

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147406.4 - phase: 0
(310 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
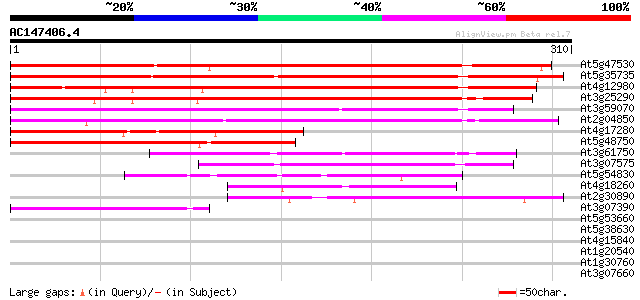
Score E
Sequences producing significant alignments: (bits) Value
At5g47530 unknown protein 311 2e-85
At5g35735 unknown protein 290 9e-79
At4g12980 unknown protein 254 5e-68
At3g25290 unknown protein 249 1e-66
At3g59070 putative protein 211 5e-55
At2g04850 unknown protein 206 1e-53
At4g17280 unknown protein 164 7e-41
At5g48750 putative protein 126 2e-29
At3g61750 putative protein 117 6e-27
At3g07575 unknown protein 116 1e-26
At5g54830 unknown protein 69 3e-12
At4g18260 unknown protein 59 4e-09
At2g30890 hypothetical protein 55 5e-08
At3g07390 auxin-induced protein (AIR12) 44 1e-04
At5g53660 putative protein 33 0.26
At5g38630 cytochrome b-561 (gb|AAD45585.1) 30 1.3
At4g15840 unknown protein 30 1.7
At1g20540 unknown protein 30 2.2
At1g30760 putative reticuline oxidase-like protein, 3' partial 29 3.7
At3g07660 unknown protein 28 4.9
>At5g47530 unknown protein
Length = 395
Score = 311 bits (798), Expect = 2e-85
Identities = 163/305 (53%), Positives = 211/305 (68%), Gaps = 12/305 (3%)
Query: 1 MIGAQALVAIPQASGSPKAYTSNIVDTSTRLQEGTISYPVSGLSATYQNNKVTIFATLTL 60
M+GAQA+VA PQ+ G+ +AYTS I T L E +S+ VS LSATYQNN++ I+A L L
Sbjct: 88 MVGAQAIVAYPQSDGTVRAYTSPISSYQTSLLEAELSFNVSQLSATYQNNEMVIYAILNL 147
Query: 61 PNGTTSLVH-VWQDGVLSSDSTPQEHSHESSHQNSKEVLDLVSGTSQAAS---GIGSRQR 116
P +++ VWQDG LS ++ P H ++ S L+LVSG S + S G S+ R
Sbjct: 148 PLANGGIINTVWQDGSLSGNN-PLPHPTSGNNVRSVSTLNLVSGASGSTSTGAGGASKLR 206
Query: 117 RRNTHGVLNAISWGILMPTGAVIARYLKVFKSADPAWFYLHITCQVSAYIVGLSGFGTGL 176
+RN HG+LN +SWGI+MP GA+IARYLKV KSADPAWFYLH+ CQ SAYI+G++G+ TGL
Sbjct: 207 KRNIHGILNGVSWGIMMPIGAIIARYLKVSKSADPAWFYLHVFCQSSAYIIGVAGWATGL 266
Query: 177 KLGSDSEGITYDTHRALAIVLVTLATLQVFALFLRPNKDHKLRFYWNIYHHVVGYVTISI 236
KLG++S GI + HRA+ I L LAT+QVFA+FLRP +HK R YWNIYHH VGY I +
Sbjct: 267 KLGNESAGIQFTFHRAVGIALFCLATIQVFAMFLRPKPEHKYRVYWNIYHHTVGYSVIIL 326
Query: 237 SIVNVFKGFEALGDFVGDRYKNWKHAYIGIIGALGGIAVLLEAYTWMVCMKRKKAD--NK 294
++VNVFKG + L K W++AY II LG +AV+LE +TW V +KR KA+ K
Sbjct: 327 AVVNVFKGLDILSP-----EKQWRNAYTAIIVVLGIVAVVLEGFTWYVVIKRGKAEASAK 381
