

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
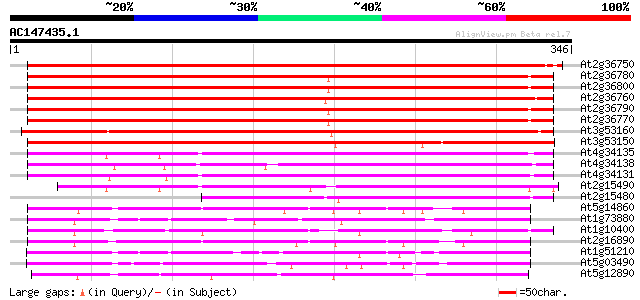
Query= AC147435.1 - phase: 0 /pseudo
(346 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At2g36750 putative glucosyl transferase 360 e-100
At2g36780 putative glucosyl transferase 350 5e-97
At2g36800 putative glucosyl transferase 349 1e-96
At2g36760 putative glucosyl transferase 348 2e-96
At2g36790 putative glucosyl transferase 348 2e-96
At2g36770 putative glucosyl transferase 345 3e-95
At3g53160 glucosyltransferase - like protein 308 2e-84
At3g53150 glucosyltransferase - like protein 294 6e-80
At4g34135 glucosyltransferase like protein 224 4e-59
At4g34138 glucosyltransferase -like protein 223 9e-59
At4g34131 glucosyltransferase -like protein 214 6e-56
At2g15490 putative glucosyltransferase 205 3e-53
At2g15480 glucosyltransferase like protein 167 1e-41
At5g14860 glucosyltransferase -like protein 136 1e-32
At1g73880 putative glucosyltransferase (At1g73880) 135 3e-32
At1g10400 glucosyl transferase - like protein 133 1e-31
At2g16890 glucosyltransferase like protein 132 3e-31
At1g51210 putative glucosyl transferase 124 6e-29
At5g03490 UDPG glucosyltransferase - like protein 119 2e-27
At5g12890 glucosyltransferase -like protein 112 4e-25
>At2g36750 putative glucosyl transferase
Length = 491
Score = 360 bits (923), Expect = e-100
Identities = 178/332 (53%), Positives = 240/332 (71%), Gaps = 5/332 (1%)
Query: 12 HFVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSGLQIKIVT 71
HFVLFP +AQGH+IPM+DIA+LLAQRGV +TI TTP+NA RF +VLSRA+ SGL I +V
Sbjct: 10 HFVLFPFMAQGHMIPMVDIARLLAQRGVTITIVTTPQNAGRFKNVLSRAIQSGLPINLVQ 69
Query: 72 LNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPSCIISDFFI 131
+ FP++++G PEG EN D++DS+ + F A +LL++P E+L + P+P+CII+D +
Sbjct: 70 VKFPSQESGSPEGQENLDLLDSLGASLTFFKAFSLLEEPVEKLLKEIQPRPNCIIADMCL 129
Query: 132 PWTIQIAENHKIPRISFHGFSCFCLHCM-LKIQTSKILERVDSESEYFTVPGIPDQIQVT 190
P+T +IA+N IP+I FHG CF L C + Q + LE ++S+ EYF +P PD+++ T
Sbjct: 130 PYTNRIAKNLGIPKIIFHGMCCFNLLCTHIMHQNHEFLETIESDKEYFPIPNFPDRVEFT 189
Query: 191 KEQIPGIL-KGELKEFGEKMHDAEMKSYGEIINTFEELEKAYVKDYKKEKNGKVWFVGPV 249
K Q+P +L G+ K+F + M + + SYG I+NTFEELE AYV+DYKK K GK+W +GPV
Sbjct: 190 KSQLPMVLVAGDWKDFLDGMTEGDNTSYGVIVNTFEELEPAYVRDYKKVKAGKIWSIGPV 249
Query: 250 SLCNKDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVYACLGSLCNLIPSQLMELALALE 309
SLCNK G D+A+RG A I + C+KWLD + SV+Y CLGS+CNL SQL EL L LE
Sbjct: 250 SLCNKLGEDQAERGNKADIDQDECIKWLDSKEEGSVLYVCLGSICNLPLSQLKELGLGLE 309
Query: 310 ATNRPFIWVIREGNKSSEELEKWISEERNGYR 341
+ RPFIWVIR K +E LE WISE +GY+
Sbjct: 310 ESQRPFIWVIRGWEKYNELLE-WISE--SGYK 338
>At2g36780 putative glucosyl transferase
Length = 496
Score = 350 bits (899), Expect = 5e-97
Identities = 176/327 (53%), Positives = 234/327 (70%), Gaps = 4/327 (1%)
Query: 12 HFVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSGLQIKIVT 71
HFVLFP +AQGH+IPMIDIA+LLAQRGV +TI TTP NA+RF +VL+RA+ SGL I I+
Sbjct: 14 HFVLFPFMAQGHMIPMIDIARLLAQRGVTITIVTTPHNAARFKNVLNRAIESGLAINILH 73
Query: 72 LNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPSCIISDFFI 131
+ FP ++ GLPEG EN D +DS ++ + F A+ LL+ P +L + + P+PSC+ISD+ +
Sbjct: 74 VKFPYQEFGLPEGKENIDSLDSTELMVPFFKAVNLLEDPVMKLMEEMKPRPSCLISDWCL 133
Query: 132 PWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTS-KILERVDSESEYFTVPGIPDQIQVT 190
P+T IA+N IP+I FHG CF L CM ++ + +ILE V S+ EYF VP PD+++ T
Sbjct: 134 PYTSIIAKNFNIPKIVFHGMGCFNLLCMHVLRRNLEILENVKSDEEYFLVPSFPDRVEFT 193
Query: 191 KEQIP--GILKGELKEFGEKMHDAEMKSYGEIINTFEELEKAYVKDYKKEKNGKVWFVGP 248
