

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147430.8 + phase: 1 /pseudo
(437 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
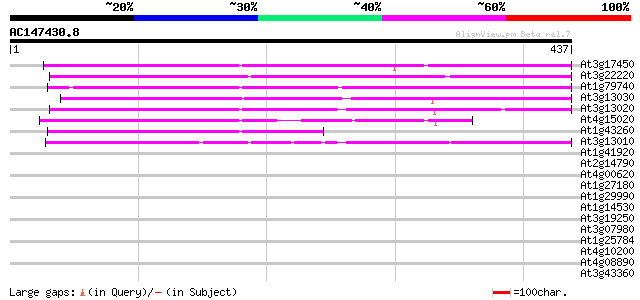
Score E
Sequences producing significant alignments: (bits) Value
At3g17450 unknown protein 278 3e-75
At3g22220 hypothetical protein 245 4e-65
At1g79740 hypothetical protein 200 1e-51
At3g13030 unknown protein (At3g13030) 199 2e-51
At3g13020 hypothetical protein 196 3e-50
At4g15020 putative protein 167 8e-42
At1g43260 hypothetical protein 167 1e-41
At3g13010 hypothetical protein 164 9e-41
At1g41920 hypothetical protein 31 1.2
At2g14790 Mutator-like transposase 30 2.0
At4g00620 putative tetrahydrofolate synthase 30 2.6
At1g27180 disease resistance protein, putative 30 2.6
At1g29990 hydrophilic like protein 29 4.5
At1g14530 unknown protein 29 4.5
At3g19250 hypothetical protein 29 5.8
At3g07980 putative MAP3K epsilon protein kinase 29 5.8
At1g25784 mutator-like transposase, putative 29 5.8
At4g10200 putative protein 28 7.6
At4g08890 putative transposon protein 28 7.6
At3g43360 putative transposase 28 7.6
>At3g17450 unknown protein
Length = 877
Score = 278 bits (712), Expect = 3e-75
Identities = 149/418 (35%), Positives = 239/418 (56%), Gaps = 10/418 (2%)
Query: 27 FNIMCDMISRHGNGFKPPSYYDIRVKLLNQEVKSTNDALEEHKREWKKTSCTIMTDEWTD 86
F M ++I +G GF PS +LL +E+ + L E++ W T C+IM D WT+
Sbjct: 325 FQKMIELIGMYGEGFVVPSSQLFSGRLLQEEMSTIKSYLREYRSSWVVTGCSIMADTWTN 384
Query: 87 RRRRTILNFLVNSPRGTVFLKSVDASNICKTSEKIFEMIDAVVEEVGEDNVVQVVTDNAA 146
+ +++FLV+ PRG F S+DA++I + + +F+ +D +V+++GE+NVVQV+T N A
Sbjct: 385 TEGKKMISFLVSCPRGVYFHSSIDATDIVEDALSLFKCLDKLVDDIGEENVVQVITQNTA 444
Query: 147 NYKAAGEMLMRKRKKLYWTPCAAHCLDLMLEDLEKKIPIHGETIPKGKKITTFIYARTSL 206
+++AG++L KRK LYWTPCA HC +L+LED K+ E + K ++IT FIY +T L
Sbjct: 445 IFRSAGKLLEEKRKNLYWTPCAIHCTELVLEDF-SKLEFVSECLEKAQRITRFIYNQTWL 503
Query: 207 ISIL-HSHTKNRDLVRPGATRFATSYLTLGCLYENKEALIRMFTSKEW-KSGKCAKLRDG 264
++++ + T+ DL+RP R A+ + TL L ++K +L +F S W S AK +G
Sbjct: 504 LNLMKNEFTQGLDLLRPAVMRHASGFTTLQSLMDHKASLRGLFQSDGWILSQTAAKSEEG 563
Query: 265 KALEDIILDKKFWKDIVICLKGATPLMKVLRLAD-----LDKKTSHGIHL*SNGSSKGAN 319
+ +E ++L FWK + LK P+M+V+ + + L ++G + + K +
Sbjct: 564 REVEKMVLSAVFWKKVQYVLKSVDPVMQVIHMINDGGDRLSMPYAYGYMCCAKMAIKSIH 623
Query: 320 SKEF*WCQKEPVWNIIDDRWDRMLHRPLHAAAYYLNPQLHYRPGFKADIEVRKGLMDCIT 379
