILCFDEIQTVDVFAIVALSGILSRLLSSGTIIV 238
++QMKHILP VADKF +D++ +KGA+ILCFDEIQTVDVFAIVALSGI+SRLL++GT++V
Sbjct: 331 QLQMKHILPAVADKFLVDQQSSKKGASILCFDEIQTVDVFAIVALSGIMSRLLTTGTVLV 390
Query: 239 ATSNRAPKDLNEANMVPEFFQNLLSNLEEHCEKVLVGSEIDYRRFIAQRSENRVNYLWPI 298
ATSNRAP++LN+ M E F +S LE+HCE + +GSE+DYRR A+ S V+YLWP+
Sbjct: 391 ATSNRAPRELNQDGMQKEIFDKFISKLEKHCEIISIGSEVDYRRVAAKNSAENVHYLWPL 450
Query: 299 ERETINKFEKKWQDATGRFGGKVISNTISVMFGRTLEVPESCEGVARFTFDYLCGRPLGA 358
+ +FEK W+ T ++GG++ S T+ VMFGRT+EVPESC GVARFTF+YLCGRP+GA
Sbjct: 451 NDAVLEEFEKMWRQVTDQYGGEITSATLPVMFGRTVEVPESCSGVARFTFEYLCGRPVGA 510
Query: 359 ADYIAVAENYHTVFISDIPMMSMRIRDKTS--LSYFEQQKKKRSLLFRHGDLSLLLTNYT 416
ADYIAVA+NYHT+FISDIP MSM IRDK ++ ++ L+ H L+ + T
Sbjct: 511 ADYIAVAKNYHTIFISDIPAMSMEIRDKARRFITLVDE-------LYNH-HCCLVSSAET 562
Query: 417 IIILVYVAWHPLQLMNYFKELRRAHFLTWREAEGSKLRRDVLAEGNVGSGGTPVGITSIL 476
I ++ L +L F T E E S+LRRDVLAEG++ + G+P I S+L
Sbjct: 563 AIDELFQGTAEGTLF----DLESFQFET--ETEDSRLRRDVLAEGSISAAGSPSSIVSML 616
Query: 477 SGQEELFTFQRAVSRLIEMQTQLYLDGVSNVHPYFQTQ 514
SG+EE+F F RA SRLIEMQT LYL+GV +HPYF Q
Sbjct: 617 SGEEEMFAFARAASRLIEMQTPLYLEGVRFLHPYFHQQ 654
>At4g30490 unknown protein
Length = 497
Score = 132 bits (333), Expect = 4e-31
Identities = 113/428 (26%), Positives = 175/428 (40%), Gaps = 103/428 (24%)
Query: 90 KGLYIYGNVGSGKTMLMDMFYSATEGIVKHRRRYHFHEAMLRINEHMHKTWKKQMEEKPL 149
KGLY+YG VG+GKTMLMD+F+ K ++R HFH+ ML ++ + K
Sbjct: 162 KGLYLYGGVGTGKTMLMDLFFDQLPCTWK-KQRIHFHDFMLSVHSRLQK----------- 209
Query: 150 QSGISSWIMNLPFDTKAKEWLAAEERYKKEVQMKHILPDVADKFFLDREGEEKGANILCF 209
G+S P + A+E I D A +LC
Sbjct: 210 HKGLSD-----PLEVVAQE----------------IAHD---------------AILLCL 233
Query: 210 DEIQTVDVFAIVALSGILSRLLSSGTIIVATSNRAPKDLNEANMVPEFFQNLLSNLEEHC 269
DE DV + L+ + L S+G I+VATSNR P L E + + F +S+L+E
Sbjct: 234 DEFMVTDVADALILNRLFGHLFSNGVILVATSNRNPDKLYEGGLQRDLFLPFISSLKERS 293
Query: 270 EKVLVGSEIDYRRFIAQRSENRVNYLWPIERETINKFEKKWQDATGRFGGKVIS--NTIS 327
+GS +DYR+ + + I ++ ++K++ G V++ +
Sbjct: 294 VVHEIGSAVDYRKLTSAEQG-----FYFIGKDLSTLLKQKFRQL---IGDNVVARPQVVE 345
Query: 328 VMFGRTLEVPESCEGVARFTFDYLCGRPLGAADYIAVAENYHTVFISDIPMMSMRIRDKT 387
V+ GR L++P G A F F+ LC RPLGAADY + + +HT+ + +IP+ + R
Sbjct: 346 VVMGRKLQIPLGANGCAYFPFEELCDRPLGAADYFGLFKKFHTLALDEIPVFGLHNRTAA 405
Query: 388 SLSYFEQQKKKRSLLFRHGDLSLLLTNYTIIILVYVAWHPLQLMNYFKELRRAHFLTWRE 447
Y + LV V + RA L E
Sbjct: 406 ---------------------------YRFVTLVDVMYE-----------NRARLLCTAE 427
Query: 448 AEGSKLRRDVLAEGNVGSGGTPVG-------ITSILSGQEELFTFQRAVSRLIEMQTQLY 500
A +L ++ S G +T + E F R +SRL EM ++ Y
Sbjct: 428 ANPQELLEKIITISEAKSMGPRTSSRSRKNDVTELCVDNELGFAKDRTISRLTEMNSKEY 487
Query: 501 LDGVSNVH 508
L+ + H
Sbjct: 488 LEHHAITH 495
>At4g28070 unknown protein
Length = 464
