

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147429.11 - phase: 0
(149 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
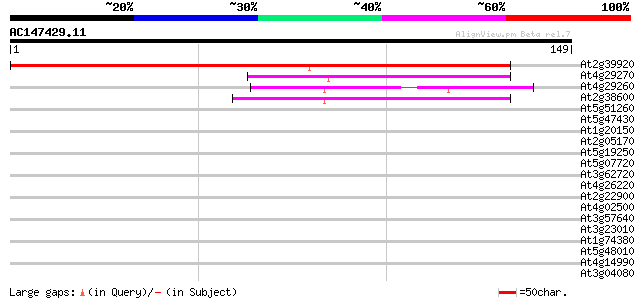
Score E
Sequences producing significant alignments: (bits) Value
At2g39920 unknown protein 130 2e-31
At4g29270 acid phosphatase-like protein 49 1e-06
At4g29260 acid phosphatase-like protein 41 2e-04
At2g38600 putative acid phosphatase 40 5e-04
At5g51260 acid phosphatase 38 0.002
At5g47430 DNA-binding protein-like 28 1.8
At1g20150 unknown protein 28 1.8
At2g05170 unknown protein 28 2.4
At5g19250 unknown protein 27 3.1
At5g07720 alpha galactosyltransferase protein 27 3.1
At3g62720 alpha galactosyltransferase-like protein 27 3.1
At4g26220 27 4.1
At2g22900 unknown protein 27 4.1
At4g02500 unknown protein 27 5.3
At3g57640 putative protein 27 5.3
At3g23010 disease resistance protein, putative 27 5.3
At1g74380 putative alpha galactosyltransferase 27 5.3
At5g48010 cycloartenol synthase 26 6.9
At4g14990 unknown protein 26 6.9
At3g04080 apyrase (Atapy1) 26 6.9
>At2g39920 unknown protein
Length = 283
Score = 130 bits (328), Expect = 2e-31
Identities = 63/134 (47%), Positives = 94/134 (70%), Gaps = 1/134 (0%)
Query: 1 MGSSFVLESGFYITSFSATIFIAGFAALGLLLVTLLVSMAMMLQSCQNSNGGIIELRNIN 60
+GS + +ESG Y+TS +A+IFIA G+L++TLL++++ MLQSC+N N I+E + ++
Sbjct: 23 LGSRYSIESGCYMTSLAASIFIASLVTFGVLMITLLIALSTMLQSCENRNIAIVEAQRLD 82
Query: 61 NDYSYCKIHSLHAKLNNL-EEHNVPNICKDLALQYIKGGQYARDLDSTKSVIEDYFNGVK 119
+ YCKI SLH++LN+L EE +P +C+D+AL IK G Y R+L+ T + YF +K
Sbjct: 83 ESFGYCKILSLHSQLNSLDEESELPLLCRDVALHRIKQGIYLRELNFTIQMALTYFQTIK 142
Query: 120 PSEDGFDVVLIDID 133
P D DVV+IDID
Sbjct: 143 PMNDNCDVVVIDID 156
>At4g29270 acid phosphatase-like protein
Length = 256
Score = 48.9 bits (115), Expect = 1e-06
Identities = 28/71 (39%), Positives = 37/71 (51%), Gaps = 1/71 (1%)
Query: 64 SYCKIHSLHAKLNNLEEHNV-PNICKDLALQYIKGGQYARDLDSTKSVIEDYFNGVKPSE 122
SYC+ L A+ NN+ V P+ C++ YI GGQ+ +D D S DY VK
Sbjct: 42 SYCESWRLAAETNNVGPWKVIPSQCENYIKNYINGGQFDKDYDVVASYAIDYAKTVKVGG 101
Query: 123 DGFDVVLIDID 133
DG D + DID
Sbjct: 102 DGKDAWVFDID 112
>At4g29260 acid phosphatase-like protein
Length = 255
Score = 41.2 bits (95), Expect = 2e-04
Identities = 29/80 (36%), Positives = 39/80 (48%), Gaps = 9/80 (11%)
Query: 65 YCKIHSLHAKLNNLEEHN-VPNICKDLALQYIKGGQYARDLDSTKSVIEDYF----NGVK 119