S + + P W +I+ RW+ + H PL+ AAY+ NP YRP F A EV +G+ +CI
Sbjct: 624 SDDA--RKYGPFWRVIEYRWNPLFHHPLYVAAYFFNPAYKYRPDFMAQSEVVRGVNECIV 681
Query: 380 RMVEDPEEQAKIEVQMHDFKKQIGYFGTKIAKLTLDKSTPADWWESVVFEYLELQRFA 437
R+ D + +Q+ D+ FGT IA T + P+ WW+ LELQR A
Sbjct: 682 RLEPDNTRRITALMQIPDYTCAKADFGTDIAIGTRTELDPSAWWQQHGISCLELQRVA 739
>At3g22220 hypothetical protein
Length = 759
Score = 245 bits (625), Expect = 4e-65
Identities = 141/407 (34%), Positives = 216/407 (52%), Gaps = 5/407 (1%)
Query: 32 DMISRHGNGFKPPSYYDIRVKLLNQEVKSTNDALEEHKREWKKTSCTIMTDEWTDRRRRT 91
D I G G P++ D+R +L V+ ++E K WK+T C+++ E
Sbjct: 218 DAIVSGGFGVSIPTHEDLRGWILKSCVEEVKKEIDECKTLWKRTGCSVLVQELNSNEGPL 277
Query: 92 ILNFLVNSPRGTVFLKSVDASNICKTSEKIFEMIDAVVEEVGEDNVVQVVTDNAANYKAA 151
IL FLV P VFLKSVDAS I + +K++E++ VVEE+G+ NVVQV+T +Y AA
Sbjct: 278 ILKFLVYCPEKVVFLKSVDASEILDSEDKLYELLKEVVEEIGDTNVVQVITKCEDHYAAA 337
Query: 152 GEMLMRKRKKLYWTPCAAHCLDLMLEDLEKKIPIHGETIPKGKKITTFIYARTSLISILH 211
G+ LM LYW PCAAHC+D MLE+ K I E I + + +T IY + +++++
Sbjct: 338 GKKLMDVYPSLYWVPCAAHCIDKMLEEFGKMDWIR-EIIEQARTVTRIIYNHSGVLNLMR 396
Query: 212 SHTKNRDLVRPGATRFATSYLTLGCLYENKEALIRMFTSKEWKSGKCAKLRDGKALEDII 271
T D+V+P T AT++ T+G + + K L M TS EW +K G A+ + I
Sbjct: 397 KFTFGNDIVQPVCTSSATNFTTMGRIADLKPYLQAMVTSSEWNDCSYSKEAGGLAMTETI 456
Query: 272 LDKKFWKDIVICLKGATPLMKVLRLADLDKKTSHGIHL*SNGSSKGANSKEF*WCQKEPV 331
D+ FWK + + P+++VLR+ ++K + G + +K A ++ V
Sbjct: 457 NDEDFWKALTLANHITAPILRVLRIVCSERKPAMGYVYAAMYRAKEAIKTNLAHREEYIV 516
Query: 332 -WNIIDDRWDRMLHRPLHAAAYYLNPQLHYRPGFKADIEVRKGLMDCITRMVEDPEEQAK 390
W IID W L +PL+AA +YLNP+ Y + E+ ++DCI ++V D Q
Sbjct: 517 YWKIIDRWW---LQQPLYAAGFYLNPKFFYSIDEEMRSEIHLAVVDCIEKLVPDVNIQDI 573
Query: 391 IEVQMHDFKKQIGYFGTKIAKLTLDKSTPADWWESVVFEYLELQRFA 437
+ ++ +K +G FG +A D PA+WW + L L RFA
Sbjct: 574 VIKDINSYKNAVGIFGRNLAIRARDTMLPAEWWSTYGESCLNLSRFA 620
>At1g79740 hypothetical protein
Length = 518
Score = 200 bits (508), Expect = 1e-51
Identities = 114/409 (27%), Positives = 212/409 (50%), Gaps = 7/409 (1%)
Query: 30 MCDMISRHGNGFKPPSYYDIRVKLLNQEVKSTNDALEEHKREWKKTSCTIMTDEWTDRRR 89
M D +++ G GF PS + + L++ + L++ ++EW T CTI+ + WTD +
Sbjct: 1 MLDAVAKCGPGFVAPSP---KTEWLDRVKSDISLQLKDTEKEWVTTGCTIIAEAWTDNKS 57
Query: 90 RTILNFLVNSPRGTVFLKSVDASNICKTSEKIFEMIDAVVEEVGEDNVVQVVTDNAANYK 149
R ++NF V+SP F KSVDAS+ K S+ + ++ D+V++++G++++VQ++ DN+ Y
Sbjct: 58 RALINFSVSSPSRIFFHKSVDASSYFKNSKCLADLFDSVIQDIGQEHIVQIIMDNSFCYT 117