K Q+P G+ KE ++M AE SYG I+NTF+ELE YVKDYK+ +GKVW +GP
Sbjct: 194 KLQLPVKANASGDWKEIMDEMVKAEYTSYGVIVNTFQELEPPYVKDYKEAMDGKVWSIGP 253
Query: 249 VSLCNKDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVYACLGSLCNLIPSQLMELALAL 308
VSLCNK G DKA+RG A+I + CL+WLD + SV+Y CLGS+CNL SQL EL L L
Sbjct: 254 VSLCNKAGADKAERGSKAAIDQDECLQWLDSKEEGSVLYVCLGSICNLPLSQLKELGLGL 313
Query: 309 EATNRPFIWVIREGNKSSEELEKWISE 335
E + R FIWVIR G++ +EL +W+ E
Sbjct: 314 EESRRSFIWVIR-GSEKYKELFEWMLE 339
>At2g36800 putative glucosyl transferase
Length = 495
Score = 349 bits (896), Expect = 1e-96
Identities = 170/328 (51%), Positives = 235/328 (70%), Gaps = 5/328 (1%)
Query: 12 HFVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSGLQIKIVT 71
HFVLFP +AQGH+IPM+DIA+LLAQRGVI+TI TTP NA+RF +VL+RA+ SGL I +V
Sbjct: 12 HFVLFPFMAQGHMIPMVDIARLLAQRGVIITIVTTPHNAARFKNVLNRAIESGLPINLVQ 71
Query: 72 LNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPSCIISDFFI 131
+ FP +AGL EG EN D +D+++ + F A+ L++P ++L + + P+PSC+ISDF +
Sbjct: 72 VKFPYLEAGLQEGQENIDSLDTMERMIPFFKAVNFLEEPVQKLIEEMNPRPSCLISDFCL 131
Query: 132 PWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTSK-ILERVDSESEYFTVPGIPDQIQVT 190
P+T +IA+ IP+I FHG CFCL CM ++ ++ IL+ + S+ E FTVP PD+++ T
Sbjct: 132 PYTSKIAKKFNIPKILFHGMGCFCLLCMHVLRKNREILDNLKSDKELFTVPDFPDRVEFT 191
Query: 191 KEQIP---GILKGELKEFGEKMHDAEMKSYGEIINTFEELEKAYVKDYKKEKNGKVWFVG 247
+ Q+P + G+ K+ + M +A SYG I+N+F+ELE AY KDYK+ ++GK W +G
Sbjct: 192 RTQVPVETYVPAGDWKDIFDGMVEANETSYGVIVNSFQELEPAYAKDYKEVRSGKAWTIG 251
Query: 248 PVSLCNKDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVYACLGSLCNLIPSQLMELALA 307
PVSLCNK G DKA+RG + I + CLKWLD + SV+Y CLGS+CNL SQL EL L
Sbjct: 252 PVSLCNKVGADKAERGNKSDIDQDECLKWLDSKKHGSVLYVCLGSICNLPLSQLKELGLG 311
Query: 308 LEATNRPFIWVIREGNKSSEELEKWISE 335
LE + RPFIWVIR G + +EL +W SE
Sbjct: 312 LEESQRPFIWVIR-GWEKYKELVEWFSE 338
>At2g36760 putative glucosyl transferase
Length = 496
Score = 348 bits (894), Expect = 2e-96
Identities = 172/327 (52%), Positives = 232/327 (70%), Gaps = 4/327 (1%)
Query: 12 HFVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSGLQIKIVT 71
HFVLFP +AQGH+IPM+DIA++LAQRGV +TI TTP NA+RF VL+RA+ SGL I++
Sbjct: 14 HFVLFPFMAQGHMIPMVDIARILAQRGVTITIVTTPHNAARFKDVLNRAIQSGLHIRVEH 73
Query: 72 LNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPSCIISDFFI 131
+ FP ++AGL EG EN D +DS+++ ++ F A+ +L+ P +L + + PKPSC+ISDF +
Sbjct: 74 VKFPFQEAGLQEGQENVDFLDSMELMVHFFKAVNMLENPVMKLMEEMKPKPSCLISDFCL 133
Query: 132 PWTIQIAENHKIPRISFHGFSCFCLHCM-LKIQTSKILERVDSESEYFTVPGIPDQIQVT 190
P+T +IA+ IP+I FHG SCFCL M + + IL + S+ EYF VP PD+++ T
Sbjct: 134 PYTSKIAKRFNIPKIVFHGVSCFCLLSMHILHRNHNILHALKSDKEYFLVPSFPDRVEFT 193
Query: 191 KEQ--IPGILKGELKEFGEKMHDAEMKSYGEIINTFEELEKAYVKDYKKEKNGKVWFVGP 248
K Q + G+ KE ++ DA+ SYG I+NTF++LE AYVK+Y + + GKVW +GP
Sbjct: 194 KLQVTVKTNFSGDWKEIMDEQVDADDTSYGVIVNTFQDLESAYVKNYTEARAGKVWSIGP 253
Query: 249 VSLCNKDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVYACLGSLCNLIPSQLMELALAL 308
VSLCNK G DKA+RG A+I + C+KWLD +SV+Y CLGS+CNL +QL EL L L
Sbjct: 254 VSLCNKVGEDKAERGNKAAIDQDECIKWLDSKDVESVLYVCLGSICNLPLAQLRELGLGL 313
Query: 309 EATNRPFIWVIREGNKSSEELEKWISE 335
EAT RPFIWVIR G K EL +WI E
Sbjct: 314 EATKRPFIWVIRGGGK-YHELAEWILE 339
>At2g36790 putative glucosyl transferase
Length = 495
Score = 348 bits (893), Expect = 2e-96
Identities = 167/327 (51%), Positives = 238/327 (72%), Gaps = 4/327 (1%)
Query: 12 HFVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSGLQIKIVT 71
HFVLFP +AQGH+IPM+DIA+LLAQRGV++TI TTP NA+RF +VL+RA+ SGL I +V
Sbjct: 13 HFVLFPFMAQGHMIPMVDIARLLAQRGVLITIVTTPHNAARFKNVLNRAIESGLPINLVQ 72
