

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
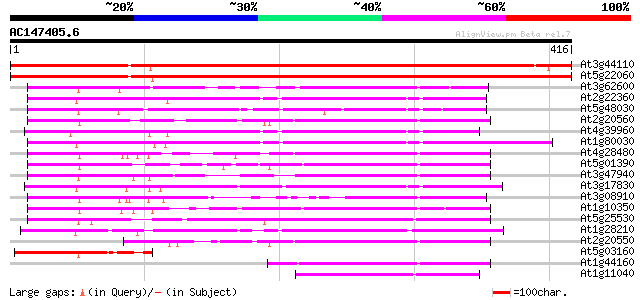
Query= AC147405.6 - phase: 0
(416 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At3g44110 dnaJ protein homolog atj3 730 0.0
At5g22060 DNAJ PROTEIN HOMOLOG ATJ 708 0.0
At3g62600 unknown protein 207 9e-54
At2g22360 DnaJ like protein 191 5e-49
At5g48030 DnaJ protein-like 189 3e-48
At2g20560 putative heat shock protein 182 2e-46
At4g39960 DnaJ like protein 179 3e-45
At1g80030 unknown protein 179 3e-45
At4g28480 heat-shock protein 169 3e-42
At5g01390 heat shock protein 40-like 167 1e-41
At3g47940 heat shock protein-like protein 166 2e-41
At3g17830 DnaJ, putative 165 5e-41
At3g08910 putative heat shock protein 162 2e-40
At1g10350 putative heat-shock protein 154 7e-38
At5g25530 heat-shock protein - like 139 3e-33
At1g28210 AtJ1 (atj) 137 1e-32
At2g20550 putative heat shock protein 108 4e-24
At5g03160 unknown protein 96 3e-20
At1g44160 putative protein 89 6e-18
At1g11040 DnaJ isolog 88 1e-17
>At3g44110 dnaJ protein homolog atj3
Length = 420
Score = 730 bits (1884), Expect = 0.0
Identities = 363/422 (86%), Positives = 388/422 (91%), Gaps = 8/422 (1%)
Query: 1 MFGRAP-KKSDSTRYYEILGVSKTASQDDLKKAYKKAAIKNHPDKGGDPEKFKELAQAYE 59
MFGR P KKSD+T++YEILGV K+AS +DLKKAYKKAAIKNHPDKGGDPEKFKELAQAYE
Sbjct: 1 MFGRGPSKKSDNTKFYEILGVPKSASPEDLKKAYKKAAIKNHPDKGGDPEKFKELAQAYE 60
Query: 60 VLSDPEKREIYDTYGEDALKEGMGGGGGGGHDPFDIFSSFFGGG--GGGSSRGRRQRRGE 117
VLSDPEKREIYD YGEDALKEGMGGGGGG HDPFDIFSSFFGGG GG +SR RRQRRGE
Sbjct: 61 VLSDPEKREIYDQYGEDALKEGMGGGGGG-HDPFDIFSSFFGGGPFGGNTSRQRRQRRGE 119
Query: 118 DVVHPLKVSLEDLYLGTSKKLSLSRNVLCSKCSGKGSKSGASMKCAGCQGTGMKVSIRHL 177
DVVHPLKVSLED+YLGT KKLSLSRN LCSKC+GKGSKSGAS+KC GCQG+GMKVSIR L
Sbjct: 120 DVVHPLKVSLEDVYLGTMKKLSLSRNALCSKCNGKGSKSGASLKCGGCQGSGMKVSIRQL 179
Query: 178 GPSMIQQMQHPCNECKGTGETINDKDRCPQCKGEKVVQEKKVLEVHVEKGMQNSQKITFP 237
GP MIQQMQH CNECKGTGETIND+DRCPQCKG+KV+ EKKVLEV+VEKGMQ+SQKITF
Sbjct: 180 GPGMIQQMQHACNECKGTGETINDRDRCPQCKGDKVIPEKKVLEVNVEKGMQHSQKITFE 239
Query: 238 GEADEAPDTVTGDIVFVLQQKEHPKFKRKSEDLFVEHTLSLTEALCGFQFVLTHLDGRQL 297
G+ADEAPDTVTGDIVFVLQQKEHPKFKRK EDLFVEHTLSLTEALCGFQFVLTHLDGR L
Sbjct: 240 GQADEAPDTVTGDIVFVLQQKEHPKFKRKGEDLFVEHTLSLTEALCGFQFVLTHLDGRSL 299
Query: 298 LIKSNPGEVVKPDSYKAINDEGMPMYQRPFMKGKLYIHFTVEFPDTLSLDQVKGLEAVLP 357
LIKSNPGEVVKPDSYKAI+DEGMP+YQRPFMKGKLYIHFTVEFPD+LS DQ K LEAVLP
Sbjct: 300 LIKSNPGEVVKPDSYKAISDEGMPIYQRPFMKGKLYIHFTVEFPDSLSPDQTKALEAVLP 359
Query: 358 AKPSSQLTDMEIDECEETTLHDVNMEEENRRKQQQQQQEAY---DEDDDMPGGAQRVQCA 414
++QL+DMEIDECEETTLHDVN+E+E RRK Q Q+EAY DEDDD PGGAQRVQCA
Sbjct: 360 KPSTAQLSDMEIDECEETTLHDVNIEDEMRRK-AQAQREAYDDDDEDDDHPGGAQRVQCA 418
Query: 415 QQ 416
QQ
Sbjct: 419 QQ 420
>At5g22060 DNAJ PROTEIN HOMOLOG ATJ
Length = 419
Score = 708 bits (1828), Expect = 0.0
Identities = 343/420 (81%), Positives = 381/420 (90%), Gaps = 5/420 (1%)
Query: 1 MFGRAP-KKSDSTRYYEILGVSKTASQDDLKKAYKKAAIKNHPDKGGDPEKFKELAQAYE 59
MFGR P +KSD+T++YEILGV KTA+ +DLKKAYKKAAIKNHPDKGGDPEKFKELAQAYE
Sbjct: 1 MFGRGPSRKSDNTKFYEILGVPKTAAPEDLKKAYKKAAIKNHPDKGGDPEKFKELAQAYE 60
Query: 60 VLSDPEKREIYDTYGEDALKEGMGGGGGGGHDPFDIFSSFFGGGG---GGSSRGRRQRRG 116
VLSDPEKREIYD YGEDALKEGMGGGGGG HDPFDIFSSFFG GG G SRGRRQRRG
Sbjct: 61 VLSDPEKREIYDQYGEDALKEGMGGGGGG-HDPFDIFSSFFGSGGHPFGSHSRGRRQRRG 119
Query: 117 EDVVHPLKVSLEDLYLGTSKKLSLSRNVLCSKCSGKGSKSGASMKCAGCQGTGMKVSIRH 176
EDVVHPLKVSLED+YLGT+KKLSLSR LCSKC+GKGSKSGASMKC GCQG+GMK+SIR
Sbjct: 120 EDVVHPLKVSLEDVYLGTTKKLSLSRKALCSKCNGKGSKSGASMKCGGCQGSGMKISIRQ 179