Query: 150 AAGEMLMRKRKKLYWTPCAAHCLDLMLEDLEKKIPIHGETIPKGKKITTFIYARTSLISI 209
L++ ++ +PCA+ CL+++LE+ K+ + I + + I+ F+Y + ++ +
Sbjct: 118 GISNHLLQNYATIFVSPCASQCLNIILEEF-SKVDWVNQCISQAQVISKFVYNNSPVLDL 176
Query: 210 LHSHTKNRDLVRPGATRFATSYLTLGCLYENKEALIRMFTSKEWKSGKCAKLRDGKALED 269
L T +D++R G TR +++L+L + + K L MF E+ + + +
Sbjct: 177 LRKLTGGQDIIRSGVTRSVSNFLSLQSMMKQKARLKHMFNCPEYTTN--TNKPQSISCVN 234
Query: 270 IILDKKFWKDIVICLKGATPLMKVLRLADLDKKTSHGIHL*SNGSSKGANSKEF*WCQKE 329
I+ D FW+ + + + P++KVLR K I+ + + + + K
Sbjct: 235 ILEDNDFWRAVEESVAISEPILKVLREVSTGKPAVGSIYELMSKAKESIRTYYIMDENKH 294
Query: 330 PVW-NIIDDRWDRMLHRPLHAAAYYLNPQLHYRPGFKADIEVRKGLMDCITRMVEDPEEQ 388
V+ +I+D W LH PLHAAA +LNP + Y P K +++ + +++ + +
Sbjct: 295 KVFSDIVDTNWCEHLHSPLHAAAAFLNPSIQYNPEIKFLTSLKEDFFKVLEKLLPTSDLR 354
Query: 389 AKIEVQMHDFKKQIGYFGTKIAKLTLDKSTPADWWESVVFEYLELQRFA 437
I Q+ F + G FG +A D +P WWE LQR A
Sbjct: 355 RDITNQIFTFTRAKGMFGCNLAMEARDSVSPGLWWEQFGDSAPVLQRVA 403
>At3g13030 unknown protein (At3g13030)
Length = 544
Score = 199 bits (507), Expect = 2e-51
Identities = 119/402 (29%), Positives = 195/402 (47%), Gaps = 11/402 (2%)
Query: 40 GFKPPSYYDIRVKLLNQEVKSTNDALEEHKREWKKTSCTIMTDEWTDRRRRTILNFLVNS 99
G + +D+ L ++ D +E+ K W T C+I+ D W D++ R ++ F+ +
Sbjct: 84 GLESSDCHDLNGWRLQDALEEVQDRVEKIKESWAITGCSILLDAWVDQKGRDLVTFVADC 143
Query: 100 PRGTVFLKSVDASNICKTSEKIFEMIDAVVEEVGEDNVVQVVTDNAANYKAA-GEMLMRK 158
P G V+L S D S+ + +++ +VEEVG NV Q++ + + + GE+
Sbjct: 144 PAGLVYLISFDVSDFKDDVTALLSLVNGLVEEVGVRNVTQIIACSTSGWVGELGELFAGH 203
Query: 159 RKKLYWTPCAAHCLDLMLEDLEKKIPIHGETIPKGKKITTFIYARTSLISILHSHTKNRD 218
++++W+ +HC +LML + K I G+ K I FI S+++I D
Sbjct: 204 DREVFWSVSVSHCFELMLVKISK-IRSFGDIFDKVNNIWLFINNNPSVLNIFRDQCHGID 262
Query: 219 L-VRPGATRFATSYLTLGCLYENKEALIRMFTSKEWKSGKCAKLRDGKALEDIILDKKFW 277
+ V F T YL L +++ K+ L MF S W + +C A+ +++ D FW
Sbjct: 263 ITVSSSEFEFVTPYLILESIFKAKKNLTAMFASSNWNNEQCI------AISNLVSDSSFW 316
Query: 278 KDIVICLKGATPLMKVLRLADLDKKTSHGIHL*SNGSSKGANSKEF*WCQK--EPVWNII 335
+ + LK +PL+ L L G + S K + ++EF + +P+W++I
Sbjct: 317 ETVESVLKCTSPLIHGLLLFSTANNQHLGYVYDTMDSIKESIAREFNHKPQFYKPLWDVI 376
Query: 336 DDRWDRMLHRPLHAAAYYLNPQLHYRPGFKADIEVRKGLMDCITRMVEDPEEQAKIEVQM 395
DD W++ LH PLHAA Y+LNP Y F DIEV GL+ + MVED Q KI Q+
Sbjct: 377 DDVWNKHLHNPLHAAGYFLNPTAFYSTNFHLDIEVVTGLISSLIHMVEDCHVQFKISTQI 436
Query: 396 HDFKKQIGYFGTKIAKLTLDKSTPADWWESVVFEYLELQRFA 437