Query: 295 TSDGV 299
TS V
Sbjct: 382 TSQRV 386
>At5g35735 unknown protein
Length = 404
Score = 290 bits (741), Expect = 9e-79
Identities = 153/317 (48%), Positives = 201/317 (63%), Gaps = 19/317 (5%)
Query: 1 MIGAQALVAIPQASGSP-KAYTSNIVDTSTRLQEGTISYPVSGLSATYQNNKVTIFATLT 59
M+G QALVA + + +AYTS++ TRL+ ++S+ VSGLSAT + +VTIFATL
Sbjct: 90 MVGTQALVAFTNTTTNQFQAYTSSVSSYGTRLERSSLSFGVSGLSATLVSGEVTIFATLE 149
Query: 60 LPNGTTSLVHVWQDGVLSSDSTPQEHSHESSHQNSKEVLDLVSGTSQAASG-IGSRQRRR 118
L + +WQ G + + P H + S +D +G + A G G R R+R
Sbjct: 150 LSPNLITANQLWQVGPVVN-GVPASHQTSGDNMRSSGRIDFRTGQASAGGGGSGDRLRKR 208
Query: 119 NTHGVLNAISWGILMPTGAVIARYLKVFKSADPAWFYLHITCQVSAYIVGLSGFGTGLKL 178
NTHGVLNA+SWG+LMP GA++ARY+KVF ADP WFYLHI QVS Y++G++G+ TG+KL
Sbjct: 209 NTHGVLNAVSWGVLMPMGAMMARYMKVF--ADPTWFYLHIAFQVSGYVIGVAGWATGIKL 266
Query: 179 GSDSEGITYDTHRALAIVLVTLATLQVFALFLRPNKDHKLRFYWNIYHHVVGYVTISISI 238
G+DS G +Y THR L I L T ATLQVFAL +RP DHK R YWN+YHH VGY TI +SI
Sbjct: 267 GNDSPGTSYSTHRNLGIALFTFATLQVFALLVRPKPDHKYRTYWNVYHHTVGYTTIILSI 326
Query: 239 VNVFKGFEALGDFVGDRYKNWKHAYIGIIGALGGIAVLLEAYTWMVCMKRKK-------- 290
VN+FKGF+ L D W+ AYIGI+ LG ++LE TW + ++RK
Sbjct: 327 VNIFKGFDIL-----DPEDKWRWAYIGILIFLGACVLILEPLTWFIVLRRKSRGGNTVAA 381
Query: 291 -ADNKTSDGVNGANGHG 306
+K S+GVNG G
Sbjct: 382 PTSSKYSNGVNGTTTTG 398
>At4g12980 unknown protein
Length = 394
Score = 254 bits (648), Expect = 5e-68
Identities = 142/303 (46%), Positives = 188/303 (61%), Gaps = 18/303 (5%)
Query: 1 MIGAQALVAIPQAS-GSPKAYTSNIVDTSTRLQEGTISYPVSGLSATYQNNK---VTIFA 56
M G+QALVA S G T NIV S+ L +S+ V + A N + IFA
Sbjct: 90 MAGSQALVASKDPSTGVASVTTLNIVSYSS-LVPSKLSFDVWDVKAEEAANDGGALRIFA 148
Query: 57 TLTLPNGTTS---LVHVWQDGVLSSDSTPQEHSHESSHQNSKEVLDLVSGTS----QAAS 109
+ +P + + VWQ G S+ Q H + NS LDL T
Sbjct: 149 KVKVPADLAASGKVNQVWQVGPGVSNGRIQAHDFSGPNLNSVGSLDLTGTTPGVPVSGGG 208
Query: 110 GIG-SRQRRRNTHGVLNAISWGILMPTGAVIARYLKVFKSADPAWFYLHITCQVSAYIVG 168
G G SR +RN HG+LNA+SWG+L P GA+IARY+++F+SADPAWFYLH++CQ SAY++G
Sbjct: 209 GAGNSRIHKRNIHGILNAVSWGLLFPIGAMIARYMRIFESADPAWFYLHVSCQFSAYVIG 268
Query: 169 LSGFGTGLKLGSDSEGITYDTHRALAIVLVTLATLQVFALFLRPNKDHKLRFYWNIYHHV 228