Query: 177 LGPSMIQQMQHPCNECKGTGETINDKDRCPQCKGEKVVQEKKVLEVHVEKGMQNSQKITF 236
GP M+QQ+QH CN+CKGTGETIND+DRCPQCKGEKVV EKKVLEV+VEKGMQ++QKITF
Sbjct: 180 FGPGMMQQVQHACNDCKGTGETINDRDRCPQCKGEKVVSEKKVLEVNVEKGMQHNQKITF 239
Query: 237 PGEADEAPDTVTGDIVFVLQQKEHPKFKRKSEDLFVEHTLSLTEALCGFQFVLTHLDGRQ 296
G+ADEAPDTVTGDIVFV+QQKEHPKFKRK EDLFVEHT+SLTEALCGFQFVLTHLD RQ
Sbjct: 240 SGQADEAPDTVTGDIVFVIQQKEHPKFKRKGEDLFVEHTISLTEALCGFQFVLTHLDKRQ 299
Query: 297 LLIKSNPGEVVKPDSYKAINDEGMPMYQRPFMKGKLYIHFTVEFPDTLSLDQVKGLEAVL 356
LLIKS PGEVVKPDSYKAI+DEGMP+YQRPFMKGKLYIHFTVEFP++LS DQ K +EAVL
Sbjct: 300 LLIKSKPGEVVKPDSYKAISDEGMPIYQRPFMKGKLYIHFTVEFPESLSPDQTKAIEAVL 359
Query: 357 PAKPSSQLTDMEIDECEETTLHDVNMEEENRRKQQQQQQEAYDEDDDMPGGAQRVQCAQQ 416
P + ++DMEID+CEETTLHDVN+E+E +RK Q Q++ D+++D PGGAQRVQCAQQ
Sbjct: 360 PKPTKAAISDMEIDDCEETTLHDVNIEDEMKRKAQAQREAYDDDEEDHPGGAQRVQCAQQ 419
>At3g62600 unknown protein
Length = 346
Score = 207 bits (527), Expect = 9e-54
Identities = 132/350 (37%), Positives = 190/350 (53%), Gaps = 40/350 (11%)
Query: 14 YYEILGVSKTASQDDLKKAYKKAAIKNHPDKGGDPE----KFKELAQAYEVLSDPEKREI 69
YY++L V K AS + +K+AY+K A+K HPDK E KF E+ AYEVLSD EKREI
Sbjct: 27 YYDVLQVPKGASDEQIKRAYRKLALKYHPDKNQGNEEATRKFAEINNAYEVLSDEEKREI 86
Query: 70 YDTYGEDALKE----GMGGGGGGGHDPFDIFSSFFGGGGGGSSRGRRQRRGEDVVHPLKV 125
Y+ YGE+ LK+ G GGGGGG + DIFSSFFGGG + +G+DV+ L+
Sbjct: 87 YNKYGEEGLKQFSANGGRGGGGGGMNMQDIFSSFFGGGS--MEEEEKVVKGDDVIVELEA 144
Query: 126 SLEDLYLGTSKKLSLSRNVLCSKCSGKGSKSGASMKCAGCQGTGMKVSIRHLGPSMIQQM 185
+LEDLY+G S K+ +NV+ K + C+ +V R +GP M QQM
Sbjct: 145 TLEDLYMGGSMKVWREKNVI---------KPAPGKRKCNCRN---EVYHRQIGPGMFQQM 192
Query: 186 QHPCNECKGTGETINDKDRCPQCKGEKVVQEKKVLEVHVEKGMQNSQKITFPGEADEAPD 245
T D+CP K E+ E + V +EKGM++ ++++F + + D
Sbjct: 193 ------------TEQVCDKCPNVKYER---EGYFVTVDIEKGMKDGEEVSFYEDGEPILD 237
Query: 246 TVTGDIVFVLQQKEHPKFKRKSEDLFVEHTLSLTEALCGFQFVLTHLDGRQLLIKSNPGE 305
GD+ F ++ H +F+R DL + ++L EAL GF+ HLD ++ I S
Sbjct: 238 GDPGDLKFRIRTAPHARFRRDGNDLHMNVNITLVEALVGFEKSFKHLDDHEVDISSK--G 295
Query: 306 VVKPDSYKAINDEGMPMYQRPFMKGKLYIHFTVEFPDTLSLDQVKGLEAV 355
+ KP K EGMP++ KG L++ F V FP +L+ DQ K ++ V
Sbjct: 296 ITKPKEVKKFKGEGMPLHYST-KKGNLFVTFEVLFPSSLTDDQKKKIKEV 344
>At2g22360 DnaJ like protein
Length = 442
Score = 191 bits (486), Expect = 5e-49
Identities = 118/347 (34%), Positives = 177/347 (51%), Gaps = 14/347 (4%)
Query: 14 YYEILGVSKTASQDDLKKAYKKAAIKNHPDKGGDP---EKFKELAQAYEVLSDPEKREIY 70
YY +LGVSK A++ ++K AY+K A HPD DP EKFKE++ AYEVLSD EK+ +Y
Sbjct: 87 YYSVLGVSKNATKAEIKSAYRKLARNYHPDVNKDPGAEEKFKEISNAYEVLSDDEKKSLY 146
Query: 71 DTYGEDALKEGMGGGGGGGHDPFDIFSSFFGGGGGGSSRGRRQRR--GEDVVHPLKVSLE 128
D YGE LK G G G +PFD+F S F G GGG RG R R G+D + L ++ +
Sbjct: 147 DRYGEAGLKGAAGFGNGDFSNPFDLFDSLFEGFGGGMGRGSRSRAVDGQDEYYTLILNFK 206
Query: 129 DLYLGTSKKLSLSRNVLCSKCSGKGSKSGAS-MKCAGCQGTGMKVSIRHLGPSMIQQMQH 187
+ G K++ +SR C C G G+K G KC C G G VS + QQ+
Sbjct: 207 EAVFGMEKEIEISRLESCGTCEGSGAKPGTKPTKCTTCGGQGQVVSAARTPLGVFQQVM- 265
Query: 188 PCNECKGTGETINDKDRCPQCKGEKVVQEKKVLEVHVEKGMQNSQKITFPGEADEAP-DT 246
C+ C GTGE C C G+ V++ K + + V G+ + ++ GE +
Sbjct: 266 TCSSCNGTGEI---STPCGTCSGDGRVRKTKRISLKVPAGVDSGSRLRVRGEGNAGKRGG 322
Query: 247 VTGDIVFVLQQKEHPKFKRKSEDLFVEHTLSLTEALCGFQFVLTHLDGRQLLIKSNPGEV 306
GD+ V++ P KR ++ +S +A+ G + +DG + +K G
Sbjct: 323 SPGDLFVVIEVIPDPILKRDDTNILYTCKISYIDAILGTTLKVPTVDG-TVDLKVPAG-- 379
Query: 307 VKPDSYKAINDEGMPMYQRPFMKGKLYIHFTVEFPDTLSLDQVKGLE 353
+P + + +G+P+ + M+G + VE P LS ++ K +E
Sbjct: 380 TQPSTTLVMAKKGVPVLNKSNMRGDQLVRVQVEIPKRLSKEEKKLIE 426