++ F + +PA+WW +Y ELQ A
Sbjct: 437 DMYRLGKDCFNEASQADQITGISPAEWWAHKASQYPELQSLA 478
>At3g13020 hypothetical protein
Length = 605
Score = 196 bits (497), Expect = 3e-50
Identities = 122/411 (29%), Positives = 202/411 (48%), Gaps = 14/411 (3%)
Query: 32 DMISRHGNGFKPPSYYDIRVKLLNQEVKSTNDALEEHKREWKKTSCTIMTDEWTDRRRRT 91
+M+ G G K P +D+ +LL + +K D ++ K WK T C+I+ D W D +
Sbjct: 149 EMMMALGVGQKIPDSHDLNGRLLQEAMKEVQDYVKNIKDSWKITGCSILLDAWIDPKGHD 208
Query: 92 ILNFLVNSPRGTVFLKSVDASNICKTSEKIFEMIDAVVEEVGEDNVVQVVTDNAANYKA- 150
+++F+ + P G V+LKS+D S + + +++ +VEEVG NV Q++ + + +
Sbjct: 209 LVSFVADCPAGPVYLKSIDVSVVKNDVTALLSLVNGLVEEVGVHNVTQIIACSTSGWVGE 268
Query: 151 AGEMLMRKRKKLYWTPCAAHCLDLMLEDLEKKIPIHGETIPKGKKITTFIYARTSLISIL 210
G++ ++++W+ +HC +LML + K+ G+ + K I FI S + I
Sbjct: 269 LGKLFSGHDREVFWSVSLSHCFELMLVKI-GKMRSFGDILDKVNTIWEFINNNPSALKIY 327
Query: 211 HSHTKNRDL-VRPGATRFATSYLTLGCLYENKEALIRMFTSKEWKSGKCAKLRDGKALED 269
+ +D+ V F YL L +++ K+ L MF S WK +GK++ +
Sbjct: 328 RDQSHGKDITVSSSEFEFVKPYLILKSVFKAKKNLAAMFASSVWKK------EEGKSVSN 381
Query: 270 IILDKKFWKDIVICLKGATPLMKVLRL-ADLDKKTSHGIHL*SNGSSKGANSKEF*WCQK 328
++ D FW+ + LK +PL LRL ++ D G + K + KEF +K
Sbjct: 382 LVNDSSFWEAVEEILKCTSPLTDGLRLFSNADNNQHVGYIYDTLDGIKLSIKKEFNDEKK 441
Query: 329 E--PVWNIIDDRWDRMLHRPLHAAAYYLNPQLHYRPGFKADIEVRKGLMDCITRMVEDPE 386
+W++IDD W++ LH PLHAA YYLNP Y F D EV GL + + + E
Sbjct: 442 HYLTLWDVIDDVWNKHLHNPLHAAGYYLNPTSFYSTDFHLDPEVSSGLTHSLVHVAK--E 499
Query: 387 EQAKIEVQMHDFKKQIGYFGTKIAKLTLDKSTPADWWESVVFEYLELQRFA 437
Q KI Q+ ++ F + +P DWW ++ ELQ FA
Sbjct: 500 GQIKIASQLDRYRLGKDCFNEASQPDQISGISPIDWWTEKASQHPELQSFA 550
>At4g15020 putative protein
Length = 522
Score = 167 bits (424), Expect = 8e-42
Identities = 112/340 (32%), Positives = 168/340 (48%), Gaps = 25/340 (7%)
Query: 24 SKEFNIMCDMISRHGNGFKPPSYYDIRVKLLNQEVKSTNDALEEHKREWKKTSCTIMTDE 83
S F M D I+ G G P++ D+R +L V+ ++E K WK+T C+I+ +E
Sbjct: 198 SVNFQPMIDAIASGGFGVSAPTHDDLRGWILKNCVEEMAKEIDECKAMWKRTGCSILVEE 257
Query: 84 WTDRRRRTILNFLVNSPRGTVFLKSVDASNICKTSEKIFEMIDAVVEEVGEDNVVQVVTD 143
+ +LNFLV P VFLKSVDAS + +++K+FE++ +VEEVG NVVQV+T
Sbjct: 258 LNSDKGFKVLNFLVYCPEKVVFLKSVDASEVLSSADKLFELLSELVEEVGSTNVVQVITK 317
Query: 144 NAANYKAAGEMLMRKRKKLYWTPCAAHCLDLMLEDLEKKIPIHGETIPKGKKITTFIYAR 203
Y AG+ LM LYW PCAAHC+D MLE+ K+ ETI + + IT F+Y
Sbjct: 318 CDDYYVDAGKRLMLVYPSLYWVPCAAHCIDQMLEEF-GKLGWISETIEQAQAITRFVYNH 376