++G+ TGLKLGS+S+GI Y+THR + I L ++ATLQ+FA+ LRP KDHK RF WNIYHH
Sbjct: 269 VAGWATGLKLGSESKGIQYNTHRNIGICLFSIATLQMFAMLLRPRKDHKFRFVWNIYHHG 328
Query: 229 VGYVTISISIVNVFKGFEALGDFVGDRYKNWKHAYIGIIGALGGIAVLLEAYTWMVCMKR 288
VGY + + I+NVFKG L + +K AYI +IG LGGI +LLE TW++ +KR
Sbjct: 329 VGYSILILGIINVFKGLSIL-----NPKHTYKTAYIAVIGTLGGITLLLEVVTWVIVLKR 383
Query: 289 KKA 291
K A
Sbjct: 384 KSA 386
>At3g25290 unknown protein
Length = 393
Score = 249 bits (636), Expect = 1e-66
Identities = 133/300 (44%), Positives = 183/300 (60%), Gaps = 16/300 (5%)
Query: 1 MIGAQALVAIPQASGSPKAYTSNIVDTSTRLQEGTISYPVSGLSA---TYQNNKVTIFAT 57
M+G+Q LVA + + + + L +++ V + A + IFA
Sbjct: 89 MVGSQTLVAYKDPGNGVAVVKTLNISSYSSLIPSKLAFDVWDMKAEEAARDGGSLRIFAR 148
Query: 58 LTLPNGTTS---LVHVWQDGV-LSSDSTPQEHSHESSHQNSKEVLDLVS----GTSQAAS 109
+ +P + + VWQ G L H+ +S++ S LDL GT
Sbjct: 149 VKVPADLVAKGKVNQVWQVGPELGPGGMIGRHAFDSANLASMSSLDLKGDNSGGTISGGD 208
Query: 110 GIGSRQRRRNTHGVLNAISWGILMPTGAVIARYLKVFKSADPAWFYLHITCQVSAYIVGL 169
+ ++ + RN HG+LNA+SWGIL P GA+IARY++VF SADPAWFYLH++CQ SAY++G+
Sbjct: 209 EVNAKIKNRNIHGILNAVSWGILFPIGAIIARYMRVFDSADPAWFYLHVSCQFSAYVIGV 268
Query: 170 SGFGTGLKLGSDSEGITYDTHRALAIVLVTLATLQVFALFLRPNKDHKLRFYWNIYHHVV 229
+G+ TGLKLG++SEGI + HR + I L TLAT+Q+FA+ LRP KDHK RFYWNIYHH V
Sbjct: 269 AGWATGLKLGNESEGIRFSAHRNIGIALFTLATIQMFAMLLRPKKDHKYRFYWNIYHHGV 328
Query: 230 GYVTISISIVNVFKGFEALGDFVGDRYKNWKHAYIGIIGALGGIAVLLEAYTWMVCMKRK 289
GY +++ I+NVFKG L D YK AYI +I LGGIA+LLEA TW+V +KRK
Sbjct: 329 GYAILTLGIINVFKGLNILKP--QDTYKT---AYIAVIAVLGGIALLLEAITWVVVLKRK 383
>At3g59070 putative protein
Length = 466
Score = 211 bits (536), Expect = 5e-55
Identities = 115/281 (40%), Positives = 158/281 (55%), Gaps = 9/281 (3%)
Query: 1 MIGAQALVAIPQA-SGSPKAYTSNIVDTSTRLQEGTISYPVSGLSATYQNNKVTIFATLT 59
M+GAQALVA + SG +AYTS+I S LQE +S V+ +SA Y N ++ IFATL
Sbjct: 98 MVGAQALVAYRNSTSGVMRAYTSSINSYSPMLQESPLSLRVTQVSAEYSNGEMMIFATLV 157
Query: 60 LPNGTTSLVHVWQDGVLSSDSTPQEHSHESSHQNSKEVLDLVSGTSQAASGIGSRQRR-R 118
LP TT + H+WQDG L H+ + S LDL+SG + +
Sbjct: 158 LPPNTTVVNHLWQDGPLKEGDRLGMHAMSGDNLKSMASLDLLSGQVTTTKSVNRNMLLVK 217
Query: 119 NTHGVLNAISWGILMPTGAVIARYLKVFKSADPAWFYLHITCQVSAYIVGL-SGFGTGLK 177
H ++NA+SWGILMP G + ARY+K ++ DP WFY+H+ CQ + Y GL G GT +