>At5g48030 DnaJ protein-like
Length = 456
Score = 189 bits (479), Expect = 3e-48
Identities = 124/357 (34%), Positives = 191/357 (52%), Gaps = 32/357 (8%)
Query: 14 YYEILGVSKTASQDDLKKAYKKAAIKNHPDKG-GDPE---KFKELAQAYEVLSDPEKREI 69
YY +LGVSK A + ++KKAY A K HPD DPE KF+E+++AYE+L D EKR++
Sbjct: 95 YYSVLGVSKNAQEGEIKKAYYGLAKKLHPDMNKDDPEAETKFQEVSKAYEILKDKEKRDL 154
Query: 70 YDTYGEDALK----------EGMGGGGGGGHDPFDIFSSFFGGGGGGSSRGRRQRRGEDV 119
YD G +A + +G GGGGGGG +PFDIF SF G + R+ G+DV
Sbjct: 155 YDQVGHEAFEQNASGGFPNDQGFGGGGGGGFNPFDIFGSF---NGDIFNMYRQDIGGQDV 211
Query: 120 VHPLKVSLEDLYLGTSKKLSLSRNVLCSKCSGKGSKSGASM-KCAGCQGTGMKVSIRHLG 178
L +S + G SK ++ + C+ C G+G G KC C G+GM S+R
Sbjct: 212 KVLLDLSFMEAVQGCSKTVTFQTEMACNTCGGQGVPPGTKREKCKACNGSGM-TSLRR-- 268
Query: 179 PSMIQQMQHPCNECKGTGETINDKDRCPQCKGEKVVQEKKVLEVHVEKGMQNSQ--KITF 236
+ +Q C +C G G+T + C C+G +VV+ +K ++V ++ G+ NS K+
Sbjct: 269 --GMLSIQTTCQKCGGAGQTFS--SICKSCRGARVVRGQKSVKVTIDPGVDNSDTLKVAR 324
Query: 237 PGEADEAPDTVTGDIVFVLQQKEHPKFKRKSEDLFVEHTLSLTEALCGFQFVLTHLDGRQ 296
G AD D GD+ L+ +E P F+R+ D+ V+ LS+T+A+ G + L G
Sbjct: 325 VGGADPEGDQ-PGDLYVTLKVREDPVFRREGSDIHVDAVLSVTQAILGGTIQVPTLTG-D 382
Query: 297 LLIKSNPGEVVKPDSYKAINDEGMPMYQRPFMKGKLYIHFTVEFPDTLSLDQVKGLE 353
+++K PG +P + ++G+ ++ G Y+HF V P ++ Q + LE
Sbjct: 383 VVVKVRPG--TQPGHKVVLRNKGI-RARKSTKFGDQYVHFNVSIPANITQRQRELLE 436
>At2g20560 putative heat shock protein
Length = 337
Score = 182 bits (463), Expect = 2e-46
Identities = 120/353 (33%), Positives = 182/353 (50%), Gaps = 31/353 (8%)
Query: 14 YYEILGVSKTASQDDLKKAYKKAAIKNHPDKGGDPEK-----FKELAQAYEVLSDPEKRE 68
YY++L V ++AS DDLKKAY+K A+K HPDK + +K FK++++AYEVLSDP+K+
Sbjct: 5 YYKVLQVDRSASDDDLKKAYRKLAMKWHPDKNPNNKKDAEAMFKQISEAYEVLSDPQKKA 64
Query: 69 IYDTYGEDALKEGMGGGGGGGHDPFDIFSSFFGGGGGGSSRGRRQRRGEDVVHPLKVSLE 128
+YD YGE+ LK + GG +++F G G +S R +D+
Sbjct: 65 VYDQYGEEGLKGNVPPPDAGG-------ATYFSTGDGPTSFRFNPRNADDIFAE------ 111
Query: 129 DLYLGTSKKLSLSRNVLCSKCSGKGSKSGASMKCAGCQGTGMKVSIRHLGPSMIQQMQH- 187
+ G S R S G AS G G G S+ H G +++
Sbjct: 112 --FFGFSSPFGGGRGGTRFSSSMFGDNMFASFGEGGGGGGG---SMHHGGARKAAPIENK 166
Query: 188 -PCNE---CKGTGETINDKDRCPQCKGEKVVQEKKVLEVHVEKGMQNSQKITFPGEADEA 243
PC+ KGT + + G K +Q +++L + V+ G + KITFP + +E
Sbjct: 167 LPCSLEDLYKGTTKKMRISREIADVSG-KTMQVEEILTIDVKPGWKKGTKITFPEKGNEQ 225
Query: 244 PDTVTGDIVFVLQQKEHPKFKRKSEDLFVEHTLSLTEALCGFQFVLTHLDGRQLLIKSNP 303
P + D+VF++ +K HP F R+ DL V +SL EAL G+ LT LDGR+L I
Sbjct: 226 PGVIPADLVFIIDEKPHPVFTREGNDLIVTQKISLVEALTGYTVNLTTLDGRRLTIPVT- 284
Query: 304 GEVVKPDSYKAINDEGMPMYQRPFMKGKLYIHFTVEFPDTLSLDQVKGLEAVL 356
VV P+ + + EGMP+ + +G L I F ++FP L+ +Q G++ +L
Sbjct: 285 -NVVHPEYEEVVPKEGMPLQKDQTKRGNLRIKFNIKFPTRLTSEQKTGVKKLL 336
>At4g39960 DnaJ like protein
Length = 447
Score = 179 bits (453), Expect = 3e-45
Identities = 114/351 (32%), Positives = 177/351 (49%), Gaps = 21/351 (5%)
Query: 12 TRYYEILGVSKTASQDDLKKAYKKAAIKNHPD---KGGDPEKFKELAQAYEVLSDPEKRE 68
T +Y +LGVSK A++ ++K AY+K A HPD G +KFKE++ AYE+LSD EKR
Sbjct: 84 TDFYSVLGVSKNATKAEIKSAYRKLARSYHPDVNKDAGAEDKFKEISNAYEILSDDEKRS 143
Query: 69 IYDTYGEDALKEGMGGGGGGGHDPFDIFSSFFGG-------GGGGSSRGRRQRR--GEDV 119
+YD YGE +K GG G +PFD+F S F G GGG SRG R R GED
Sbjct: 144 LYDRYGEAGVKGAGMGGMGDYSNPFDLFESLFEGMGGMGGMGGGMGSRGSRSRAIDGEDE 203
Query: 120 VHPLKVSLEDLYLGTSKKLSLSRNVLCSKCSGKGSKSGAS-MKCAGCQGTGMKVSIRHLG 178
+ L ++ ++ G K++ +SR C C+G G+K+G KC C G G V+
Sbjct: 204 YYSLILNFKEAVFGIEKEIEISRLESCGTCNGSGAKAGTKPTKCKTCGGQGQVVASTRTP 263
Query: 179 PSMIQQMQHPCNECKGTGETINDKDRCPQCKGEKVVQEKKVLEVHVEKGMQNSQKITFPG 238