Query: 204 TSLISILHSHTKNRDLVRPGATRFATSYLTLGCLYENKEALIRMFTSKEWKSGKCAKLRD 263
++ S AT++ TLG + E K L M TS EW ++
Sbjct: 377 SAFSS------------------SATNFATLGRIAELKSNLQAMVTSAEWNECSYSEEPS 418
Query: 264 GKALEDIILDKKFWKDIVICLKGATPLMKVLRLADLDKKTSHGIHL*SNGSSKGANSKEF 323
G + + + D+ FWK + + +PL++ LR+ +K+ + G + +K A
Sbjct: 419 GLVM-NALTDEAFWKAVALVNHLTSPLLRALRIVCSEKRPAMGYVYAALYRAKDAIKTHL 477
Query: 324 *WCQKEP---VWNIIDDRWDRMLHRPLHAAAYYLNPQLHY 360
+E W IID ++LNP+L Y
Sbjct: 478 --VNREDYIIYWKIIDHGGSNNSIFLCLLLGFFLNPKLFY 515
>At1g43260 hypothetical protein
Length = 294
Score = 167 bits (422), Expect = 1e-41
Identities = 85/216 (39%), Positives = 128/216 (58%), Gaps = 2/216 (0%)
Query: 30 MCDMISRHGNGFKPPSYYDIRVKLLNQEVKSTNDALEEHKREWKKTSCTIMTDEWTDRRR 89
M ++ + G G PPS Y +R LL +EV +EE + EW+ C++ TD W+DR+R
Sbjct: 61 MLEVAGQFGPGVTPPSQYQLREPLLKEEVVRMKGLMEEQEDEWRVNGCSVTTDSWSDRKR 120
Query: 90 RTILNFLVNSPRGTVFLKSVDASNICKTSEKIFEMI-DAVVEEVGEDNVVQVVTDNAANY 148
R+I+N +N GT+FL S D + T E IF + + ++ +G D+VVQVVT+NA N
Sbjct: 121 RSIMNLCINCKEGTMFLSSKDCFDDSHTGEYIFAYVNEYCIKNLGGDHVVQVVTNNATNN 180
Query: 149 KAAGEMLMRKRKKLYWTPCAAHCLDLMLEDLEKKIPIHGETIPKGKKITTFIYARTSLIS 208
A ++L R ++WT CA H ++LM+E + K+ + E + K T FIYA +S
Sbjct: 181 ITAAKLLKEVRPTIFWTFCATHTINLMVEGI-SKLAMSDEIVKMAKAFTIFIYAHHQTLS 239
Query: 209 ILHSHTKNRDLVRPGATRFATSYLTLGCLYENKEAL 244
++ S TK RD+VRPG FA+++ TL L E +E L
Sbjct: 240 MMRSFTKRRDIVRPGIIGFASAFRTLKSLVEKEENL 275
>At3g13010 hypothetical protein
Length = 572
Score = 164 bits (415), Expect = 9e-41
Identities = 109/410 (26%), Positives = 186/410 (44%), Gaps = 15/410 (3%)
Query: 29 IMCDMISRHGNGFKPPSYYDIRVKLLNQEVKSTNDALEEHKREWKKTSCTIMTDEWTDRR 88
+M + G G P D+ + + +K D ++E K W+ T C+I+ D W +
Sbjct: 122 MMIKTVGGEGGGQMIPDSRDLNGWMFQEALKKVQDRVKEIKASWEITGCSILFDAWIGPK 181
Query: 89 RRTILNFLVNSPRGTVFLKSVDASNICKTSEKIFEMIDAVVEEVGEDNVVQVVTDNAANY 148
R ++ F+ + P G V+LKS D S+I + +++ +VEEVG NV Q++ + + +
Sbjct: 182 GRDLVTFVADCPAGAVYLKSADVSDIKTDVTALTSLVNGIVEEVGVRNVTQIIACSTSGW 241
Query: 149 KAAGEMLMRKRKKLYWTPCAAHCLDLMLEDLEKKIPIHGETIPKGKKITTFIYARTSLIS 208
G++ + +++W+ ++CL LML ++ K + K K + I S +
Sbjct: 242 --VGDLGKQLAGQVFWSVSLSYCLKLMLVEIGKMYSFE-DIFEKVKLLLDLINNNPSFLY 298
Query: 209 ILHSHTKNRDLVRPGATRFATSYLTLGCLYENKEALIRMFTSKEWKSGKCAKLRDGKALE 268
+ ++ D+ F YLTL +Y + A +F S EWK G A+
Sbjct: 299 VFRENSHKVDV--SSECEFVMPYLTLEHIYWVRRA--GLFASPEWKK------EQGIAIS 348
Query: 269 DIILDKKFWKDIVICLKGATPLMK-VLRLADLDKKTSHGIHL*SNGSSKGANSKEF*WCQ 327