Query: 72 LNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPSCIISDFFI 131
+ FP ++AGL EG EN D++ +++ + F A+ LL++P + L + ++P+PSC+ISD +
Sbjct: 73 VKFPYQEAGLQEGQENMDLLTTMEQITSFFKAVNLLKEPVQNLIEEMSPRPSCLISDMCL 132
Query: 132 PWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTSK-ILERVDSESEYFTVPGIPDQIQVT 190
+T +IA+ KIP+I FHG CFCL C+ ++ ++ IL+ + S+ EYF VP PD+++ T
Sbjct: 133 SYTSEIAKKFKIPKILFHGMGCFCLLCVNVLRKNREILDNLKSDKEYFIVPYFPDRVEFT 192
Query: 191 KEQIP--GILKGELKEFGEKMHDAEMKSYGEIINTFEELEKAYVKDYKKEKNGKVWFVGP 248
+ Q+P + KE E M +A+ SYG I+N+F+ELE AY KD+K+ ++GK W +GP
Sbjct: 193 RPQVPVETYVPAGWKEILEDMVEADKTSYGVIVNSFQELEPAYAKDFKEARSGKAWTIGP 252
Query: 249 VSLCNKDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVYACLGSLCNLIPSQLMELALAL 308
VSLCNK G+DKA+RG + I + CL+WLD +P SV+Y CLGS+CNL SQL+EL L L
Sbjct: 253 VSLCNKVGVDKAERGNKSDIDQDECLEWLDSKEPGSVLYVCLGSICNLPLSQLLELGLGL 312
Query: 309 EATNRPFIWVIREGNKSSEELEKWISE 335
E + RPFIWVIR G + +EL +W SE
Sbjct: 313 EESQRPFIWVIR-GWEKYKELVEWFSE 338
>At2g36770 putative glucosyl transferase
Length = 496
Score = 345 bits (884), Expect = 3e-95
Identities = 174/327 (53%), Positives = 229/327 (69%), Gaps = 4/327 (1%)
Query: 12 HFVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSGLQIKIVT 71
HF+LFP +AQGH+IPMIDIA+LLAQRG VTI TT NA RF +VLSRA+ SGL I IV
Sbjct: 14 HFILFPFMAQGHMIPMIDIARLLAQRGATVTIVTTRYNAGRFENVLSRAMESGLPINIVH 73
Query: 72 LNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPSCIISDFFI 131
+NFP ++ GLPEG EN D DS+++ + F A+ +L+ P +L + + P+PSCIISD +
Sbjct: 74 VNFPYQEFGLPEGKENIDSYDSMELMVPFFQAVNMLEDPVMKLMEEMKPRPSCIISDLLL 133
Query: 132 PWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTS-KILERVDSESEYFTVPGIPDQIQVT 190
P+T +IA IP+I FHG CF L CM ++ + +IL+ + S+ +YF VP PD+++ T
Sbjct: 134 PYTSKIARKFSIPKIVFHGTGCFNLLCMHVLRRNLEILKNLKSDKDYFLVPSFPDRVEFT 193
Query: 191 KEQIP--GILKGELKEFGEKMHDAEMKSYGEIINTFEELEKAYVKDYKKEKNGKVWFVGP 248
K Q+P G+ K F ++M +AE SYG I+NTF+ELE AYVKDY K + GKVW +GP
Sbjct: 194 KPQVPVETTASGDWKAFLDEMVEAEYTSYGVIVNTFQELEPAYVKDYTKARAGKVWSIGP 253
Query: 249 VSLCNKDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVYACLGSLCNLIPSQLMELALAL 308
VSLCNK G DKA+RG A+I + CL+WLD + SV+Y CLGS+CNL SQL EL L L
Sbjct: 254 VSLCNKAGADKAERGNQAAIDQDECLQWLDSKEDGSVLYVCLGSICNLPLSQLKELGLGL 313
Query: 309 EATNRPFIWVIREGNKSSEELEKWISE 335
E + R FIWVIR G + EL +W+ E
Sbjct: 314 EKSQRSFIWVIR-GWEKYNELYEWMME 339
>At3g53160 glucosyltransferase - like protein
Length = 490
Score = 308 bits (790), Expect = 2e-84
Identities = 158/332 (47%), Positives = 228/332 (68%), Gaps = 6/332 (1%)
Query: 8 NDVPHFVLFPLIAQGHIIPMIDIAKLLAQR-GVIVTIFTTPKNASRFTSVLSRAVSSGLQ 66
+D HFV+ P +AQGH+IP++DI++LL+QR GV V I TT +N ++ + LS + S
Sbjct: 4 HDPLHFVVIPFMAQGHMIPLVDISRLLSQRQGVTVCIITTTQNVAKIKTSLSFS-SLFAT 62
Query: 67 IKIVTLNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALT-PKPSCI 125
I IV + F ++Q GLPEGCE+ DM+ S+ + F A L++ E+ + + P+PSCI
Sbjct: 63 INIVEVKFLSQQTGLPEGCESLDMLASMGDMVKFFDAANSLEEQVEKAMEEMVQPRPSCI 122
Query: 126 ISDFFIPWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTSKILERVDSESEYFTVPGIPD 185
I D +P+T ++A+ KIP++ FHGFSCF L + ++ S IL+ ++S EYF +PG+PD
Sbjct: 123 IGDMSLPFTSRLAKKFKIPKLIFHGFSCFSLMSIQVVRESGILKMIESNDEYFDLPGLPD 182
Query: 186 QIQVTKEQIPGI--LKGELKEFGEKMHDAEMKSYGEIINTFEELEKAYVKDYKKEKNGKV 243
+++ TK Q+ + ++G +KE K+ +A+ SYG I+NTFEELE Y ++Y+K + GKV
Sbjct: 183 KVEFTKPQVSVLQPVEGNMKESTAKIIEADNDSYGVIVNTFEELEVDYAREYRKARAGKV 242
Query: 244 WFVGPVSLCNKDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVYACLGSLCNLIPSQLME 303
W VGPVSLCN+ GLDKA+RG ASI + CL+WLD + SV+Y CLGSLCNL +QL E
Sbjct: 243 WCVGPVSLCNRLGLDKAKRGDKASIGQDQCLQWLDSQETGSVLYVCLGSLCNLPLAQLKE 302
Query: 304 LALALEATNRPFIWVIREGNKSSEELEKWISE 335
L L LEA+N+PFIWVIRE K +L W+ +
Sbjct: 303 LGLGLEASNKPFIWVIREWGKYG-DLANWMQQ 333
>At3g53150 glucosyltransferase - like protein
Length = 507
Score = 294 bits (752), Expect = 6e-80
Identities = 154/331 (46%), Positives = 211/331 (63%), Gaps = 7/331 (2%)
Query: 12 HFVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRA-VSSGLQIKIV 70
HFVL PL+AQGH+IPM+DI+K+LA++G IVTI TTP+NASRF + RA + SGL+I +V
Sbjct: 13 HFVLIPLMAQGHLIPMVDISKILARQGNIVTIVTTPQNASRFAKTVDRARLESGLEINVV 72
Query: 71 TLNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPSCIISDFF 130
P K+ GLP+ CE D + S D+ + A+ LQ+P E + PSCIISD
Sbjct: 73 KFPIPYKEFGLPKDCETLDTLPSKDLLRRFYDAVDKLQEPMERFLEQQDIPPSCIISDKC 132
Query: 131 IPWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTSKILERVDSESEYFTVPGIPDQIQVT 190
+ WT + A+ KIPRI FHG CF L I V S E F +PG+P +I++
Sbjct: 133 LFWTSRTAKRFKIPRIVFHGMCCFSLLSSHNIHLHSPHLSVSSAVEPFPIPGMPHRIEIA 192
Query: 191 KEQIPGILK--GELKEFGEKMHDAEMKSYGEIINTFEELEKAYVKDYKKEKNGKVWFVGP 248
+ Q+PG + + + EKM ++E +++G I+N+F+ELE Y + Y + N KVWFVGP
Sbjct: 193 RAQLPGAFEKLANMDDVREKMRESESEAFGVIVNSFQELEPGYAEAYAEAINKKVWFVGP 252
Query: 249 VSLCN---KDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVYACLGSLCNLIPSQLMELA 305
VSLCN D D+ G IA ISE CL++LD +P+SV+Y LGSLC LIP+QL+EL
Sbjct: 253 VSLCNDRMADLFDRGSNGNIA-ISETECLQFLDSMRPRSVLYVSLGSLCRLIPNQLIELG 311
Query: 306 LALEATNRPFIWVIREGNKSSEELEKWISEE 336
L LE + +PFIWVI+ K EL++W+ E
Sbjct: 312 LGLEESGKPFIWVIKTEEKHMIELDEWLKRE 342
>At4g34135 glucosyltransferase like protein
Length = 483
Score = 224 bits (572), Expect = 4e-59
Identities = 125/333 (37%), Positives = 185/333 (55%), Gaps = 14/333 (4%)
Query: 12 HFVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLS--RAVSSGLQIKI 69
H + FP +A GH+IP +D+AKL + RG TI TT N+ + + ++ GL+I I
Sbjct: 11 HVMFFPFMAYGHMIPTLDMAKLFSSRGAKSTILTTSLNSKILQKPIDTFKNLNPGLEIDI 70
Query: 70 VTLNFPTKQAGLPEGCENFDMV------DSIDMRMNLFHAITLLQKPAEELFDALTPKPS 123
NFP + GLPEGCEN D D +M + F + + E+L T +P
Sbjct: 71 QIFNFPCVELGLPEGCENVDFFTSNNNDDKNEMIVKFFFSTRFFKDQLEKLLG--TTRPD 128
Query: 124 CIISDFFIPWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTSKILERVDSESEYFTVPGI 183
C+I+D F PW + A +PR+ FHG F L I K +RV S SE F +P +
Sbjct: 129 CLIADMFFPWATEAAGKFNVPRLVFHGTGYFSLCAGYCIGVHKPQKRVASSSEPFVIPEL 188
Query: 184 PDQIQVTKEQI-PGILKGELKEFGEKMHDAEMKSYGEIINTFEELEKAYVKDYKKEKNGK 242
P I +T+EQI G + ++ +F ++ ++E+KS G ++N+F ELE Y YK +
Sbjct: 189 PGNIVITEEQIIDGDGESDMGKFMTEVRESEVKSSGVVLNSFYELEHDYADFYKSCVQKR 248
Query: 243 VWFVGPVSLCNKDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVYACLGSLCNLIPSQLM 302
W +GP+S+ N+ +KA+RG A+I E CLKWLD +P SV+Y GS+ QL
Sbjct: 249 AWHIGPLSVYNRGFEEKAERGKKANIDEAECLKWLDSKKPNSVIYVSFGSVAFFKNEQLF 308
Query: 303 ELALALEATNRPFIWVIREGNKSSEELEKWISE 335
E+A LEA+ FIWV+R K+ ++ E+W+ E
Sbjct: 309 EIAAGLEASGTSFIWVVR---KTKDDREEWLPE 338
>At4g34138 glucosyltransferase -like protein
Length = 488
Score = 223 bits (569), Expect = 9e-59
Identities = 127/339 (37%), Positives = 194/339 (56%), Gaps = 25/339 (7%)
Query: 12 HFVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSS---GLQ-I 67
HF+LFP +A GH+IP +D+AKL A +G TI TTP NA F ++ + GL+ I
Sbjct: 11 HFLLFPFMAHGHMIPTLDMAKLFATKGAKSTILTTPLNAKLFFEKPIKSFNQDNPGLEDI 70
Query: 68 KIVTLNFPTKQAGLPEGCENFDMVDSI------DMRMNLFHAITLLQKPAEELFDALTPK 121
I LNFP + GLP+GCEN D + S D+ A+ ++P EEL +T +
Sbjct: 71 TIQILNFPCTELGLPDGCENTDFIFSTPDLNVGDLSQKFLLAMKYFEEPLEELL--VTMR 128