+ QQ+ C+ C GTGE C C G+ V+ K + + V G+ + ++ G
Sbjct: 264 LGVFQQVM-TCSPCNGTGEI---SKPCGACSGDGRVRRTKRISLKVPAGVDSGSRLRVRG 319
Query: 239 EADEAP-DTVTGDIVFVLQQKEHPKFKRKSEDLFVEHTLSLTEALCGFQFVLTHLDGRQL 297
E + GD+ V++ P KR ++ +S +A+ G + +DG ++
Sbjct: 320 EGNAGKRGGSPGDLFAVIEVIPDPVLKRDDTNILYTCKISYVDAILGTTLKVPTVDG-EV 378
Query: 298 LIKSNPGEVVKPDSYKAINDEGMPMYQRPFMKGKLYIHFTVEFPDTLSLDQ 348
+K G +P + + +G+P+ + M+G + VE P LS ++
Sbjct: 379 DLKVPAG--TQPSTTLVMAKKGVPVLNKSKMRGDQLVRVQVEIPKRLSKEE 427
>At1g80030 unknown protein
Length = 500
Score = 179 bits (453), Expect = 3e-45
Identities = 121/406 (29%), Positives = 202/406 (48%), Gaps = 23/406 (5%)
Query: 14 YYEILGVSKTASQDDLKKAYKKAAIKNHPDKGGDP---EKFKELAQAYEVLSDPEKREIY 70
YY LGVSK+A+ ++K AY++ A + HPD +P EKFKE++ AYEVLSD +KR +Y
Sbjct: 76 YYATLGVSKSANNKEIKAAYRRLARQYHPDVNKEPGATEKFKEISAAYEVLSDEQKRALY 135
Query: 71 DTYGEDALKEGMGGGGGG-GHDPFDIFSSFFGGGGGG--------SSRGRRQR--RGEDV 119
D YGE +K +GG G +PFD+F +FFG GG R RR R +GED+
Sbjct: 136 DQYGEAGVKSTVGGASGPYTSNPFDLFETFFGASMGGFPGMDQADFGRTRRSRVTKGEDL 195
Query: 120 VHPLKVSLEDLYLGTSKKLSLSRNVLCSKCSGKGSKSGASMK-CAGCQGTGMKVSIRHLG 178
+ + + L + G+ K+ L+ C C+G G+K+G+ M+ C+ C G G +
Sbjct: 196 RYDITLELSEAIFGSEKEFDLTHLETCEACAGTGAKAGSKMRICSTCGGRGQVMRTEQTP 255
Query: 179 PSMIQQMQHPCNECKGTGETINDKDRCPQCKGEKVVQEKKVLEVHVEKGMQNSQKITFPG 238
M Q+ C C G GE I+ + C +C GE V+ KK ++V + G+ + G
Sbjct: 256 FGMFSQVS-ICPNCGGDGEVIS--ENCRKCSGEGRVRIKKSIKVKIPPGVSAGSILRVAG 312
Query: 239 EADEAP-DTVTGDIVFVLQQKEHPKFKRKSEDLFVEHTLSLTEALCGFQFVLTHLDGRQL 297
E D P GD+ L ++ +R +L ++S +A+ G + ++G
Sbjct: 313 EGDSGPRGGPPGDLYVYLDVEDVRGIERDGINLLSTLSISYLDAILGAVVKVKTVEG-DT 371
Query: 298 LIKSNPGEVVKPDSYKAINDEGMPMYQRPFMKGKLYIHFTVEFPDTLSLDQVKGLEAVLP 357
++ PG +P + +G+P RP ++G V P+ +S + + LE +
Sbjct: 372 ELQIPPG--TQPGDVLVLAKKGVPKLNRPSIRGDHLFTVKVSVPNQISAGERELLEELAS 429
Query: 358 AK-PSSQLTDMEIDECEETTLHDVNMEEENRRKQQQQQQEAYDEDD 402
K SS + + +TL EN++ + +++ E ++++
Sbjct: 430 LKDTSSNRSRTRAKPQQPSTLSTAPSGSENKKDEVKEENEEPEQEN 475
>At4g28480 heat-shock protein
Length = 348
Score = 169 bits (428), Expect = 3e-42
Identities = 119/376 (31%), Positives = 186/376 (48%), Gaps = 66/376 (17%)
Query: 14 YYEILGVSKTASQDDLKKAYKKAAIKNHPDKGGDPEK-----FKELAQAYEVLSDPEKRE 68
YY++L V ++A+ DDLKKAY+K A+K HPDK + +K FK++++AY+VLSDP+KR
Sbjct: 5 YYKVLQVDRSANDDDLKKAYRKLAMKWHPDKNPNNKKDAEAKFKQISEAYDVLSDPQKRA 64
Query: 69 IYDTYGEDALKEGM-------GGGG----GGGHDPF--------DIFSSFFG-----GGG 104
+YD YGE+ LK + G G G F DIF+ FFG GGG
Sbjct: 65 VYDQYGEEGLKGNVPPPNAATSGASYFSTGDGSSSFRFNPRSADDIFAEFFGFSTPFGGG 124
Query: 105 GGSSRGRRQRRGEDVVHPLKVSLEDLYLGTSKKLSLSRNVLCSKCSGKGSKSGASMKCAG 164
GG + G+R ++ +D+Y G+G+ G +M
Sbjct: 125 GGGTGGQR--------FASRMFGDDMYASF----------------GEGAGGGGAMHHHH 160
Query: 165 CQ---GTGMKVS-IRHLGPSMIQQMQHPCNECKGTGETINDKDRCPQCKGEKVVQEKKVL 220
KV+ I + P ++ + KGT + + G K +Q +++L
Sbjct: 161 HHHHHAAARKVAPIENKLPCSLEDLY------KGTTKKMKISREIVDVSG-KAMQVEEIL 213
Query: 221 EVHVEKGMQNSQKITFPGEADEAPDTVTGDIVFVLQQKEHPKFKRKSEDLFVEHTLSLTE 280
+ V+ G + KITFP + +E P + D+VF++ +K HP F R+ DL V +SL +
Sbjct: 214 TIGVKPGWKKGTKITFPEKGNEHPGVIPADLVFIIDEKPHPVFTREGNDLIVTQKVSLAD 273
Query: 281 ALCGFQFVLTHLDGRQLLIKSNPGEVVKPDSYKAINDEGMPMYQRPFMKGKLYIHFTVEF 340
AL G+ + LDGR L I V+ P+ + + EGMP+ + KG L I F ++F
Sbjct: 274 ALTGYTANIATLDGRTLTIPIT--NVIHPEYEEVVPKEGMPLQKDQTKKGNLRIKFNIKF 331
Query: 341 PDTLSLDQVKGLEAVL 356
P L+ +Q G + ++
Sbjct: 332 PARLTAEQKAGFKKLI 347
>At5g01390 heat shock protein 40-like
Length = 335
Score = 167 bits (423), Expect = 1e-41