Sbjct: 218 QIHAIVNALSWGILMPIGVMAARYMKNYEVLDPTWFYIHVVCQTTGYFSGLIGGLGTAIY 277
Query: 178 LGSDSEGITYDTHRALAIVLVTLATLQVFALFLRPNKDHKLRFYWNIYHHVVGYVTISIS 237
+ + G+ H + ++L L LQ+ +L RPNKDHK R YWN YHH +GY+ I +S
Sbjct: 278 MARHT-GMRTTLHTVIGLLLFALGFLQILSLKARPNKDHKYRKYWNWYHHTMGYIVIVLS 336
Query: 238 IVNVFKGFEALGDFVGDRYKNWKHAYIGIIGALGGIAVLLE 278
I N++KG L WK AY II + AV++E
Sbjct: 337 IYNIYKGLSIL-----QPGSIWKIAYTTIICCIAAFAVVME 372
>At2g04850 unknown protein
Length = 404
Score = 206 bits (524), Expect = 1e-53
Identities = 115/318 (36%), Positives = 169/318 (52%), Gaps = 20/318 (6%)
Query: 1 MIGAQALVAIPQASGSPKAYTSNIVDTSTRLQEGTI-SYPVS-------------GLSAT 46
M G++ L+A P + ++D+S +LQ+G + S P+ G AT
Sbjct: 85 MTGSRVLIAFPDPNSGQLILLPYVLDSSVKLQKGPLLSRPLDLVRLSSSSASLYGGKMAT 144
Query: 47 YQNN-KVTIFATLTLPNGTTSLVHVWQDGVLSSDSTPQEHSHESSHQNSKEVLDLVSGTS 105
+N V I+A++ L + T + HVW G+ +P H S+ +S D+ SG +
Sbjct: 145 IRNGASVQIYASVKLSSNNTKIHHVWNRGLYVQGYSPTIHPTTSTDLSSFSTFDVTSGFA 204
Query: 106 QAASGIGSRQRRRNTHGVLNAISWGILMPTGAVIARYLKVFKSADPAWFYLHITCQVSAY 165
GSR + THGV+NAISWG L+P GAV ARYL+ +S P WFY+H Q++ +
Sbjct: 205 TVNQNSGSRALKV-THGVVNAISWGFLLPAGAVTARYLRQMQSIGPTWFYIHAAIQLTGF 263
Query: 166 IVGLSGFGTGLKLGSDSEGITYDTHRALAIVLVTLATLQVFALFLRPNKDHKLRFYWNIY 225
++G GF G+ LG +S G+TY HR+L I T A LQ AL RP +K R YW Y
Sbjct: 264 LLGTIGFSIGIVLGHNSPGVTYGLHRSLGIATFTAAALQTLALLFRPKTTNKFRRYWKSY 323
Query: 226 HHVVGYVTISISIVNVFKGFEALGDFVGDRYKNWKHAYIGIIGALGGIAVLLEAYTWMVC 285
HH VGY + + +VNVF+GFE L + G Y K Y + L G+ V +E +W+V
Sbjct: 324 HHFVGYACVVMGVVNVFQGFEVLRE--GRSYA--KLGYCLCLSTLVGVCVAMEVNSWVVF 379
Query: 286 MKRKKADNKTSDGVNGAN 303
++ K + DG+ G +
Sbjct: 380 CRKAKEEKMKRDGLTGVD 397
>At4g17280 unknown protein
Length = 297
Score = 164 bits (414), Expect = 7e-41
Identities = 87/168 (51%), Positives = 116/168 (68%), Gaps = 8/168 (4%)
Query: 1 MIGAQALVAIPQASGSPKAYTSNIVDTSTRLQEGTISYPVSGLSATYQNNKVTIFATLTL 60
M+GAQA+VA PQ+ G+ + YTS I T L EG +S+ VSGLSATYQNN++ + A+L L
Sbjct: 95 MVGAQAIVAYPQSDGTVRVYTSPIRSYQTSLLEGDLSFNVSGLSATYQNNEIVVLASLKL 154
Query: 61 P----NGTTSLVHVWQDGVLSSDSTPQEHSHESSHQNSKEVLDLVSGTSQAASGIG--SR 114
NG T + VWQDG +S +S H ++ S L+LVSG S AA G G S+