YC L A+ NN+ + +P+IC D +Y+ G Q+ D SVI DY V+
Sbjct: 42 YCDSWRLAAETNNVGTWDLIPSICVDSVAEYLNGDQFLSDY----SVIVDYALAFAKSVE 97
Query: 120 PSEDGFDVVLIDIDSLFQWN 139
S DG DV + DID N
Sbjct: 98 ISGDGKDVWIFDIDETLLTN 117
>At2g38600 putative acid phosphatase
Length = 251
Score = 40.0 bits (92), Expect = 5e-04
Identities = 25/75 (33%), Positives = 34/75 (45%), Gaps = 1/75 (1%)
Query: 60 NNDYSYCKIHSLHAKLNNLEEHN-VPNICKDLALQYIKGGQYARDLDSTKSVIEDYFNGV 118
N SYC L + NN+ VP C Y+ GQY RD+ T I+ Y N +
Sbjct: 32 NYGASYCLSWRLAVETNNVRAWRIVPLQCLRYVEVYMLAGQYDRDVQLTVDQIKVYLNEI 91
Query: 119 KPSEDGFDVVLIDID 133
DG D ++D+D
Sbjct: 92 ILPGDGMDAWILDVD 106
>At5g51260 acid phosphatase
Length = 257
Score = 38.1 bits (87), Expect = 0.002
Identities = 22/79 (27%), Positives = 34/79 (42%), Gaps = 1/79 (1%)
Query: 65 YCKIHSLHAKLNNLEE-HNVPNICKDLALQYIKGGQYARDLDSTKSVIEDYFNGVKPSED 123
+C A++NNL +P C D Y+ G Y DL+ + ++ S D
Sbjct: 44 HCTTWRFAAEMNNLAPWKTIPVECADYVKDYVMGKGYLTDLERVSEEALIFARSIEFSGD 103
Query: 124 GFDVVLIDIDSLFQWNPPH 142
G D+ + DID N P+
Sbjct: 104 GKDIWIFDIDETLLSNLPY 122
>At5g47430 DNA-binding protein-like
Length = 889
Score = 28.1 bits (61), Expect = 1.8
Identities = 15/51 (29%), Positives = 28/51 (54%), Gaps = 1/51 (1%)
Query: 76 NNLEEHNVPNIC-KDLALQYIKGGQYARDLDSTKSVIEDYFNGVKPSEDGF 125
N+++ + VP++ D +Q G QY + + ++ FNGV+P +GF
Sbjct: 485 NDMQWNPVPDLAGPDYMMQMGPGPQYFNGMQPGFNGVQPGFNGVQPGFNGF 535
>At1g20150 unknown protein
Length = 780
Score = 28.1 bits (61), Expect = 1.8
Identities = 25/83 (30%), Positives = 37/83 (44%), Gaps = 2/83 (2%)
Query: 54 IELRNINNDYSYCKIHSLHAKLNNLEEHNVPNICKD-LALQYIKGGQYARDLDSTKSVIE 112
I + NI+ +Y IH+ AK + E N D L +KG D D VI+
Sbjct: 363 INIANIDKTQAYPLIHARSAKKIDANEEAARNCAPDTLDQTIVKGKIVVCDSDLDNQVIQ 422
Query: 113 DYFNGVKPSEDGFDVVLIDIDSL 135
+ VK G +VL+D +S+
Sbjct: 423 WKSDEVK-RLGGIGMVLVDDESM 444
>At2g05170 unknown protein
Length = 932
Score = 27.7 bits (60), Expect = 2.4
Identities = 17/61 (27%), Positives = 27/61 (43%), Gaps = 4/61 (6%)
Query: 68 IHSLHAKLNNLEEHNVPNICKDLALQYIKGGQYARDLDSTKSVIEDYFNGVKPSEDGFDV 127
+HS H + E C + A +Y + R L+ + +F VK S+DGF V
Sbjct: 859 MHSFHQRCLGDNEKE----CPECAPEYRSVMEMKRSLEQNSKDQDLFFQQVKGSKDGFSV 914
Query: 128 V 128
+
Sbjct: 915 I 915
>At5g19250 unknown protein
Length = 196
Score = 27.3 bits (59), Expect = 3.1