Query: 122 PSCIISDFFIPWTIQIAENHKIPRISFHGFSCFCL---HCMLKIQTSKILERVDSESEYF 178
P C++ + F PW+ ++AE +PR+ FHG F L HC+ ++ + V + SE F
Sbjct: 129 PDCLVGNMFFPWSTKVAEKFGVPRLVFHGTGYFSLCASHCI------RLPKNVATSSEPF 182
Query: 179 TVPGIPDQIQVTKEQIPGILKGELK-EFGEKMHDAEMKSYGEIINTFEELEKAYVKDYKK 237
+P +P I +T+EQ+ + + F + + D+E S+G ++N+F ELE+AY +K
Sbjct: 183 VIPDLPGDILITEEQVMETEEESVMGRFMKAIRDSERDSFGVLVNSFYELEQAYSDYFKS 242
Query: 238 EKNGKVWFVGPVSLCNKDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVYACLGSLCNLI 297
+ W +GP+SL N+ +KA+RG ASI EH CLKWLD + SV+Y G++ +
Sbjct: 243 FVAKRAWHIGPLSLGNRKFEEKAERGKKASIDEHECLKWLDSKKCDSVIYMAFGTMSSFK 302
Query: 298 PSQLMELALALEATNRPFIWVI-REGNKSSEELEKWISE 335
QL+E+A L+ + F+WV+ R+G S E E W+ E
Sbjct: 303 NEQLIEIAAGLDMSGHDFVWVVNRKG--SQVEKEDWLPE 339
>At4g34131 glucosyltransferase -like protein
Length = 481
Score = 214 bits (545), Expect = 6e-56
Identities = 123/333 (36%), Positives = 180/333 (53%), Gaps = 13/333 (3%)
Query: 12 HFVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRA--VSSGLQIKI 69
H V FP +A GH+IP +D+AKL + RG TI TTP N+ F + R ++ +I I
Sbjct: 10 HVVFFPFMAYGHMIPTLDMAKLFSSRGAKSTILTTPLNSKIFQKPIERFKNLNPSFEIDI 69
Query: 70 VTLNFPTKQAGLPEGCENFDMVDSID------MRMNLFHAITLLQKPAEELFDALTPKPS 123
+FP GLPEGCEN D S + + + F + + E+L + T +P
Sbjct: 70 QIFDFPCVDLGLPEGCENVDFFTSNNNDDRQYLTLKFFKSTRFFKDQLEKLLE--TTRPD 127
Query: 124 CIISDFFIPWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTSKILERVDSESEYFTVPGI 183
C+I+D F PW + AE +PR+ FHG F L I+ V S E F +P +
Sbjct: 128 CLIADMFFPWATEAAEKFNVPRLVFHGTGYFSLCSEYCIRVHNPQNIVASRYEPFVIPDL 187
Query: 184 PDQIQVTKEQIPGI-LKGELKEFGEKMHDAEMKSYGEIINTFEELEKAYVKDYKKEKNGK 242
P I +T+EQI + E+ +F ++ ++++KS G I+N+F ELE Y YK +
Sbjct: 188 PGNIVITQEQIADRDEESEMGKFMIEVKESDVKSSGVIVNSFYELEPDYADFYKSVVLKR 247
Query: 243 VWFVGPVSLCNKDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVYACLGSLCNLIPSQLM 302
W +GP+S+ N+ +KA+RG ASI+E CLKWLD +P SV+Y GS+ QL
Sbjct: 248 AWHIGPLSVYNRGFEEKAERGKKASINEVECLKWLDSKKPDSVIYISFGSVACFKNEQLF 307
Query: 303 ELALALEATNRPFIWVIREGNKSSEELEKWISE 335
E+A LE + FIWV+R+ E E+W+ E
Sbjct: 308 EIAAGLETSGANFIWVVRK--NIGIEKEEWLPE 338
>At2g15490 putative glucosyltransferase
Length = 460
Score = 205 bits (521), Expect = 3e-53
Identities = 122/329 (37%), Positives = 175/329 (53%), Gaps = 28/329 (8%)
Query: 30 IAKLLAQRGVIVTIFTTPKNASRFTSVLS--RAVSSGLQIKIVTLNFPTKQAGLPEGCEN 87
+AKL A+RG T+ TTP NA + + + L+I I LNFP + GLPEGCEN
Sbjct: 1 MAKLFARRGAKSTLLTTPINAKILEKPIEAFKVQNPDLEIGIKILNFPCVELGLPEGCEN 60
Query: 88 FDMV------DSIDMRMNLFHAITLLQKPAEELFDALTPKPSCIISDFFIPWTIQIAENH 141
D + DS D+ + + +++ E + T KPS +++D F PW + AE
Sbjct: 61 RDFINSYQKSDSFDLFLKFLFSTKYMKQQLESFIE--TTKPSALVADMFFPWATESAEKI 118
Query: 142 KIPRISFHGFSCFCLHCMLKIQTSKILERVDSESEYFTVPGIP-------DQIQVTKEQI 194
+PR+ FHG S F L C ++ K ++V S S F +PG+P DQ VT E+
Sbjct: 119 GVPRLVFHGTSSFALCCSYNMRIHKPHKKVASSSTPFVIPGLPGDIVITEDQANVTNEET 178
Query: 195 PGILKGELKEFGEKMHDAEMKSYGEIINTFEELEKAYVKDYKKEKNGKVWFVGPVSLCNK 254
P +F +++ ++E S+G ++N+F ELE +Y Y+ K W +GP+SL N+
Sbjct: 179 P------FGKFWKEVRESETSSFGVLVNSFYELESSYADFYRSFVAKKAWHIGPLSLSNR 232
Query: 255 DGLDKAQRGIIASISEHHCLKWLDLHQPKSVVYACLGSLCNLIPSQLMELALALEATNRP 314
+KA RG A+I E CLKWLD P SVVY GS L QL+E+A LE + +
Sbjct: 233 GIAEKAGRGKKANIDEQECLKWLDSKTPGSVVYLSFGSGTGLPNEQLLEIAFGLEGSGQN 292
Query: 315 FIWVI--REGNKSSEELEKWIS---EERN 338
FIWV+ E + E E W+ EERN
Sbjct: 293 FIWVVSKNENQVGTGENEDWLPKGFEERN 321
>At2g15480 glucosyltransferase like protein
Length = 372
Score = 167 bits (422), Expect = 1e-41
Identities = 85/219 (38%), Positives = 133/219 (59%), Gaps = 4/219 (1%)
Query: 119 TPKPSCIISDFFIPWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTSKILERVDSESEYF 178
T KPS +++D F PW + AE +PR+ FHG S F L C ++ K ++V + S F
Sbjct: 11 TTKPSALVADMFFPWATESAEKLGVPRLVFHGTSFFSLCCSYNMRIHKPHKKVATSSTPF 70
Query: 179 TVPGIPDQIQVTKEQIPGILKGE--LKEFGEKMHDAEMKSYGEIINTFEELEKAYVKDYK 236
+PG+P I +T++Q + K E + +F +++ ++E S+G ++N+F ELE AY Y+
Sbjct: 71 VIPGLPGDIVITEDQA-NVAKEETPMGKFMKEVRESETNSFGVLVNSFYELESAYADFYR 129
Query: 237 KEKNGKVWFVGPVSLCNKDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVYACLGSLCNL 296
+ W +GP+SL N++ +KA+RG A+I E CLKWLD P SVVY GS N
Sbjct: 130 SFVAKRAWHIGPLSLSNRELGEKARRGKKANIDEQECLKWLDSKTPGSVVYLSFGSGTNF 189
Query: 297 IPSQLMELALALEATNRPFIWVIREGNKSSEELEKWISE 335
QL+E+A LE + + FIWV+R+ N++ + E+W+ E
Sbjct: 190 TNDQLLEIAFGLEGSGQSFIWVVRK-NENQGDNEEWLPE 227
>At5g14860 glucosyltransferase -like protein
Length = 492
Score = 136 bits (343), Expect = 1e-32
Identities = 114/335 (34%), Positives = 164/335 (48%), Gaps = 42/335 (12%)
Query: 12 HFVLFPLIAQGHIIPMIDIAKLLAQRGVIV-----------TIFTTPKNASRFTSVLSRA 60
H VLFP +++GH IP++ A+LL + IV T+FTTPKN ++ LS
Sbjct: 8 HAVLFPYMSKGHTIPLLQFARLLLRHRRIVSVDDEEPTISVTVFTTPKNQPFVSNFLSDV 67
Query: 61 VSSGLQIKIVTLNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTP 120
SS IK+++L FP AG+P G E+ DM+ SI + + A LQ E L
Sbjct: 68 ASS---IKVISLPFPENIAGIPPGVESTDMLPSISLYVPFTRATKSLQPFFEAELKNL-E 123
Query: 121 KPSCIISDFFIPWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTSKIL---ERVDSESEY 177
K S ++SD F+ WT + A +IPR++F+G + + I ++ E V S++E
Sbjct: 124 KVSFMVSDGFLWWTSESAAKFEIPRLAFYGMNSYASAMCSAISVHELFTKPESVKSDTEP 183
Query: 178 FTVPGIPDQIQVTKEQIPGIL--KGELKEFGEKMHDAEM---KSYGEIINTFEELEKAYV 232
TVP P I V K + +L + E + D M KS G I+N+F ELE +V
Sbjct: 184 VTVPDFP-WICVKKCEFDPVLTEPDQSDPAFELLIDHLMSTKKSRGVIVNSFYELESTFV 242
Query: 233 KDYKKEKNG--KVWFVGPVSLCN--KDGLDKAQRGIIASISEHHCLKWLD--LHQPKSVV 286
DY+ N K W VGP+ L N K DK + WLD L + V+
Sbjct: 243 -DYRLRDNDEPKPWCVGPLCLVNPPKPESDKPD-----------WIHWLDRKLEERCPVM 290
Query: 287 YACLGSLCNLIPSQLMELALALEATNRPFIWVIRE 321
Y G+ + QL E+AL LE + F+WV R+
Sbjct: 291 YVAFGTQAEISNEQLKEIALGLEDSKVNFLWVTRK 325
>At1g73880 putative glucosyltransferase (At1g73880)
Length = 473
Score = 135 bits (340), Expect = 3e-32
Identities = 100/322 (31%), Positives = 156/322 (48%), Gaps = 29/322 (9%)
Query: 12 HFVLFPLIAQGHIIPMIDIAKLLAQRG---VIVTIFTTPKNASRFTSVLSRAVSSGLQIK 68
H ++FP AQGH+IP++D LA RG + +T+ TPKN + +LS V+ I+
Sbjct: 14 HVLIFPFPAQGHMIPLLDFTHRLALRGGAALKITVLVTPKNLPFLSPLLSAVVN----IE 69
Query: 69 IVTLNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPSCIISD 128
+ L FP+ + +P G EN + + + HA+ L P + P I+SD
Sbjct: 70 PLILPFPSHPS-IPSGVENVQDLPPSGFPL-MIHALGNLHAPLISWITSHPSPPVAIVSD 127
Query: 129 FFIPWTIQIAENHKIPRISFH---GFSCFCLHCMLKIQTSKILERVDSESEYFTVPGIPD 185
FF+ WT +N IPR F +C L+ + +KI E D+E +F P IP+
Sbjct: 128 FFLGWT----KNLGIPRFDFSPSAAITCCILNTLWIEMPTKINEDDDNEILHF--PKIPN 181
Query: 186 QIQVTKEQIPGILKGELK-----EFGEKMHDAEMKSYGEIINTFEELEKAYVKDYKKEK- 239
+ +QI + + + EF + S+G ++N+F +E Y++ K+E
Sbjct: 182 CPKYRFDQISSLYRSYVHGDPAWEFIRDSFRDNVASWGLVVNSFTAMEGVYLEHLKREMG 241
Query: 240 NGKVWFVGPVSLCNKDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVYACLGSLCNLIPS 299
+ +VW VGP+ + D RG S+S H + WLD + VVY C GS L
Sbjct: 242 HDRVWAVGPIIPLSGDN-----RGGPTSVSVDHVMSWLDAREDNHVVYVCFGSQVVLTKE 296
Query: 300 QLMELALALEATNRPFIWVIRE 321
Q + LA LE + FIW ++E
Sbjct: 297 QTLALASGLEKSGVHFIWAVKE 318
>At1g10400 glucosyl transferase - like protein
Length = 467
Score = 133 bits (335), Expect = 1e-31
Identities = 104/350 (29%), Positives = 166/350 (46%), Gaps = 51/350 (14%)