Score = 119 bits (297), Expect = 5e-27
Identities = 85/275 (30%), Positives = 132/275 (47%), Gaps = 55/275 (20%)
Query: 90 KGLYIYGNVGSGKTMLMDMFYSATEGIVKHRRRYHFHEAMLRINEHMHKTWKKQMEEKPL 149
KGLY+YG VG+GKTMLMD+F+ + +R HFH ML ++ + K K +E+ PL
Sbjct: 134 KGLYLYGGVGTGKTMLMDLFFHQLPASWR-TQRIHFHNFMLSVHSRLQK--HKGLED-PL 189
Query: 150 QSGISSWIMNLPFDTKAKEWLAAEERYKKEVQMKHILPDVADKFFLDREGEEKGANILCF 209
+ ++ L ++AD+ A +LC
Sbjct: 190 E------VVGL---------------------------EIADE-----------AILLCL 205
Query: 210 DEIQTVDVFAIVALSGILSRLLSSGTIIVATSNRAPKDLNEANMVPEFFQNLLSNLEEHC 269
DE DV + L+ + L ++G I+VATSNRAP +L E + + F +S L+E C
Sbjct: 206 DEFMVNDVADALILNRLFRHLFNNGIILVATSNRAPDNLYEGGLQRDLFLPFISTLKERC 265
Query: 270 EKVLVGSEIDYRRFIAQRSENRVNYLWPIERETINKFEKKWQDATGRFGGKVISNTISVM 329
+GS +DYR+ + + I ++ ++K+Q G + V+
Sbjct: 266 VVREIGSSVDYRKLTSAEEG-----FYFIGKDISGLLKQKFQLLVG--DQPAGPQVVEVV 318
Query: 330 FGRTLEVPESCEGVARFTFDYLCGRPLGAADYIAV 364
GR L+VP + +G A F F+ LC RPLGAADY+ +
Sbjct: 319 MGRKLQVPLAADGCAYFLFEELCDRPLGAADYLGL 353
>At3g57290 p48 protein
Length = 441
Score = 33.1 bits (74), Expect = 0.39
Identities = 27/102 (26%), Positives = 44/102 (42%), Gaps = 24/102 (23%)
Query: 91 GLYIYGNVGSGKTMLMDMF------------------YSATEGIVKHRRRYHFHEAMLRI 132
GLYI+ N +G+T ++D+F Y AT IV RRR E +++
Sbjct: 220 GLYIFFNHDNGRTQIIDLFNQDKYLNAIQTSAPHLLRYLATAFIVNKRRRPQLKE-FIKV 278
Query: 133 NEHMHKTWKKQMEEKPLQSGISSWIMNLPFDTKAKEWLAAEE 174
+ H ++K P+ ++ +N FD K+ EE
Sbjct: 279 IQQEHYSYK-----DPIIEFLACVFVNYDFDGAQKKMKECEE 315
>At2g30960 hypothetical protein
Length = 260
Score = 33.1 bits (74), Expect = 0.39
Identities = 22/74 (29%), Positives = 36/74 (47%), Gaps = 2/74 (2%)
Query: 1 MEEYHVNLSNWEKKRENERRRILMDEVEKQQNDKDWWKRLNNKITERWTNSR--KRPENV 58
++E +N ++K E ERRR L E EK+ ++D K K R ++ E++
Sbjct: 29 LQERWLNEKEKKRKEEEERRRKLQAEEEKKIEEEDLKKAEEEKRMNRSNRKHFGRKKESI 88
Query: 59 DPGVGKWVSYLKRE 72
D G ++V K E
Sbjct: 89 DGGEARFVEKEKPE 102
>At5g22090 unknown protein
Length = 463
Score = 31.2 bits (69), Expect = 1.5
Identities = 21/78 (26%), Positives = 36/78 (45%), Gaps = 12/78 (15%)
Query: 14 KRENERRRILMDEVEKQQNDK---DWWKRLNNKITERWTNSRKRPENVDPGVGK-WVSYL 69
KR + ++ DE E ++ + D W ++ N +K+ E ++PG W S L
Sbjct: 62 KRISSSEKLRPDEEEAEEESRSGVDIWAQIQQD-----KNDKKKEEEIEPGQSDVWSSIL 116
Query: 70 KREKKLDSLVGRRPTAPP 87
+KK +S + T PP
Sbjct: 117 SEKKKTES---SKDTVPP 131
>At5g52800 unknown protein
Length = 618
Score = 30.4 bits (67), Expect = 2.6
Identities = 27/98 (27%), Positives = 45/98 (45%), Gaps = 9/98 (9%)
Query: 227 LSRLLSSGTIIVATSNRAPKDLNEANMVPEFFQNLLSNLEEHCEKVLV-GSEIDYRRFIA 285
LS ++++ T KD+ E ++ F +L+ N+E CEK+LV E D + +
Sbjct: 326 LSSKAGKTSVLLPTGRFKCKDMGERDV---FMTSLICNVESDCEKLLVCKMESDCMKTLC 382
Query: 286 QRSENRVNYLWPIERETINKFEKKWQD---ATGRFGGK 320