Identities = 24/94 (25%), Positives = 41/94 (43%), Gaps = 13/94 (13%)
Query: 44 QSCQNSNGGIIE------LRNINNDYSYCKIHSLHAKLNNLEEHNVPNICKDLALQYIKG 97
Q C N+ G N+ N S C+++ + + VPN+ L L
Sbjct: 69 QPCTNTTGSASVPGTTPGFPNLPNLLSKCRLNPTVTRDGAILPACVPNLDPSLVLTNFTM 128
Query: 98 GQYARDLDSTKSVIEDYFNGVK-PSEDGFDVVLI 130
QY++DL+ +K F G+ S+D + VV++
Sbjct: 129 SQYSKDLNDSK------FTGIGIGSDDNWIVVVL 156
>At5g07720 alpha galactosyltransferase protein
Length = 457
Score = 27.3 bits (59), Expect = 3.1
Identities = 14/51 (27%), Positives = 25/51 (48%), Gaps = 6/51 (11%)
Query: 46 CQNSNGGIIELRNINNDYSYCKIHSLHAKLNNLEEHNVPNICKDLALQYIK 96
C N G L+++ N YC+IH + +N+ ++ K+LA + K
Sbjct: 162 CDNPIGDHYLLKSVKNKIDYCRIHGIEI------VYNMAHLDKELAGYWAK 206
>At3g62720 alpha galactosyltransferase-like protein
Length = 460
Score = 27.3 bits (59), Expect = 3.1
Identities = 11/31 (35%), Positives = 16/31 (51%)
Query: 46 CQNSNGGIIELRNINNDYSYCKIHSLHAKLN 76
C+N G L++I N YC+IH + N
Sbjct: 162 CENPVGDHYLLKSIKNKIDYCRIHGIEIFYN 192
>At4g26220
Length = 232
Score = 26.9 bits (58), Expect = 4.1
Identities = 16/55 (29%), Positives = 25/55 (45%), Gaps = 5/55 (9%)
Query: 89 DLALQYIKGGQYARDLDSTKS----VIEDYFNGVKPSEDGFDVVLIDIDSLFQWN 139
++ L IK +D +S +++ N K +E GFD +D D L WN
Sbjct: 104 EIGLPVIKKAGVEHKIDFKESEALPALDELLNN-KVNEGGFDFAFVDADKLNYWN 157
>At2g22900 unknown protein
Length = 449
Score = 26.9 bits (58), Expect = 4.1
Identities = 14/41 (34%), Positives = 19/41 (46%), Gaps = 9/41 (21%)
Query: 46 CQNSNGGIIELRNINNDYSYCKIHS---------LHAKLNN 77
C+N G + LR N YC+IH LH K+N+
Sbjct: 136 CKNPIGDHLLLRFFKNKVDYCRIHGHDIFYSNALLHPKMNS 176
>At4g02500 unknown protein
Length = 461
Score = 26.6 bits (57), Expect = 5.3
Identities = 10/31 (32%), Positives = 16/31 (51%)
Query: 46 CQNSNGGIIELRNINNDYSYCKIHSLHAKLN 76
C+N G L++I N YC++H + N
Sbjct: 163 CENPVGDHYLLKSIKNKIDYCRLHGIEIFYN 193
>At3g57640 putative protein
Length = 356
Score = 26.6 bits (57), Expect = 5.3
Identities = 12/39 (30%), Positives = 19/39 (47%)
Query: 60 NNDYSYCKIHSLHAKLNNLEEHNVPNICKDLALQYIKGG 98
NN + + K+ + EHN N C+D+A+ I G
Sbjct: 82 NNHHHHHKLLLIKKWRYRFSEHNGGNFCRDIAISSIVSG 120
>At3g23010 disease resistance protein, putative
Length = 595
Score = 26.6 bits (57), Expect = 5.3
Identities = 24/83 (28%), Positives = 39/83 (46%), Gaps = 13/83 (15%)
Query: 28 LGLLLVTLLVSMAMMLQSCQNSNGGIIELRNINNDYSYCKIHSLHAKLNNLEEHNVPNIC 87