Query: 12 HFVLFPLIAQGHIIPMIDIAKLLAQRG----VIVTIFTTPKNASRFTSVLSRAVSSGLQI 67
H VLFP +++GH+IPM+ +A+LL + VT+FTTP N LS G +
Sbjct: 7 HVVLFPYLSKGHMIPMLQLARLLLSHSFAGDISVTVFTTPLNRPFIVDSLS-----GTKA 61
Query: 68 KIVTLNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDA---LTPKPSC 124
IV + FP +P G E D + ++ +LF T K + F+ P+ S
Sbjct: 62 TIVDVPFPDNVPEIPPGVECTDKLPALSS--SLFVPFTRATKSMQADFERELMSLPRVSF 119
Query: 125 IISDFFIPWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTSKILERVDSESEYFTVPGIP 184
++SD F+ WT + A PR+ F G +C + +++L V SE+E +VP P
Sbjct: 120 MVSDGFLWWTQESARKLGFPRLVFFGMNCASTVICDSVFQNQLLSNVKSETEPVSVPEFP 179
Query: 185 DQIQVTKEQIPGILKGELKEFGEKMHDAEM-----------------KSYGEIINTFEEL 227
I+V K +F + M D + +S G I NTF++L
Sbjct: 180 -WIKVRK-----------CDFVKDMFDPKTTTDPGFKLILDQVTSMNQSQGIIFNTFDDL 227
Query: 228 EKAYVKDYKKEKNGKVWFVGPVSLCNKDGLDKAQRGIIASISEHHCLKWLDLHQPK--SV 285
E ++ YK+++ K+W VGP+ N D+ + + S +KWLD + K +V
Sbjct: 228 EPVFIDFYKRKRKLKLWAVGPLCYVNNFLDDEVEEKVKPS-----WMKWLDEKRDKGCNV 282
Query: 286 VYACLGSLCNLIPSQLMELALALEATNRPFIWVIREGNKSSEELEKWISE 335
+Y GS + QL E+AL LE + F+WV++ GN+ + E+ + E
Sbjct: 283 LYVAFGSQAEISREQLEEIALGLEESKVNFLWVVK-GNEIGKGFEERVGE 331
>At2g16890 glucosyltransferase like protein
Length = 478
Score = 132 bits (332), Expect = 3e-31
Identities = 103/326 (31%), Positives = 155/326 (46%), Gaps = 32/326 (9%)
Query: 12 HFVLFPLIAQGHIIPMIDIAKLLAQRG-----VIVTIFTTPKNASRFTSVLSRAVSSGLQ 66
H VLFP +++GHIIP++ +LL + + VT+FTTPKN + LS +
Sbjct: 9 HVVLFPFMSKGHIIPLLQFGRLLLRHHRKEPTITVTVFTTPKNQPFISDFLSDTP----E 64
Query: 67 IKIVTLNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPSCII 126
IK+++L FP G+P G EN + + S+ + + A LLQ EE L PK S ++
Sbjct: 65 IKVISLPFPENITGIPPGVENTEKLPSMSLFVPFTRATKLLQPFFEETLKTL-PKVSFMV 123
Query: 127 SDFFIPWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTSKILERVDSES--EYFTVPGIP 184
SD F+ WT + A IPR +G + + + + ++ +S+S E TVP P
Sbjct: 124 SDGFLWWTSESAAKFNIPRFVSYGMNSYSAAVSISVFKHELFTEPESKSDTEPVTVPDFP 183
Query: 185 DQIQVTKEQIP-GILK----GELKEFGEKMHDAEMKSYGEIINTFEELEKAYVKDYKKEK 239
I+V K G + G E + S+G ++N+F ELE A+V DY
Sbjct: 184 -WIKVKKCDFDHGTTEPEESGAALELSMDQIKSTTTSHGFLVNSFYELESAFV-DYNNNS 241
Query: 240 NG--KVWFVGPVSLCNKDGLDKAQRGIIASISEHHCLKWLD--LHQPKSVVYACLGSLCN 295
K W VGP+ L + A+ I WLD + + V+Y G+
Sbjct: 242 GDKPKSWCVGPLCLTDPPKQGSAKPAWI---------HWLDQKREEGRPVLYVAFGTQAE 292
Query: 296 LIPSQLMELALALEATNRPFIWVIRE 321
+ QLMELA LE + F+WV R+
Sbjct: 293 ISNKQLMELAFGLEDSKVNFLWVTRK 318
>At1g51210 putative glucosyl transferase
Length = 433
Score = 124 bits (312), Expect = 6e-29
Identities = 98/316 (31%), Positives = 156/316 (49%), Gaps = 23/316 (7%)
Query: 11 PHFVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSGLQIKIV 70
PH ++FP AQGH++P++D+ L RG+ V+I TPKN + +LS S+ + +V
Sbjct: 19 PHIMVFPYPAQGHLLPLLDLTHQLCLRGLTVSIIVTPKNLPYLSPLLSAHPSA---VSVV 75
Query: 71 TLNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPSCIISDFF 130
TL FP +P G EN + + + ++ L++P + P +ISDFF
Sbjct: 76 TLPFPHHPL-IPSGVENVKDLGGYGNPL-IMASLRQLREPIVNWLSSHPNPPVALISDFF 133
Query: 131 IPWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTSKILERVDSESEYFTVPGIPDQIQVT 190
+ WT + IPR +F F L +L + K + +E + +P
Sbjct: 134 LGWTKDLG----IPRFAFFSSGAF-LASILHFVSDK--PHLFESTEPVCLSDLPRSPVFK 186
Query: 191 KEQIPGIL-KGELKEFGEKMHDAEM--KSYGEIINTFEELEKAYVKDYKKEK--NGKVWF 245
E +P ++ + L + E + D+ M SYG I NT E LE+ Y+ +Y K+K +V+
Sbjct: 187 TEHLPSLIPQSPLSQDLESVKDSTMNFSSYGCIFNTCECLEEDYM-EYVKQKVSENRVFG 245
Query: 246 VGPVSLCNKDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVYACLGSLCNLIPSQLMELA 305
VGP+S GL K ++++ L WLD SV+Y C GS L Q +LA