+E N L + + KF+ +T FGGK
Sbjct: 383 FDTEVNSNNL--VRDQNAQKFQLNASTSDMSTSYFGGK 418
>At3g28770 hypothetical protein
Length = 2081
Score = 30.4 bits (67), Expect = 2.6
Identities = 14/48 (29%), Positives = 32/48 (66%), Gaps = 1/48 (2%)
Query: 13 KKRENERRRILMDEVEKQQ-NDKDWWKRLNNKITERWTNSRKRPENVD 59
KK+E +++ + +E++KQ+ N K+ K N+K+ E +++++ E+ D
Sbjct: 957 KKKEEDKKEYVNNELKKQEDNKKETTKSENSKLKEENKDNKEKKESED 1004
>At2g18540 putative vicilin storage protein (globulin-like)
Length = 699
Score = 30.4 bits (67), Expect = 2.6
Identities = 19/57 (33%), Positives = 29/57 (50%), Gaps = 1/57 (1%)
Query: 2 EEYHVNLSNWEKKRENERRRILMDEVEKQQNDKDWWKRLNNKITERWTNSRKRPENV 58
EE KKRE ER+R +EVE+++ ++ KR + +R RKR E +
Sbjct: 512 EEEREKEEEMAKKREEERQRKEREEVERKRREEQERKRREEEARKR-EEERKREEEM 567
>At5g66930 unknown protein
Length = 215
Score = 30.0 bits (66), Expect = 3.3
Identities = 13/55 (23%), Positives = 25/55 (44%)
Query: 10 NWEKKRENERRRILMDEVEKQQNDKDWWKRLNNKITERWTNSRKRPENVDPGVGK 64
NW +K N++ +I + E + W+ ++ E+W + + P VGK
Sbjct: 73 NWIEKHPNKKSQICLSFYEVKSKQPSWFTKIERLYWEQWYINLNVLQPTKPPVGK 127
>At5g53170 cell division protein FtsH protease-like
Length = 806
Score = 30.0 bits (66), Expect = 3.3
Identities = 24/72 (33%), Positives = 35/72 (48%), Gaps = 10/72 (13%)
Query: 38 KRLNNKIT-ERWTNSRKRPENVDPG---VGKWVSYLKREKKLDSLVGRRPTAPPAPKGLY 93
K LN +IT E+ + K + D + + V YLK K L G+ PKG+
Sbjct: 346 KELNKEITPEKNVKTFKDVKGCDDAKQELEEVVEYLKNPSKFTRLGGK------LPKGIL 399
Query: 94 IYGNVGSGKTML 105
+ G G+GKT+L
Sbjct: 400 LTGAPGTGKTLL 411
>At1g71140 hypothetical protein
Length = 485
Score = 29.6 bits (65), Expect = 4.4
Identities = 10/30 (33%), Positives = 22/30 (73%)
Query: 6 VNLSNWEKKRENERRRILMDEVEKQQNDKD 35
V L+NW+K+ R R++ DE E+++++++
Sbjct: 451 VILTNWKKQARKARERVMGDEYEEKESEEE 480
>At5g13010 pre-mRNA splicing factor ATP-dependent RNA helicase -like
protein
Length = 1226
Score = 28.9 bits (63), Expect = 7.4
Identities = 16/61 (26%), Positives = 35/61 (57%), Gaps = 1/61 (1%)
Query: 13 KKRENERRRILMDEVEKQQNDKDWWKRLNNKITERWTNSRKRPENVDPGVGKWVSYLKRE 72
KK++ E + + +E+EK + D+ L +K ER ++++ + PG+ K ++L+ +
Sbjct: 1164 KKKQKEEKSGMEEEMEKLRRDQVE-SELRSKERERKKRAKQQQQISGPGLKKGTTFLRPK 1222
Query: 73 K 73
K
Sbjct: 1223 K 1223
>At5g03340 transitional endoplasmic reticulum ATPase
Length = 810
Score = 28.9 bits (63), Expect = 7.4
Identities = 14/35 (40%), Positives = 21/35 (60%), Gaps = 5/35 (14%)
Query: 71 REKKLDSLVGRRPTAPPAPKGLYIYGNVGSGKTML 105
R +L +G +P PKG+ +YG GSGKT++
Sbjct: 228 RHPQLFKSIGVKP-----PKGILLYGPPGSGKTLI 257
>At3g53230 CDC48 - like protein
Length = 815
Score = 28.9 bits (63), Expect = 7.4
Identities = 14/35 (40%), Positives = 21/35 (60%), Gaps = 5/35 (14%)
Query: 71 REKKLDSLVGRRPTAPPAPKGLYIYGNVGSGKTML 105