+ D FW+ + + + L+ L + K ++ H + A + ++
Sbjct: 349 SFVNDSTFWESLDKIVGSTSSLVHGWLWFSRGSKHVAYAYHFIESIKKNVAWTFKYERQF 408
Query: 328 KEPVWNIIDDRWDRMLHRPLHAAAYYLNPQLHYRPGFKADIEVRKGLMDCITRMVEDPEE 387
EP WN+IDD W H PLHAA Y+LNP +Y F V GL + +V++P
Sbjct: 409 YEPTWNVIDDVWHNN-HNPLHAAGYFLNPMAYYSDDFHTYQHVYTGLAFSLVHLVKEPHL 467
Query: 388 QAKIEVQMHDFKKQIGYFGTKIAKLTLDKSTPADWWESVVFEYLELQRFA 437
Q KI Q+ ++ G F L+ +P +WW +Y ELQ A
Sbjct: 468 QVKIGTQLDVYRYGRGCFMKASQAGQLNGVSPVNWWTQKANQYPELQNLA 517
>At1g41920 hypothetical protein
Length = 496
Score = 31.2 bits (69), Expect = 1.2
Identities = 47/213 (22%), Positives = 74/213 (34%), Gaps = 37/213 (17%)
Query: 154 MLMRKRKKLYWTPCAAHCLDLMLEDLEKKIPIHGETIP-------------KGKKITTFI 200
+++R+ Y+ C AH L L++ + KK G+ KGK
Sbjct: 175 LILRENSSAYYIHCFAHQLQLVVVAVAKKQFEVGDFFDMISALLNVVGASCKGKDRIREE 234
Query: 201 YARTSLISILHSHTKNR-------DLVRPGATRFATSYLTLGCLYENKEALIRMFTSKEW 253
Y + I K L RPG TR+ T Y TL L +I++ E
Sbjct: 235 YLKEIEEGINQGEIKTGKGLNQELSLQRPGNTRWGTHYTTLHRLAHLFSVIIKLLEFVEE 294
Query: 254 KSGKCAKLRDGKALEDII--LDKKFWKDIVICLKGATPLMKVLRLADLDKKTSHGIHL*S 311
+ K R L D F+ +++ L G T + V L KK +
Sbjct: 295 EGTDSTKRRQANGLLKYFNTFDFAFYLQLMLLLLGLTNSLSVA----LQKKDQDIL---- 346
Query: 312 NGSSKGANSKEF*WCQKEPVWNIIDDRWDRMLH 344
N K+ + + DD WD +++
Sbjct: 347 -------NPMSLVKSTKQQLCKLRDDGWDSLVN 372
>At2g14790 Mutator-like transposase
Length = 895
Score = 30.4 bits (67), Expect = 2.0
Identities = 26/92 (28%), Positives = 41/92 (44%), Gaps = 7/92 (7%)
Query: 146 ANYKAAGEMLMRKRKKLYWTPCAAHCLDLMLEDLEKKIPIHGETIPKGKKITTFIY--AR 203
ANYK GE++ + P A L+L DL K+ G+ I + + T++Y
Sbjct: 386 ANYKLLGEVVRSRYSSTQGGPRAVDLPQLLLNDLNWKM-YGGDEIANYRFLPTYLYLLQL 444
Query: 204 TSLISILHSHTKNRDLVRPGATRFATSYLTLG 235
+ +I H H D G RF +++LG
Sbjct: 445 ANPGTITHLHYTEDD----GKQRFKYVFVSLG 472
>At4g00620 putative tetrahydrofolate synthase
Length = 360
Score = 30.0 bits (66), Expect = 2.6
Identities = 20/70 (28%), Positives = 34/70 (48%), Gaps = 6/70 (8%)
Query: 152 GEMLMRKRKKLYWTPCAAH-CLDLM----LEDLEKKIPIHGETIPKGKKITTFIYARTSL 206
G + MR R+ L+ PC C++L+ +E K+ + G + G + +
Sbjct: 195 GRLAMRGREPLF-VPCTPKGCIELLHRYNIEIKGKRAVVIGRSNIVGMPAALLLQREDAT 253
Query: 207 ISILHSHTKN 216
+SI+HS TKN
Sbjct: 254 VSIIHSRTKN 263
>At1g27180 disease resistance protein, putative
Length = 1556
Score = 30.0 bits (66), Expect = 2.6
Identities = 23/78 (29%), Positives = 38/78 (48%), Gaps = 2/78 (2%)