L LL++ LV + + QN G I+ RN +S ++ L+ NNL+ +I
Sbjct: 85 LSLLMIPSLVHIDLS----QNHFEGPIDFRNT---FSLSRLRVLYVGFNNLDGLIPESIS 137
Query: 88 KDLALQYIK------GGQYARDL 104
K + L+Y+ GGQ R +
Sbjct: 138 KLVNLEYLDVSHNNFGGQVPRSI 160
>At1g74380 putative alpha galactosyltransferase
Length = 457
Score = 26.6 bits (57), Expect = 5.3
Identities = 13/51 (25%), Positives = 25/51 (48%), Gaps = 6/51 (11%)
Query: 46 CQNSNGGIIELRNINNDYSYCKIHSLHAKLNNLEEHNVPNICKDLALQYIK 96
C N G L+++ N YC++H + +N+ ++ K+LA + K
Sbjct: 163 CDNPIGDHYLLKSVKNKIDYCRLHGIEI------VYNMAHLDKELAGYWAK 207
>At5g48010 cycloartenol synthase
Length = 769
Score = 26.2 bits (56), Expect = 6.9
Identities = 13/46 (28%), Positives = 26/46 (56%), Gaps = 3/46 (6%)
Query: 70 SLHAKLNNLEEHNVPNICKDLALQYIKGGQYARDLDSTKSVIEDYF 115
SLHA L+ +++H+V + ++ +KG Y + T++ D+F
Sbjct: 428 SLHALLDGIDDHDVDD---EIKTTLVKGYDYLKKSQITENPRGDHF 470
>At4g14990 unknown protein
Length = 787
Score = 26.2 bits (56), Expect = 6.9
Identities = 21/74 (28%), Positives = 33/74 (44%), Gaps = 15/74 (20%)
Query: 83 VPNICKDLALQYI-----KGGQYARD--LDSTKSVIEDYFNGVKPSEDGFDVVLIDIDSL 135
+P+IC+ AL + G ++ D L + + IED F G + ++DID
Sbjct: 456 LPSICRPRALLEVDSPPSSGHKHLEDEPLVAARVTIEDAF--------GVLIDIVDIDRT 507
Query: 136 FQWNPPHSSNLLLR 149
Q+N P LR
Sbjct: 508 LQFNRPQDGGAQLR 521
>At3g04080 apyrase (Atapy1)
Length = 471
Score = 26.2 bits (56), Expect = 6.9
Identities = 10/22 (45%), Positives = 14/22 (63%)
Query: 73 AKLNNLEEHNVPNICKDLALQY 94
+K +EE N+P +C DL QY
Sbjct: 406 SKFPRVEEDNLPYLCLDLVYQY 427
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.321 0.138 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,447,145
Number of Sequences: 26719
Number of extensions: 139594
Number of successful extensions: 382
Number of sequences better than 10.0: 25
Number of HSP's better than 10.0 without gapping: 13
Number of HSP's successfully gapped in prelim test: 12
Number of HSP's that attempted gapping in prelim test: 368
Number of HSP's gapped (non-prelim): 25
length of query: 149
length of database: 11,318,596
effective HSP length: 90
effective length of query: 59
effective length of database: 8,913,886
effective search space: 525919274
effective search space used: 525919274
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 55 (25.8 bits)
Medicago: description of AC147429.11