R +L +G +P PKG+ +YG GSGKT++
Sbjct: 229 RHPQLFKSIGVKP-----PKGILLYGPPGSGKTLI 258
>At3g09840 putative transitional endoplasmic reticulum ATPase
Length = 809
Score = 28.9 bits (63), Expect = 7.4
Identities = 14/35 (40%), Positives = 21/35 (60%), Gaps = 5/35 (14%)
Query: 71 REKKLDSLVGRRPTAPPAPKGLYIYGNVGSGKTML 105
R +L +G +P PKG+ +YG GSGKT++
Sbjct: 228 RHPQLFKSIGVKP-----PKGILLYGPPGSGKTLI 257
>At3g05530 26S proteasome AAA-ATPase subunit RPT5a
Length = 424
Score = 28.9 bits (63), Expect = 7.4
Identities = 20/89 (22%), Positives = 44/89 (48%), Gaps = 6/89 (6%)
Query: 17 NERRRILMDEVEKQQNDKDWWKRLNNKITERWTNSRKRPENVDPGVGKWVSYLKREKKLD 76
N+ +++D + + + + ++ K TE + + + + V V + +++ +
Sbjct: 139 NKDSYLILDTLPSEYDSRVKAMEVDEKPTEDYNDIGGLEKQIQELVEAIVLPMTHKERFE 198
Query: 77 SLVGRRPTAPPAPKGLYIYGNVGSGKTML 105
L G RP PKG+ +YG G+GKT++
Sbjct: 199 KL-GVRP-----PKGVLLYGPPGTGKTLM 221
>At1g20920 putative RNA helicase
Length = 1166
Score = 28.9 bits (63), Expect = 7.4
Identities = 16/53 (30%), Positives = 30/53 (56%), Gaps = 1/53 (1%)
Query: 12 EKKRENERRRILMDEVEKQQNDKDWWKRLNNKITERWTNSRKRPENVDPGVGK 64
E++ E+E+++ L +EVEK++ W+ L K E + S+ + +P GK
Sbjct: 265 EEELEDEQKK-LDEEVEKRRRRVQEWQELKRKKEEAESESKGDADGNEPKAGK 316
>At2g26140 FtsH like protease
Length = 717
Score = 28.5 bits (62), Expect = 9.7
Identities = 16/40 (40%), Positives = 22/40 (55%), Gaps = 6/40 (15%)
Query: 66 VSYLKREKKLDSLVGRRPTAPPAPKGLYIYGNVGSGKTML 105
V YL+ K+ L G+ PKG+ + G G+GKTML
Sbjct: 243 VHYLRDPKRFTRLGGK------LPKGVLLVGPPGTGKTML 276
>At1g09100 putative 26S protease regulatory subunit 6A
Length = 423
Score = 28.5 bits (62), Expect = 9.7
Identities = 20/89 (22%), Positives = 44/89 (48%), Gaps = 6/89 (6%)
Query: 17 NERRRILMDEVEKQQNDKDWWKRLNNKITERWTNSRKRPENVDPGVGKWVSYLKREKKLD 76
N+ +++D + + + + ++ K TE + + + + V V + +++ +
Sbjct: 138 NKDSYLILDTLPSEYDSRVKAMEVDEKPTEDYNDIGGLEKQIQELVEAIVLPMTHKEQFE 197
Query: 77 SLVGRRPTAPPAPKGLYIYGNVGSGKTML 105
L G RP PKG+ +YG G+GKT++
Sbjct: 198 KL-GIRP-----PKGVLLYGPPGTGKTLM 220
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.135 0.402
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,256,496
Number of Sequences: 26719
Number of extensions: 541535
Number of successful extensions: 1785
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 13
Number of HSP's that attempted gapping in prelim test: 1752
Number of HSP's gapped (non-prelim): 40
length of query: 532
length of database: 11,318,596
effective HSP length: 104
effective length of query: 428
effective length of database: 8,539,820
effective search space: 3655042960
effective search space used: 3655042960
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 62 (28.5 bits)
Medicago: description of AC147471.10