Query: 82 DEWTDRRRRTILNFLVNSPRGTVFLKSVDASNICKTSEKIFEMIDAVVEEV--GEDNVVQ 139
DE T +R + +N + N P K+V N EK+ +MID VV++V N +
Sbjct: 297 DEETIQRWKRAMNLVGNIPGYVCTAKTVGDDNEGINREKVDDMIDLVVKKVVAAVRNRPE 356
Query: 140 VVTDNAANYKAAGEMLMR 157
+V D ++ + LM+
Sbjct: 357 IVADYTVGLESPIKDLMK 374
>At1g29990 hydrophilic like protein
Length = 129
Score = 29.3 bits (64), Expect = 4.5
Identities = 25/99 (25%), Positives = 47/99 (47%), Gaps = 13/99 (13%)
Query: 107 KSVDASNICKTSEKIFEMIDAVVEEVGEDNVV----QVVTDNAANYKAAGEMLMRK---- 158
K+ D I K K ++ ++GE+ +V ++ ++A YK G +L+++
Sbjct: 16 KANDLGKIQKDIGKNHQLRKKYTIQLGENELVLKELDLLEEDANVYKLIGPVLVKQDLAE 75
Query: 159 -----RKKLYWTPCAAHCLDLMLEDLEKKIPIHGETIPK 192
RK++ + LD +L+D+E+K ETI K
Sbjct: 76 ANANVRKRIEYISAELKRLDAILQDMEEKQNNKRETIMK 114
>At1g14530 unknown protein
Length = 293
Score = 29.3 bits (64), Expect = 4.5
Identities = 14/31 (45%), Positives = 18/31 (57%)
Query: 84 WTDRRRRTILNFLVNSPRGTVFLKSVDASNI 114
WT ++ LNF+VN R VFL DA N+
Sbjct: 70 WTTQKVFHFLNFMVNGVRALVFLFRRDAQNM 100
>At3g19250 hypothetical protein
Length = 398
Score = 28.9 bits (63), Expect = 5.8
Identities = 26/96 (27%), Positives = 47/96 (48%), Gaps = 20/96 (20%)
Query: 39 NGFKPPSYYDIRVKL-------------LNQEVKSTNDALEE--HKREWKKTSCTIMTDE 83
+ F+ PSY+DIR +L L+QE++ N++++E R K+T+ T +
Sbjct: 80 HAFQTPSYHDIRSRLLVIDPTQENLELFLSQELRPNNESVQEALSLRHAKQTTLTNLVST 139
Query: 84 W---TDRRRRTILNFL--VNSPRGTVFLKSVDASNI 114
+ ++ R LN V+S R ++ +D NI
Sbjct: 140 YFQHSEDATRFCLNLYQNVHSARCHLYTPLLDLFNI 175
>At3g07980 putative MAP3K epsilon protein kinase
Length = 1367
Score = 28.9 bits (63), Expect = 5.8
Identities = 24/84 (28%), Positives = 38/84 (44%), Gaps = 8/84 (9%)
Query: 78 TIMTDEWTDRRRRTILNFLVNSPRGTV-FLKSVDASNICKTSEKIFEMIDAVVEEVGEDN 136
T+++ W RR + + L +S GT+ ++K D+S SEK E VVE V +
Sbjct: 267 TLLSHPWIRNSRRALRSSLRHS--GTIRYMKETDSS-----SEKDAEGSQEVVESVSAEK 319
Query: 137 VVQVVTDNAANYKAAGEMLMRKRK 160
V T++ + G R K
Sbjct: 320 VEVTKTNSKSKLPVIGGASFRSEK 343
>At1g25784 mutator-like transposase, putative
Length = 1127
Score = 28.9 bits (63), Expect = 5.8
Identities = 28/108 (25%), Positives = 45/108 (40%), Gaps = 23/108 (21%)
Query: 146 ANYKAAGEMLMRKRKKLYWTPCAAHCLDLMLEDLEKKIP--------------IHGETIP 191
ANYK GE++ + P A L+L+DL +I + G+ I
Sbjct: 461 ANYKLLGEVVRSRYSSTQGGPRAVDLPQLLLKDLNVRISYSTAWRAKEVAVENVRGDEIA 520
Query: 192 KGKKITTFIY----ARTSLISILHSHTKNRDLVRPGATRFATSYLTLG 235