Identities = 113/355 (31%), Positives = 180/355 (49%), Gaps = 38/355 (10%)
Query: 14 YYEILGVSKTASQDDLKKAYKKAAIKNHPDKGGDPEK-----FKELAQAYEVLSDPEKRE 68
+Y++L V ++A+ D+LKKAY+K A+K HPDK + +K FK++++AY+VLSDP+KR
Sbjct: 5 FYKVLEVDRSANDDELKKAYRKLAMKWHPDKNPNNKKEAEAKFKQISEAYDVLSDPQKRA 64
Query: 69 IYDTYGEDALKEGMGGGGGGGHDPFDIFSSFFGGGGGGSSRGRRQRRGEDVVHPLKVSLE 128
IY+ YGE+ L + G GGG+ GG G+S R +D+
Sbjct: 65 IYEQYGEEGLNQAPPPGAGGGYP---------GGSDAGASFRFNPRSADDI-------FS 108
Query: 129 DLYLGTSKKLSLSRNVLCSKCSGKGSKSG----ASMKCAGCQGTGMKVSIRHLGPSMIQQ 184
+ + T + S+ G + G AS + A TG + SI + I++
Sbjct: 109 EFFGFTRPSFGTGSD---SRAGPSGFRYGDDIFASFRAAT---TGGEASIPSRKSAPIER 162
Query: 185 MQHPCNE---CKGTGETINDKDRCPQCKGEKVVQEKKVLEVHVEKGMQNSQKITFPGEAD 241
Q PC+ KG + + G E+ +L + ++ G + KITF + +
Sbjct: 163 -QLPCSLEDLYKGVSKKMKISRDVLDSSGRPTPVEE-ILTIEIKPGWKKGTKITFLEKGN 220
Query: 242 EAPDTVTGDIVFVLQQKEHPKFKRKSEDLFVEHTLSLTEALCGFQFVLTHLDGRQLLIKS 301
E + D+VF++ +K HP FKR DL V +SL +AL G+ +T LDGR L +
Sbjct: 221 EHRGVIPSDLVFIVDEKPHPVFKRDGNDLVVMQKISLVDALTGYTAQVTTLDGRTLTVPV 280
Query: 302 NPGEVVKPDSYKAINDEGMPMYQRPFMKGKLYIHFTVEFPDTLSLDQVKGLEAVL 356
N V+ P + + EGMP+ + P KG L I F ++FP L+ +Q G++ +L
Sbjct: 281 N--NVISPSYEEVVKGEGMPIPKDPSRKGNLRIRFIIKFPSKLTTEQKSGIKRML 333
>At3g47940 heat shock protein-like protein
Length = 350
Score = 166 bits (421), Expect = 2e-41
Identities = 118/385 (30%), Positives = 176/385 (45%), Gaps = 85/385 (22%)
Query: 14 YYEILGVSKTASQDDLKKAYKKAAIKNHPDKGGDPE------KFKELAQAYEVLSDPEKR 67
YY IL V+ A++DDLKKAYK+ A+ HPDK KFK +++AY+VLSDP+KR
Sbjct: 5 YYNILKVNHNATEDDLKKAYKRLAMIWHPDKNPSTRRDEAEAKFKRISEAYDVLSDPQKR 64
Query: 68 EIYDTYGEDALKEGMGGGGGGGH----------------------------------DPF 93
+IYD YGE+ LK G D
Sbjct: 65 QIYDLYGEEGLKSGKIPNSSSSEASSSSSSSSSRYPHFHQHRPQHPPNASSFRFNPRDAE 124
Query: 94 DIFSSFFGG--GGGGSSRGRRQRRGEDVVHPLKVSLEDLYLGTSKKLSLSRNVLCSKCSG 151
DI++ FFG GGG ++ G R R H + Y G +K+ N L
Sbjct: 125 DIYAEFFGSENGGGSNNAGGRGNRAFRNGH-FNTGGANGYSGEMRKVPAMENPL------ 177
Query: 152 KGSKSGASMKCAGCQGTGMKVSIRHLGPSMIQQMQHPCNECKGTGETINDKDRCPQCKGE 211
VS+ L ++++M+ N +G
Sbjct: 178 -------------------PVSLEDLYKGVVKKMRITRNVYDASG--------------- 203
Query: 212 KVVQEKKVLEVHVEKGMQNSQKITFPGEADEAPDTVTGDIVFVLQQKEHPKFKRKSEDLF 271
+++ E ++L + ++ G + K+TFP + +E P + DIVFV+++K HP +KR DL
Sbjct: 204 RMMVEAEILPIEIKPGWKKGTKLTFPKKGNEEPGIIPADIVFVVEEKPHPVYKRDGNDLL 263
Query: 272 VEHTLSLTEALCGFQFVLTHLDGRQLLIKSNPGEVVKPDSYKAINDEGMPMYQRPFMKGK 331
V ++L EAL G L LDGR L+I E++KPD + +EGMP+ + P KG
Sbjct: 264 VSQEITLLEALTGKTVNLITLDGRTLMIPLT--EIIKPDHEIVVPNEGMPISKEPGKKGN 321
Query: 332 LYIHFTVEFPDTLSLDQVKGLEAVL 356
L + +V++P L+ DQ L+ VL
Sbjct: 322 LKLKLSVKYPSRLTSDQKFELKRVL 346
>At3g17830 DnaJ, putative
Length = 517
Score = 165 bits (417), Expect = 5e-41
Identities = 114/372 (30%), Positives = 185/372 (49%), Gaps = 24/372 (6%)
Query: 12 TRYYEILGVSKTASQDDLKKAYKKAAIKNHPDKGGDP---EKFKELAQAYEVLSDPEKRE 68
T +Y L V++ A+ ++K +Y+K A K HPD +P +KFK+++ AYEVLSD EKR
Sbjct: 62 TDHYSTLNVNRNATLQEIKSSYRKLARKYHPDMNKNPGAEDKFKQISAAYEVLSDEEKRS 121
Query: 69 IYDTYGEDALKEGMGGG--GGGGHDPFDIFSSFFGG-----GGGGSSRG------RRQRR 115
YD +GE L+ G G DPFD++S+FFGG GG G S G ++
Sbjct: 122 AYDRFGEAGLEGDFNGSQDTSPGVDPFDLYSAFFGGSDGFFGGMGESGGMGFDFMNKRSL 181
Query: 116 GEDVVHPLKVSLEDLYLGTSKKLSLSRNVLCSKCSGKGSKSGASMK-CAGCQGTGMKVSI 174
D+ + L++S E+ G +++ +S C C G G+KS S+K C+ C G G +V
Sbjct: 182 DLDIRYDLRLSFEEAVFGVKREIEVSYLETCDGCGGTGAKSSNSIKQCSSCDGKG-RVMN 240
Query: 175 RHLGPSMIQQMQHPCNECKGTGETINDKDRCPQCKGEKVVQEKKVLEVHVEKGMQNSQKI 234