Sbjct: 155 AQDLGNGGT-INTVWQDGSMSGNSL-LPHPTSGNNVRSVSTLNLVSGVSAAAGGAGGSSK 212
Query: 115 QRRRNTHGVLNAISWGILMPTGAVIARYLKVFKSADPAWFYLHITCQV 162
R+RN HG+LN +SWGI+MP GA+IARYL+V KSADPAWFY+H++ ++
Sbjct: 213 LRKRNIHGILNGVSWGIMMPLGAIIARYLRVAKSADPAWFYIHVSARL 260
>At5g48750 putative protein
Length = 255
Score = 126 bits (316), Expect = 2e-29
Identities = 69/162 (42%), Positives = 101/162 (61%), Gaps = 6/162 (3%)
Query: 1 MIGAQALVAIPQA-SGSPKAYTSNIVDTSTRLQEGTISYPVSGLSATYQNNKVTIFATLT 59
MIGAQ L+A + SG +AYTS+I + LQEG +S+ V+ LSA Y N ++TIFAT+
Sbjct: 95 MIGAQTLLAYRNSTSGFMRAYTSSINGYTPMLQEGPLSFRVTQLSAEYLNREMTIFATMV 154
Query: 60 LPNGTTSLVHVWQDGVLSSDSTPQEHSHESSHQNSKEVLDLVSG---TSQAASGIGSRQR 116
P+ TT + H+WQDG L H+ +H S LDL+SG T++AA+ +
Sbjct: 155 WPSNTTVVNHLWQDGPLKEGDRLGMHAMSGNHLKSMANLDLLSGQVMTTKAAN--DNMLL 212
Query: 117 RRNTHGVLNAISWGILMPTGAVIARYLKVFKSADPAWFYLHI 158
++ HG++NA+ WGI +P G + ARY++ +K DP WFY+ I
Sbjct: 213 VKSIHGLVNAVCWGIFIPIGVMAARYMRTYKGLDPTWFYILI 254
>At3g61750 putative protein
Length = 398
Score = 117 bits (294), Expect = 6e-27
Identities = 69/203 (33%), Positives = 105/203 (50%), Gaps = 8/203 (3%)
Query: 78 SDSTPQEHSHESSHQNSKEVLDLVSGTSQAASGIGSRQRRRNTHGVLNAISWGILMPTGA 137
S + P + + H + V+ S S A S + + HGV+ + WG L+P GA
Sbjct: 177 STAYPSKLGRLTKHDDKTTVIVDFSKASGATSIKTTTSTEKTKHGVMAILGWGFLLPVGA 236
Query: 138 VIARYLKVFKSADPAWFYLHITCQVSAYIVGLSGFGTGLKLGSDSEGITYDTHRALAIVL 197
++ARYL+ DP W+YLHI Q + +I GL+ G++L + + HR + I L
Sbjct: 237 ILARYLR---HKDPLWYYLHIGFQFTGFIFGLAAVILGIQLYNRIQP-DIPAHRGIGIFL 292
Query: 198 VTLATLQVFALFLRPNKDHKLRFYWNIYHHVVGYVTISISIVNVFKGFEALGDFVGDRYK 257
+ L+TLQV A F RP K+ K+R YWN YHH +G +++ VN+ G + D GD
Sbjct: 293 LVLSTLQVLAFFARPQKETKMRRYWNWYHHWIGRISLFFGAVNIVLGIR-MADNGGD--- 348
Query: 258 NWKHAYIGIIGALGGIAVLLEAY 280
WK Y ++ V+LE +
Sbjct: 349 GWKIGYGFVLSVTLLAFVVLEIF 371
>At3g07575 unknown protein
Length = 369
Score = 116 bits (291), Expect = 1e-26
Identities = 59/174 (33%), Positives = 93/174 (52%), Gaps = 8/174 (4%)
Query: 105 SQAASGIGSRQRRRNTHGVLNAISWGILMPTGAVIARYLKVFKSADPAWFYLHITCQVSA 164
SQ+ + + + THG++N WGIL+ GA++AR++K + DP WFY HI Q +
Sbjct: 197 SQSVVKVSPHSKLKKTHGLMNMFGWGILIIVGAIVARHMKQW---DPTWFYAHIALQTTG 253
Query: 165 YIVGLSGFGTGLKLGSDSEGITYDTHRALAIVLVTLATLQVFALFLRPNKDHKLRFYWNI 224