+ + T++Y A I+ LH +T D G RF +++LG
Sbjct: 521 NYRFLPTYLYLLQLANPGTITHLH-YTPEDD----GKQRFKYVFVSLG 563
>At4g10200 putative protein
Length = 733
Score = 28.5 bits (62), Expect = 7.6
Identities = 21/76 (27%), Positives = 37/76 (48%), Gaps = 8/76 (10%)
Query: 105 FLKSVDASNICKTSEKIFEMI--DAVVEEVGEDNVVQVVTDNAANYKAAGEMLMRK---- 158
FL V+ S+ K+ E +FE++ V + ++V DN N K + + +K
Sbjct: 319 FLTFVEVSD--KSGEGLFELLCDTLVALNLNINDVRGQGYDNGCNMKGKHKGVQKKLLDI 376
Query: 159 RKKLYWTPCAAHCLDL 174
+ ++TPC H L+L
Sbjct: 377 NSRAFYTPCGCHSLNL 392
>At4g08890 putative transposon protein
Length = 1028
Score = 28.5 bits (62), Expect = 7.6
Identities = 28/108 (25%), Positives = 44/108 (39%), Gaps = 23/108 (21%)
Query: 146 ANYKAAGEMLMRKRKKLYWTPCAAHCLDLMLEDLEKKIP--------------IHGETIP 191
ANYK GE++ + P A L+L DL +I + G+ I
Sbjct: 393 ANYKLLGEVVRSRYSSTQGGPQAVDLPQLLLNDLNVRISYSTAWRAKEVAVENVRGDEIA 452
Query: 192 KGKKITTFIY----ARTSLISILHSHTKNRDLVRPGATRFATSYLTLG 235
+ + T++Y A I+ LH +T D G RF +++LG
Sbjct: 453 NYRFLPTYLYLLQLANPGTITHLH-YTPEDD----GKQRFKYVFVSLG 495
>At3g43360 putative transposase
Length = 675
Score = 28.5 bits (62), Expect = 7.6
Identities = 28/108 (25%), Positives = 44/108 (39%), Gaps = 23/108 (21%)
Query: 146 ANYKAAGEMLMRKRKKLYWTPCAAHCLDLMLEDLEKKIP--------------IHGETIP 191
ANYK GE++ + P A L+L DL +I + G+ I
Sbjct: 461 ANYKLLGEVVRSRYSSTQGGPRAVDLPQLLLNDLNVRISYSTAWRAKEVVVENVRGDEIA 520
Query: 192 KGKKITTFIY----ARTSLISILHSHTKNRDLVRPGATRFATSYLTLG 235
+ + T++Y A I+ LH +T D G RF +++LG
Sbjct: 521 NYRFLPTYLYLLQLANPGTITHLH-YTPEDD----GKQRFKYVFVSLG 563
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.325 0.138 0.432
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,836,976
Number of Sequences: 26719
Number of extensions: 419426
Number of successful extensions: 1459
Number of sequences better than 10.0: 23
Number of HSP's better than 10.0 without gapping: 11
Number of HSP's successfully gapped in prelim test: 12
Number of HSP's that attempted gapping in prelim test: 1417
Number of HSP's gapped (non-prelim): 28
length of query: 437
length of database: 11,318,596
effective HSP length: 102
effective length of query: 335
effective length of database: 8,593,258
effective search space: 2878741430
effective search space used: 2878741430
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 61 (28.1 bits)
Medicago: description of AC147430.8