P I C++C G G+TI DK C +C G ++ +K ++V V G+ + +
Sbjct: 241 SQRTPFGIMSQVSTCSKCGGEGKTITDK--CRKCIGNGRLRARKKMDVVVPPGVSDRATM 298
Query: 235 TFPGEAD-EAPDTVTGDIVFVLQQKEHPKFKRKSEDLFVEHTLSLTEALCGFQFVLTHLD 293
GE + + GD+ VLQ E +R+ +L+ + T+A+ G + ++
Sbjct: 299 RIQGEGNMDKRSGRAGDLFIVLQVDEKRGIRREGLNLYSNINIDFTDAILGATTKVETVE 358
Query: 294 GRQLLIKSNPGEVVKPDSYKAINDEGMPMYQRPFMKGKLYIHFTVEFPDTLSLDQVKGLE 353
G + ++ PG +P + +G+P RP ++G + P LS + K +E
Sbjct: 359 G-SMDLRIPPG--TQPGDTVKLPRKGVPDTDRPSIRGDHCFVVKISIPKKLSERERKLVE 415
Query: 354 AVLPAKPSSQLT 365
+ SS T
Sbjct: 416 EFSSLRRSSSST 427
>At3g08910 putative heat shock protein
Length = 323
Score = 162 bits (411), Expect = 2e-40
Identities = 128/377 (33%), Positives = 176/377 (45%), Gaps = 101/377 (26%)
Query: 14 YYEILGVSKTASQDDLKKAYKKAAIKNHPDKGGDPEK-----FKELAQAYEVLSDPEKRE 68
YY++L V + A DDLKKAY+K A+K HPDK + +K FK++++AY+VLSDP+KR
Sbjct: 5 YYKVLQVDRNAKDDDLKKAYRKLAMKWHPDKNPNNKKDAEAKFKQISEAYDVLSDPQKRA 64
Query: 69 IYDTYGEDALKE-----GMGGG---GG-----GGHDPFDIFSSFFG-------GGGGGSS 108
IYD YGE+ L G GGG GG G DIFS FFG G G S
Sbjct: 65 IYDQYGEEGLTSQAPPPGAGGGFSDGGASFRFNGRSADDIFSEFFGFTRPFGDSRGAGPS 124
Query: 109 RGRR------------QRRGEDVVHPLKVSLEDLYLGTSKKLSLSRNVLCSKCSGKGSKS 156
G R R+ + L SLEDLY G SKK+ +SR+VL S SG+
Sbjct: 125 NGFRFAEDVFSSNVVPPRKAAPIERQLPCSLEDLYKGVSKKMKISRDVLDS--SGR---- 178
Query: 157 GASMKCAGCQGTGMKVSIRHLGPSMIQQMQHPCNECKGTGETINDKDRCPQCKGEKVVQE 216
P+ ++++ TI K KG K+
Sbjct: 179 ----------------------PTTVEEIL-----------TIEIKPGWK--KGTKI--- 200
Query: 217 KKVLEVHVEKGMQNSQKITFPGEADEAPDTVTGDIVFVLQQKEHPKFKRKSEDLFVEHTL 276
EKG N Q+ P D+VF++ +K H FKR DL + +
Sbjct: 201 -----TFPEKG--NEQRGIIP-----------SDLVFIVDEKPHAVFKRDGNDLVMTQKI 242
Query: 277 SLTEALCGFQFVLTHLDGRQLLIKSNPGEVVKPDSYKAINDEGMPMYQRPFMKGKLYIHF 336
L EAL G+ ++ LDGR + + N V+ P + + EGMP+ + P KG L I F
Sbjct: 243 PLVEALTGYTAQVSTLDGRSVTVPIN--NVISPSYEEVVKGEGMPIPKDPSKKGNLRIKF 300
Query: 337 TVEFPDTLSLDQVKGLE 353
TV+FP L+ +Q G++
Sbjct: 301 TVKFPSRLTTEQKSGIK 317
>At1g10350 putative heat-shock protein
Length = 349
Score = 154 bits (390), Expect = 7e-38
Identities = 117/373 (31%), Positives = 178/373 (47%), Gaps = 61/373 (16%)
Query: 14 YYEILGVSKTASQDDLKKAYKKAAIKNHPDKGGDPEK-----FKELAQAYEVLSDPEKRE 68
YY +L V++ A++DDLKK+Y++ A+K HPDK +K FK++++AY+VLSDP++R+
Sbjct: 5 YYNVLKVNRNANEDDLKKSYRRMAMKWHPDKNPTSKKEAEAKFKQISEAYDVLSDPQRRQ 64
Query: 69 IYDTYGEDALKEG--MGGGGGGGH------------------DPFDIFSSFFGGGG---G 105
IYD YGE+ LK H D DIF+ FFG G G
Sbjct: 65 IYDQYGEEGLKSTDLPTAAETAAHQQQRSYSSSNSEFRYYPRDAEDIFAEFFGESGDAFG 124
Query: 106 GSSRGRRQRRGEDVVHPLKVSLEDLYLGTSKKLSLSRNVLCSKCSGKGSKSGASMKCAGC 165
G S GR + G D G ++ K + GS++
Sbjct: 125 GGSSGRTRGDGGD--------------GGGRRF---------KSAEAGSQANRKTPPTNK 161
Query: 166 QGTGMKVSIRHLGPSMIQQMQHPCNEC-KGTGETINDKDRCPQCKGE-KVVQEKKVLEVH 223
+ T P++ ++ E KG + + P G+ K VQE +L++
Sbjct: 162 KTTP---PANRKAPAIESKLACTLEELYKGAKKKMRISRVVPDDFGKPKTVQE--ILKID 216
Query: 224 VEKGMQNSQKITFPGEADEAPDTVTGDIVFVLQQKEHPKFKRKSEDLFVEHTLSLTEALC 283
++ G + KITFP + ++ P D++FV+ +K H FKR DL +E +SL +AL
Sbjct: 217 IKPGWKKGTKITFPEKGNQEPGVTPADLIFVVDEKPHSVFKRDGNDLILEKKVSLIDALT 276
Query: 284 GFQFVLTHLDGRQLLIKSNPGEVVKPDSYKAINDEGMPMYQRPFMKGKLYIHFTVEFPDT 343
G +T LDGR L I ++VKP I +EGMP + P +G L + F + FP
Sbjct: 277 GLTISVTTLDGRSLTIPVL--DIVKPGQEIVIPNEGMPT-KDPLKRGDLRVTFEILFPSR 333
Query: 344 LSLDQVKGLEAVL 356
L+ +Q L+ VL
Sbjct: 334 LTSEQKNDLKRVL 346
>At5g25530 heat-shock protein - like
Length = 347
Score = 139 bits (350), Expect = 3e-33