+++GL+G GL L + + H+ L I ++ + LQ+ AL RP+K K R YWN
Sbjct: 254 FLLGLTGVICGLVLENRLKANNVSKHKGLGITILVMGVLQMLALLARPDKQSKYRKYWNW 313
Query: 225 YHHVVGYVTISISIVNVFKGFEALGDFVGDRYKNWKHAYIGIIGALGGIAVLLE 278
YHH +G + I ++I N+F G + +W Y + L A+ LE
Sbjct: 314 YHHNIGRLLIILAISNIFYGIH-----LAKAGTSWNGGYGFAVAVLALTAIGLE 362
>At5g54830 unknown protein
Length = 907
Score = 68.9 bits (167), Expect = 3e-12
Identities = 54/195 (27%), Positives = 85/195 (42%), Gaps = 17/195 (8%)
Query: 64 TTSLVHVWQDGVLSSDSTPQEHS-HESSHQNSKEVLDLVSGTSQAASGIGSRQRRRNTHG 122
TT L +W G +D E + H + Q V+ L G+++A + + HG
Sbjct: 635 TTPLKVIWAMGAKWTDGQLTERNMHSVTSQRPVRVM-LTRGSAEADQDL---RPVLGVHG 690
Query: 123 VLNAISWGILMPTGAVIARYLKVFKSADPAWFYLHITCQVSAYIVGLSGFGTGLKLGSDS 182
+ ++WGIL+P G + ARYLK K WF +H+ Q S + G L ++
Sbjct: 691 FMMFLAWGILLPGGILSARYLKHIKG--DGWFKIHMYLQCSGLAIVFLGL---LFAVAEL 745
Query: 183 EGITY-DTHRALAIVLVTLATLQVFALFLRPNKD------HKLRFYWNIYHHVVGYVTIS 235
G ++ TH + LA Q +LRP K R W H +VG +
Sbjct: 746 NGFSFSSTHVKFGFTAIVLACAQPVNAWLRPAKPAQGELISSKRLIWEYSHSIVGQSAVV 805
Query: 236 ISIVNVFKGFEALGD 250
+ +V +F G + LG+
Sbjct: 806 VGVVALFTGMKHLGE 820
>At4g18260 unknown protein
Length = 545
Score = 58.5 bits (140), Expect = 4e-09
Identities = 36/130 (27%), Positives = 62/130 (47%), Gaps = 6/130 (4%)
Query: 121 HGVLNAISWGILMPTGAVIARYL-KVFKSA--DPAWFYLHITCQVSAYIVGLSGFGTGLK 177
HG+L +S G LMP G + R K ++ +FYLH+ Q+ A ++ G L+
Sbjct: 64 HGILLWVSMGFLMPVGILFIRMANKAHENGIKVKVFFYLHVIFQILAVVLATIGAILSLR 123
Query: 178 LGSDSEGITYDTHRALAIVLVTLATLQVFALFLRPNKDHKLRFYWNIYHHVVGYVTISIS 237
+S + H+ L + L LQ +P++ K R W + H ++G + +
Sbjct: 124 TLENSFD---NNHQRLGLALYAAMWLQFLTGVFKPSRGSKRRLRWFLLHWILGTIVSIVG 180
Query: 238 IVNVFKGFEA 247
IVN++ G +A
Sbjct: 181 IVNIYTGIQA 190
>At2g30890 hypothetical protein
Length = 257
Score = 55.1 bits (131), Expect = 5e-08
Identities = 41/197 (20%), Positives = 81/197 (40%), Gaps = 19/197 (9%)
Query: 121 HGVLNAISWGILMPTGAVIARYLKVFKSADPAW---FYLHITCQVSAYIVGLSGFGTGLK 177
HG + + G+LMP G + R + + F+LH+T Q+ A I+ +
Sbjct: 56 HGFMLWAAMGVLMPIGIISIRLMSIKDQPIITLRRLFFLHVTSQMVAVIL--------VT 107
Query: 178 LGSDSEGITYDT-----HRALAIVLVTLATLQVFALFLRPNKDHKLRFYWNIYHHVVGYV 232