Sbjct: 246 VGPLS---SVGLSKEDS--VSNVDAKALLSWLDGCPDDSVLYICFGSQKVLTKEQCDDLA 300
Query: 306 LALEATNRPFIWVIRE 321
L LE + F+WV+++
Sbjct: 301 LGLEKSMTRFVWVVKK 316
>At5g03490 UDPG glucosyltransferase - like protein
Length = 465
Score = 119 bits (299), Expect = 2e-27
Identities = 95/320 (29%), Positives = 151/320 (46%), Gaps = 26/320 (8%)
Query: 11 PHFVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSGLQIKIV 70
PH V+FP AQGH++P++D+ L RG V++ TP N + + +LS SS + V
Sbjct: 18 PHIVVFPFPAQGHLLPLLDLTHQLCLRGFNVSVIVTPGNLTYLSPLLSAHPSS---VTSV 74
Query: 71 TLNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPSCIISDFF 130
FP L G EN V + + + ++ L++P F + P +ISDFF
Sbjct: 75 VFPFP-PHPSLSPGVENVKDVGN-SGNLPIMASLRQLREPIINWFQSHPNPPIALISDFF 132
Query: 131 IPWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTSKILERVD--SESEYFTVPGIPDQIQ 188
+ WT + IPR +F S F + + E +D ++ + +P
Sbjct: 133 LGWTHDLCNQIGIPRFAFFSISFFLVSVL-----QFCFENIDLIKSTDPIHLLDLPRAPI 187
Query: 189 VTKEQIPGILKGELKEFG---EKMHDAEMK--SYGEIINTFEELEKAYVKDYKKEKNG-- 241
+E +P I++ L+ E + D M SYG + N+ E LE Y++ Y K++ G
Sbjct: 188 FKEEHLPSIVRRSLQTPSPDLESIKDFSMNLLSYGSVFNSSEILEDDYLQ-YVKQRMGHD 246
Query: 242 KVWFVGPVSLCNKDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVYACLGSLCNLIPSQL 301
+V+ +GP LC+ K+ G + + L WLD SV+Y C GS L Q
Sbjct: 247 RVYVIGP--LCSIGSGLKSNSGSV----DPSLLSWLDGSPNGSVLYVCFGSQKALTKDQC 300
Query: 302 MELALALEATNRPFIWVIRE 321
LAL LE + F+WV+++
Sbjct: 301 DALALGLEKSMTRFVWVVKK 320
>At5g12890 glucosyltransferase -like protein
Length = 488
Score = 112 bits (279), Expect = 4e-25
Identities = 81/323 (25%), Positives = 149/323 (46%), Gaps = 30/323 (9%)
Query: 14 VLFPLIAQGHIIPMIDIAKLLAQRGVI-------VTIFTTPKNASRFTSVLSRAVSSGLQ 66
V+FP + QGHIIP + +A L + ++ +++ TP N + S L S
Sbjct: 12 VMFPFMGQGHIIPFVALALRLEKIMIMNRANKTTISMINTPSNIPKIRSNLPPESS---- 67
Query: 67 IKIVTLNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPS--- 123
I ++ L F + GLP ENFD + + ++L A L++P + + +
Sbjct: 68 ISLIELPFNSSDHGLPHDGENFDSLP-YSLVISLLEASRSLREPFRDFMTKILKEEGQSS 126
Query: 124 -CIISDFFIPWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTSKILERVDSESEYFTVPG 182
+I DFF+ W ++ + + + F F L C I + L +++ + F +
Sbjct: 127 VIVIGDFFLGWIGKVCKEVGVYSVIFSASGAFGLGCYRSIWLN--LPHKETKQDQFLLDD 184
Query: 183 IPDQIQVTKEQIPGIL-----KGELKEFGEKMHDAEMKSYGEIINTFEELEKAYVKDYKK 237
P+ ++ K Q+ + + F +K+ G + NT E+++ + +++
Sbjct: 185 FPEAGEIEKTQLNSFMLEADGTDDWSVFMKKIIPGWSDFDGFLFNTVAEIDQMGLSYFRR 244
Query: 238 EKNGKVWFVGPVSLCNKDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVYACLGSLCNLI 297
VW VGPV L + + + +E WLD SVVY C GS+ +++
Sbjct: 245 ITGVPVWPVGPV-------LKSPDKKVGSRSTEEAVKSWLDSKPDHSVVYVCFGSMNSIL 297
Query: 298 PSQLMELALALEATNRPFIWVIR 320
+ ++ELA+ALE++ + FIWV+R
Sbjct: 298 QTHMLELAMALESSEKNFIWVVR 320
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.322 0.139 0.419
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,948,769
Number of Sequences: 26719
Number of extensions: 340456
Number of successful extensions: 1177
Number of sequences better than 10.0: 130
Number of HSP's better than 10.0 without gapping: 118
Number of HSP's successfully gapped in prelim test: 12
Number of HSP's that attempted gapping in prelim test: 867
Number of HSP's gapped (non-prelim): 167
length of query: 346
length of database: 11,318,596
effective HSP length: 100
effective length of query: 246
effective length of database: 8,646,696
effective search space: 2127087216
effective search space used: 2127087216
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 60 (27.7 bits)
Medicago: description of AC147435.1