Identities = 111/361 (30%), Positives = 166/361 (45%), Gaps = 38/361 (10%)
Query: 14 YYEILGVSKTASQDDLKKAYKKAAIKNHPDKGGDPE-----KFKELAQAYE--------V 60
YY+IL V++ A++DDLKK+Y+K A+K HPDK + + KFK++++AYE V
Sbjct: 5 YYDILKVNRNATEDDLKKSYRKLAMKWHPDKNPNTKTEAEAKFKQISEAYEAKYEVMFQV 64
Query: 61 LSDPEKREIYDTYGEDALKEGMGGGGGGGHDPFDIFSSFFGGGGGGSSRGRRQRRGEDVV 120
LSDP+KR +YD YGE+ L + G G + G + G R ED+
Sbjct: 65 LSDPQKRAVYDQYGEEGLSDMPPPGSTGNN---------------GRAGGFNPRNAEDIF 109
Query: 121 HPLKVSLEDLYLGTSKKLSLSRNVLCSKCSGKGSKSGASMKCAGCQGTGMKV-SIRHLGP 179
S G R++ G G G G + + S P
Sbjct: 110 AEFFGSSP---FGFGSAGGPGRSMRFQSDGGGGMFGGFGGGNNGSENNIFRTYSEGTPAP 166
Query: 180 SMIQQMQH--PCN-ECKGTGETINDK-DRCPQCKGEKVVQEKKVLEVHVEKGMQNSQKIT 235
++ PC+ E +G T K R + QE ++L + V+ G + KI
Sbjct: 167 KKPPPVESKLPCSLEELYSGSTRKMKISRSIVDANGRQAQETEILTIVVKPGWKKGTKIK 226
Query: 236 FPGEADEAPDTVTGDIVFVLQQKEHPKFKRKSEDLFVEHTLSLTEALCGFQFVLTHLDGR 295
FP + +E + + D+VFV+ +K H F R DL ++L EA+ G + LDGR
Sbjct: 227 FPDKGNEQVNQLPADLVFVIDEKPHDLFTRDGNDLITSRRVTLAEAIGGTTVNINTLDGR 286
Query: 296 QLLIKSNPGEVVKPDSYKAINDEGMPMYQRPFMKGKLYIHFTVEFPDTLSLDQVKGLEAV 355
L + E+V P + EGMP+ + P KG L I F V+FP L+ +Q L+ V
Sbjct: 287 NLPV--GVAEIVSPGYEFVVPGEGMPIAKEPRNKGDLKIKFDVQFPARLTTEQKSALKRV 344
Query: 356 L 356
L
Sbjct: 345 L 345
>At1g28210 AtJ1 (atj)
Length = 427
Score = 137 bits (344), Expect = 1e-32
Identities = 105/373 (28%), Positives = 171/373 (45%), Gaps = 37/373 (9%)
Query: 9 SDSTRYYEILGVSKTASQDDLKKAYKKAAIKNHPDKGGD----PEKFKELAQAYEVLSDP 64
S + YY++LGVS A+++++KK++ + A K HPD + KF+E+ +AYE L +
Sbjct: 44 SSARNYYDVLGVSPKATREEIKKSFHELAKKFHPDTNRNNPSAKRKFQEIREAYETLGNS 103
Query: 65 EKREIYDTYGEDALKEGMGGGGGGGHDPF---------DIFSSFFGGGGGGSSRGRRQRR 115
E+RE YD GG + F D F F S +
Sbjct: 104 ERREEYDKL--QYRNSDYVNNDGGDSERFRRAYQSNFSDTFHKIF------SEIFENNQI 155
Query: 116 GEDVVHPLKVSLEDLYLGTSKKLSLSRNVLCSKCSGKGSKSGASMK-CAGCQGTGMKVSI 174
D+ L +SL + G +K+LS V C C G G S A+M C C+G G +V+I
Sbjct: 156 KPDIRVELSLSLSEAAEGCTKRLSFDAYVFCDSCDGLGHPSDAAMSICPTCRGVG-RVTI 214
Query: 175 RHLGPSMIQQMQHPCNECKGTGETINDKDRCPQCKGEKVVQEKKVLEVHVEKGMQNSQKI 234
S C CKGTG I K+ C C+G +V+ K E+ + G+++ I
Sbjct: 215 PPFTAS--------CQTCKGTGHII--KEYCMSCRGSGIVEGTKTAELVIPGGVESEATI 264
Query: 235 TFPGEADEAPDT-VTGDIVFVLQQKEHPKFKRKSEDLFVEHTLSLTEALCGFQFVLTHLD 293
T G + + T G++ L+ F R D++V+ +S T+A+ G + V+ L
Sbjct: 265 TIVGAGNVSSRTSQPGNLYIKLKVANDSTFTRDGSDIYVDANISFTQAILGGKVVVPTLS 324
Query: 294 GRQLLIKSNPGEVVKPDSYKAINDEGMPMYQRPFMKGKLYIHFTVEFPDTLSLDQVKGLE 353
G+ I+ + + +PD + +G+P G Y+ F V FP ++ Q LE
Sbjct: 325 GK---IQLDIPKGTQPDQLLVLRGKGLPKQGFFVDHGDQYVRFRVNFPTEVNERQRAILE 381
Query: 354 AVLPAKPSSQLTD 366
+ +++L+D
Sbjct: 382 EFAKEEINNELSD 394
>At2g20550 putative heat shock protein
Length = 284
Score = 108 bits (271), Expect = 4e-24
Identities = 87/281 (30%), Positives = 128/281 (44%), Gaps = 28/281 (9%)
Query: 85 GGGGGHDPFDIFSSFFGGGGGGSSRGRRQRRGE--DVVHPL----KVSLEDLYLGTSKKL 138
G G + DI+S FFG R + E +V +P K + Y G
Sbjct: 22 GPSGPRNADDIYSEFFGVSSPSGPRNKDDIFSEFFEVSNPSGPRDKDDISAEYFGVPSP- 80
Query: 139 SLSRNVLCSKCSGKGSKSGASMKCAGCQGTGMKVSIRHLGPSMIQQMQHPCNE---CKGT 195
SG GS SG G GT R P + + PC+ KGT
Sbjct: 81 -----------SGSGS-SGGREGGGGGGGTMHHGGARKAAPV---EKKLPCSLEDLYKGT 125
Query: 196 GETINDKDRCPQCKGEKVVQEKKVLEVHVEKGMQNSQKITFPGEADEAPDTVTGDIVFVL 255
+ + G K Q +++L V V+ G + KITF + +E P + D+VF++
Sbjct: 126 TKKMKISREIAGVFG-KTTQVQEILTVDVKPGWKTGTKITFSEKGNEQPGVIPADLVFII 184
Query: 256 QQKEHPKFKRKSEDLFVEHTLSLTEALCGFQFVLTHLDGRQLLIKSNPGEVVKPDSYKAI 315
+K HP F R+ DL V +S+ EA G+ LT LDGR+L I N V+ P+ + +