+G+ I ++ H+ L I L + Q FLRP ++ K R W + H ++G
Sbjct: 108 IGAVMSVINFNNSFSNHHQQLGIGLYVIVWFQALLGFLRPPREEKARRKWFVGHWILGTS 167
Query: 233 TISISIVNVFKGFEALGDFVGDRYKNWKHAYIGIIGALGGIAVLLEAYTWM---VCMKRK 289
+ I+N++ G A W + + + + + + ++++ R
Sbjct: 168 IAILGIINIYTGLHAYAKKTSKSANLWTILFTAQLSCIALVYLFQDKWSYIQSQATFNRN 227
Query: 290 KADNKTSDGVNGANGHG 306
++ + S+ GHG
Sbjct: 228 QSVDHNSNISTAETGHG 244
>At3g07390 auxin-induced protein (AIR12)
Length = 273
Score = 43.5 bits (101), Expect = 1e-04
Identities = 31/111 (27%), Positives = 50/111 (44%), Gaps = 4/111 (3%)
Query: 1 MIGAQALVAIPQASGSPKAYTSNIVDTSTRLQEGTISYPVSGLSA-TYQNNKVTIFATLT 59
M G+QA +A G+ + + + + L EG +++ L A + ++ IF T+
Sbjct: 112 MAGSQAFLAYRSGGGAAPVVKTYNISSYSSLVEGKLAFDFWNLRAESLSGGRIAIFTTVK 171
Query: 60 LPNGTTSLVHVWQDGVLSSDSTPQEHSHESSHQNSKEVLDLVSGTSQAASG 110
+P G S+ VWQ G ++ P H + S VL S T AA G
Sbjct: 172 VPAGADSVNQVWQIGGNVTNGRPGVHPFGPDNLGSHRVL---SFTEDAAPG 219
>At5g53660 putative protein
Length = 365
Score = 32.7 bits (73), Expect = 0.26
Identities = 26/89 (29%), Positives = 40/89 (44%), Gaps = 3/89 (3%)
Query: 17 PKAYTSNIVDTSTRLQEGTISYPVSGLSATYQNNKVTIFATLTLPNGTTSLVHVWQDGVL 76
P+ +S+ + T L+ +S +S + + K T+ AT N + + VW DGV
Sbjct: 172 PQKLSSSSIKDKT-LEPMEVSSSISNYRDSRGSEKFTVLATTEQENKYLNFIDVWSDGVR 230
Query: 77 SSD--STPQEHSHESSHQNSKEVLDLVSG 103
SS+ ST S+ S LDL G
Sbjct: 231 SSEKQSTTSTPVSSSNGNLSLYSLDLSMG 259
>At5g38630 cytochrome b-561 (gb|AAD45585.1)
Length = 230
Score = 30.4 bits (67), Expect = 1.3
Identities = 32/131 (24%), Positives = 54/131 (40%), Gaps = 9/131 (6%)
Query: 119 NTHGVLNAISWGILMPTGAVIARYLKVFKSADPAWFYLHITCQVSAYIVGLSGFGTGLKL 178
N H V+ I G+++ G + Y V + +H+T Q++A+I+ L G LK
Sbjct: 49 NVHPVMMVI--GLILFNGEAMLAYKSV-QGTKNLKKLVHLTLQLTAFILSLIGVWAALKF 105
Query: 179 GSDSEGIT--YDTHRALAIVLVTLATLQ---VFALFLRPNKDHKLRFYWNIYHHVVGYVT 233
D +GI Y H L + + L Q F + P R +H +G
Sbjct: 106 HID-KGIENFYSLHSWLGLACLFLFAFQWAAGFVTYWYPGGSRNSRASLMPWHVFLGISI 164
Query: 234 ISISIVNVFKG 244
++++V G
Sbjct: 165 YALALVTATTG 175
>At4g15840 unknown protein
Length = 298
Score = 30.0 bits (66), Expect = 1.7
Identities = 19/57 (33%), Positives = 28/57 (48%)
Query: 66 SLVHVWQDGVLSSDSTPQEHSHESSHQNSKEVLDLVSGTSQAASGIGSRQRRRNTHG 122
S V + ++G S S E + S + SK+ L + G SQ SG+G+ R T G
Sbjct: 60 SRVPINRNGQQSIASDGSEGRIDQSGEISKDGLTALVGLSQGTSGVGNNPRGEQTEG 116
>At1g20540 unknown protein
Length = 351
Score = 29.6 bits (65), Expect = 2.2
Identities = 17/59 (28%), Positives = 26/59 (43%)
Query: 58 LTLPNGTTSLVHVWQDGVLSSDSTPQEHSHESSHQNSKEVLDLVSGTSQAASGIGSRQR 116
L L GT S+V++W SSD E ES+ Q +L+ + + G+ R
Sbjct: 267 LILSAGTDSVVNLWYASATSSDDKTSESPVESTRQRVNPLLNSYTDYEDSVYGLAWSSR 325
>At1g30760 putative reticuline oxidase-like protein, 3' partial
Length = 431
Score = 28.9 bits (63), Expect = 3.7
Identities = 25/93 (26%), Positives = 40/93 (42%), Gaps = 3/93 (3%)
Query: 124 LNAISWGILMPTGAVIARYLKVFKSADPAWFYLHITCQVSAYIVGLSGFGTGLKL---GS 180
LNA S+ + + T A RYL F V A ++ L+L G
Sbjct: 58 LNASSFKLALETSAQNLRYLMPSNPKPEFIFEPLYETHVQAAVLCAKKLKLHLRLRSGGH 117
Query: 181 DSEGITYDTHRALAIVLVTLATLQVFALFLRPN 213
D EG++Y + A V+V L+ L+ ++ + N
Sbjct: 118 DYEGLSYVSEMETAFVIVDLSKLRQISVDIESN 150
>At3g07660 unknown protein
Length = 782
Score = 28.5 bits (62), Expect = 4.9
Identities = 26/104 (25%), Positives = 44/104 (42%), Gaps = 17/104 (16%)
Query: 11 PQASGSPKAYTSNIVDTSTRLQEGTISYPVSGL-----SATYQNNKVTIFA--------- 56
P++ S +SN+ S +L+ G S P + + S N+V + +
Sbjct: 97 PKSGSSDVRQSSNLPAGSGQLKAGRSSVPSNNIDGAPSSVELSKNRVAVGSRVVHTEQKA 156
Query: 57 --TLTLPNGTTSLVHVWQDGVLSSDSTPQEHSHES-SHQNSKEV 97
+L LP ++S V S + P++H ES SH S+ V
Sbjct: 157 VDSLPLPRPSSSEVRFTSSNPKSEQTAPEQHIGESKSHTRSRGV 200
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.134 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,901,306
Number of Sequences: 26719
Number of extensions: 286052
Number of successful extensions: 833
Number of sequences better than 10.0: 21
Number of HSP's better than 10.0 without gapping: 16
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 790
Number of HSP's gapped (non-prelim): 22
length of query: 310
length of database: 11,318,596
effective HSP length: 99
effective length of query: 211
effective length of database: 8,673,415
effective search space: 1830090565
effective search space used: 1830090565
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 60 (27.7 bits)
Medicago: description of AC147406.4