Sbjct: 185 DEKPHPVFTREGNDLVVTQKISVLEAFTGYTVNLTTLDGRRLTIPVN--TVIHPEYVEVV 242
Query: 316 NDEGMPMYQRPFMKGKLYIHFTVEFPDTLSLDQVKGLEAVL 356
+EGMP+ + KG L I F ++FP TL+ +Q GL+ +L
Sbjct: 243 PNEGMPLQKDQAKKGNLRIKFNIKFPTTLTSEQKTGLKKLL 283
Score = 39.7 bits (91), Expect = 0.003
Identities = 37/139 (26%), Positives = 57/139 (40%), Gaps = 29/139 (20%)
Query: 6 PKKSDSTRYYEILGVSKTASQDDLKKAYKKAAIKNHPDKGGDPEKFKELAQAYEVLSDPE 65
P+ +D Y E GVS + + + + ++P D + +++ Y + P
Sbjct: 26 PRNADDI-YSEFFGVSSPSGPRNKDDIFSEFFEVSNPSGPRDKD---DISAEYFGVPSPS 81
Query: 66 KREIYDTYGEDALKEGMGGGGGGGHDPFDIFSSFFGGGGGGSSRGRRQRRGEDVVHPLKV 125
G +EG GGGGG H GG+ R+ V L
Sbjct: 82 GS------GSSGGREGGGGGGGTMHH-------------GGA------RKAAPVEKKLPC 116
Query: 126 SLEDLYLGTSKKLSLSRNV 144
SLEDLY GT+KK+ +SR +
Sbjct: 117 SLEDLYKGTTKKMKISREI 135
>At5g03160 unknown protein
Length = 482
Score = 96.3 bits (238), Expect = 3e-20
Identities = 56/108 (51%), Positives = 73/108 (66%), Gaps = 14/108 (12%)
Query: 4 RAPKKSDSTRYYEILGVSKTASQDDLKKAYKKAAIKNHPDKG-GDPE----KFKELAQAY 58
+A K S +Y+ILG+S+TAS ++KKAYKK A++ HPDK G+ E KF+E+A AY
Sbjct: 361 KALKMSKRKDWYKILGISRTASISEIKKAYKKLALQWHPDKNVGNREEAENKFREIAAAY 420
Query: 59 EVLSDPEKREIYDTYGEDALKEGMGGGGGGGHDPFDIFSSFFGGGGGG 106
E+L D +KR +D GED E MGGGGGGG++P F GGGGGG
Sbjct: 421 EILGDDDKRARFDR-GEDL--EDMGGGGGGGYNP------FHGGGGGG 459
>At1g44160 putative protein
Length = 352
Score = 88.6 bits (218), Expect = 6e-18
Identities = 51/165 (30%), Positives = 89/165 (53%), Gaps = 3/165 (1%)
Query: 192 CKGTGETINDKDRCPQCKGEKVVQEKKVLEVHVEKGMQNSQKITFPGEADEAPDTVTGDI 251
C G + I K GEK +E++++E+ V+ G + K+TF G+ +EA +V D+
Sbjct: 187 CNGCTKKIKIKRDVITSLGEKC-EEEEMVEIKVKPGWKGGTKVTFEGKGNEAMRSVPADL 245
Query: 252 VFVLQQKEHPKFKRKSEDLFVEHTLSLTEALCGFQFVLTHLDGRQLLIKSNPGEVVKPDS 311
FV+ +KEH FKR+ +DL + +SL EAL G + + LDG + ++ +V+ P
Sbjct: 246 TFVIVEKEHEVFKREGDDLEMAVEVSLLEALTGCELSVALLDGDNMRLRIE--DVIHPGY 303
Query: 312 YKAINDEGMPMYQRPFMKGKLYIHFTVEFPDTLSLDQVKGLEAVL 356
+ +GMP + +G L + F +FP L+ +Q + ++L
Sbjct: 304 VTVVQGKGMPNLKEKGKRGDLRVRFRTKFPQHLTDEQRAEIHSIL 348
>At1g11040 DnaJ isolog
Length = 438
Score = 87.8 bits (216), Expect = 1e-17
Identities = 48/136 (35%), Positives = 78/136 (57%), Gaps = 2/136 (1%)
Query: 213 VVQEKKVLEVHVEKGMQNSQKITFPGEADEAPDTVTGDIVFVLQQKEHPKFKRKSEDLFV 272
++Q++++L V+++ G + KITF G +E P + DI FV+++K HP FKR+ +DL +
Sbjct: 291 IMQQEEMLRVNIQPGWKKGTKITFEGVGNEKPGYLPEDITFVVEEKRHPLFKRRGDDLEI 350
Query: 273 EHTLSLTEALCGFQFVLTHLDGRQLLIKSNPGEVVKPDSYKAINDEGMPMYQRPFMKGKL 332
+ L +AL G + + L G + I G+V+ KAI +GMP + +G L
Sbjct: 351 AVEIPLLKALTGCKLSVPLLSGESMSI--TVGDVIFHGFEKAIKGQGMPNAKEEGKRGDL 408
Query: 333 YIHFTVEFPDTLSLDQ 348
I F V FP+ LS +Q
Sbjct: 409 RITFLVNFPEKLSEEQ 424
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.314 0.134 0.389
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,241,311
Number of Sequences: 26719
Number of extensions: 495853
Number of successful extensions: 3880
Number of sequences better than 10.0: 223
Number of HSP's better than 10.0 without gapping: 132
Number of HSP's successfully gapped in prelim test: 96
Number of HSP's that attempted gapping in prelim test: 2531
Number of HSP's gapped (non-prelim): 788
length of query: 416
length of database: 11,318,596
effective HSP length: 102
effective length of query: 314
effective length of database: 8,593,258
effective search space: 2698283012
effective search space used: 2698283012
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 61 (28.1 bits)
Medicago: description of AC147405.6


